Download This PDF File
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Algorithms for Computational Biology 8Th International Conference, Alcob 2021 Missoula, MT, USA, June 7–11, 2021 Proceedings
Lecture Notes in Bioinformatics 12715 Subseries of Lecture Notes in Computer Science Series Editors Sorin Istrail Brown University, Providence, RI, USA Pavel Pevzner University of California, San Diego, CA, USA Michael Waterman University of Southern California, Los Angeles, CA, USA Editorial Board Members Søren Brunak Technical University of Denmark, Kongens Lyngby, Denmark Mikhail S. Gelfand IITP, Research and Training Center on Bioinformatics, Moscow, Russia Thomas Lengauer Max Planck Institute for Informatics, Saarbrücken, Germany Satoru Miyano University of Tokyo, Tokyo, Japan Eugene Myers Max Planck Institute of Molecular Cell Biology and Genetics, Dresden, Germany Marie-France Sagot Université Lyon 1, Villeurbanne, France David Sankoff University of Ottawa, Ottawa, Canada Ron Shamir Tel Aviv University, Ramat Aviv, Tel Aviv, Israel Terry Speed Walter and Eliza Hall Institute of Medical Research, Melbourne, VIC, Australia Martin Vingron Max Planck Institute for Molecular Genetics, Berlin, Germany W. Eric Wong University of Texas at Dallas, Richardson, TX, USA More information about this subseries at http://www.springer.com/series/5381 Carlos Martín-Vide • Miguel A. Vega-Rodríguez • Travis Wheeler (Eds.) Algorithms for Computational Biology 8th International Conference, AlCoB 2021 Missoula, MT, USA, June 7–11, 2021 Proceedings 123 Editors Carlos Martín-Vide Miguel A. Vega-Rodríguez Rovira i Virgili University University of Extremadura Tarragona, Spain Cáceres, Spain Travis Wheeler University of Montana Missoula, MT, USA ISSN 0302-9743 ISSN 1611-3349 (electronic) Lecture Notes in Bioinformatics ISBN 978-3-030-74431-1 ISBN 978-3-030-74432-8 (eBook) https://doi.org/10.1007/978-3-030-74432-8 LNCS Sublibrary: SL8 – Bioinformatics © Springer Nature Switzerland AG 2021 This work is subject to copyright. -

Microblogging the ISMB: a New Approach to Conference Reporting
Message from ISCB Microblogging the ISMB: A New Approach to Conference Reporting Neil Saunders1*, Pedro Beltra˜o2, Lars Jensen3, Daniel Jurczak4, Roland Krause5, Michael Kuhn6, Shirley Wu7 1 School of Molecular and Microbial Sciences, University of Queensland, St. Lucia, Brisbane, Queensland, Australia, 2 Department of Cellular and Molecular Pharmacology, University of California San Francisco, San Francisco, California, United States of America, 3 Novo Nordisk Foundation Center for Protein Research, Panum Institute, Copenhagen, Denmark, 4 Department of Bioinformatics, University of Applied Sciences, Hagenberg, Freistadt, Austria, 5 Max-Planck-Institute for Molecular Genetics, Berlin, Germany, 6 European Molecular Biology Laboratory, Heidelberg, Germany, 7 Stanford Medical Informatics, Stanford University, Stanford, California, United States of America Cameron Neylon entitled FriendFeed for Claire Fraser-Liggett opened the meeting Scientists: What, Why, and How? (http:// with a review of metagenomics and an blog.openwetware.org/scienceintheopen/ introduction to the human microbiome 2008/06/12/friendfeed-for-scientists-what- project (http://friendfeed.com/search?q = why-and-how/) for an introduction. room%3Aismb-2008+microbiome+OR+ We—a group of science bloggers, most fraser). The subsequent Q&A session of whom met in person for the first time at covered many of the exciting challenges The International Conference on Intel- ISMB 2008—found FriendFeed a remark- for those working in this field. Clearly, ligent Systems for Molecular Biology -

Complete Vertebrate Mitogenomes Reveal Widespread Gene Duplications and Repeats
bioRxiv preprint doi: https://doi.org/10.1101/2020.06.30.177956; this version posted July 1, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. Complete vertebrate mitogenomes reveal widespread gene duplications and repeats Giulio Formenti+*1,2,3, Arang Rhie4, Jennifer Balacco1, Bettina Haase1, Jacquelyn Mountcastle1, Olivier Fedrigo1, Samara Brown2,3, Marco Capodiferro5, Farooq O. Al-Ajli6,7,8, Roberto Ambrosini9, Peter Houde10, Sergey Koren4, Karen Oliver11, Michelle Smith11, Jason Skelton11, Emma Betteridge11, Jale Dolucan11, Craig Corton11, Iliana Bista11, James Torrance11, Alan Tracey11, Jonathan Wood11, Marcela Uliano-Silva11, Kerstin Howe11, Shane McCarthy12, Sylke Winkler13, Woori Kwak14, Jonas Korlach15, Arkarachai Fungtammasan16, Daniel Fordham17, Vania Costa17, Simon Mayes17, Matteo Chiara18, David S. Horner18, Eugene Myers13, Richard Durbin12, Alessandro Achilli5, Edward L. Braun19, Adam M. Phillippy4, Erich D. Jarvis*1,2,3, and The Vertebrate Genomes Project Consortium +First author *Corresponding authors; [email protected]; [email protected] Author affiliations 1. The Vertebrate Genome Lab, Rockefeller University, New York, NY, USA 2. Laboratory of Neurogenetics of Language, Rockefeller University, New York, NY, USA 3. The Howards Hughes Medical Institute, Chevy Chase, MD, USA 4. Genome Informatics Section, Computational and Statistical Genomics Branch, National Human Genome Research Institute, National Institutes of Health, Bethesda, MD USA 5. Department of Biology and Biotechnology “L. Spallanzani”, University of Pavia, Pavia, Italy 6. Monash University Malaysia Genomics Facility, School of Science, Selangor Darul Ehsan, Malaysia 7. -

PROGRAM CHAIR: Eben Rosenthal, MD PROFFERED PAPERS CHAIR: Ellie Maghami, MD POSTER CHAIR: Maie St
AHNS 10TH INTERNATIONAL CONFERENCE ON HEAD & NECK CANCER “Survivorship through Quality & Innovation” JULY 22-25, 2021 • VIRTUAL CONFERENCE AHNS PRESIDENT: Cherie-Ann Nathan, MD, FACS CONFERENCE/DEVELOPMENT CHAIR: Robert Ferris, MD, PhD PROGRAM CHAIR: Eben Rosenthal, MD PROFFERED PAPERS CHAIR: Ellie Maghami, MD POSTER CHAIR: Maie St. John, MD Visit www.ahns2021.org for more information. WELCOME LETTER Dear Colleagues, The American Head and Neck Society (AHNS) is pleased to invite you to the virtual AHNS 10th International Conference on Head and Neck Cancer, which will be held July 22-25, 2021. The theme is Survivorship through Quality & Innovation and the scientific program has been thoughtfully designed to bring together all disciplines related to the treatment of head and neck cancer. Our assembled group of renowned head and neck surgeons, radiologists and oncologists have identified key areas of interest and major topics for us to explore. The entire conference will be presented LIVE online July 22-25, 2021. We encourage you to attend the live sessions in order to engage with the faculty and your colleagues. After the live meeting, all of the meeting content will be posted on the conference site and remain open for on-demand viewing through October 1, 2021. Attendees may earn up to 42.25 AMA PRA Category 1 Credit(s)TM as well as earn re- quired annual part II self-assessment credit in the American Board of Otolaryngology – Head and Neck Surgery’s Continu- ing Certification program (formerly known as MOC). At the conclusion of the activity, -

The Johns Hopkins University
THE JOHNS HOPKINS UNIVERSITY COMMENCEMENT 2019 Conferring of degrees at the close of the 143rd academic year MAY 23, 2019 CONTENTS Order of Candidate Procession ......................................................... 11 Order of Procession .......................................................................... 12 Order of Events ................................................................................. 13 Conferring of Degrees ....................................................................... 14 Honorary Degrees ............................................................................. 17 Academic Garb .................................................................................. 14 Awards ............................................................................................... 16 Honor Societies ................................................................................. 28 Student Honors ................................................................................. 34 Candidates for Degrees ..................................................................... 43 Divisional Ceremonies Information ............................................... 121 Please note that while all degrees are conferred, only doctoral and bachelor’s graduates process across the stage. Taking photos from your seats during the ceremony is allowed, but we request that guests respect one another’s comfort and enjoyment by not standing and blocking other people’s views. Photos of graduates can be purchased from GradImages®: gradimages.com -

LNBI 9044), Entitled “Advances on Computational Intelligence
Lecture Notes in Bioinformatics 9044 Subseries of Lecture Notes in Computer Science LNBI Series Editors Sorin Istrail Brown University, Providence, RI, USA Pavel Pevzner University of California, San Diego, CA, USA Michael Waterman University of Southern California, Los Angeles, CA, USA LNBI Editorial Board Alberto Apostolico Georgia Institute of Technology, Atlanta, GA, USA Søren Brunak Technical University of Denmark Kongens Lyngby, Denmark Mikhail S. Gelfand IITP, Research and Training Center on Bioinformatics, Moscow, Russia Thomas Lengauer Max Planck Institute for Informatics, Saarbrücken, Germany Satoru Miyano University of Tokyo, Japan Eugene Myers Max Planck Institute of Molecular Cell Biology and Genetics Dresden, Germany Marie-France Sagot Université Lyon 1, Villeurbanne, France David Sankoff University of Ottawa, Canada Ron Shamir Tel Aviv University, Ramat Aviv, Tel Aviv, Israel Terry Speed Walter and Eliza Hall Institute of Medical Research Melbourne, VIC, Australia Martin Vingron Max Planck Institute for Molecular Genetics, Berlin, Germany W. Eric Wong University of Texas at Dallas, Richardson, TX, USA Francisco Ortuño Ignacio Rojas (Eds.) Bioinformatics and Biomedical Engineering Third International Conference, IWBBIO 2015 Granada, Spain, April 15-17, 2015 Proceedings, Part II 13 Volume Editors Francisco Ortuño Ignacio Rojas Universidad de Granada Dpto. de Arquitectura y Tecnología de Computadores (ATC) E.T.S. de Ingenierías en Informática y Telecomunicación, CITIC-UGR Granada, Spain E-mail: {fortuno, irojas}@ugr.es ISSN 0302-9743 e-ISSN 1611-3349 ISBN 978-3-319-16479-3 e-ISBN 978-3-319-16480-9 DOI 10.1007/978-3-319-16480-9 Springer Cham Heidelberg New York Dordrecht London Library of Congress Control Number: 2015934926 LNCS Sublibrary: SL 8 – Bioinformatics © Springer International Publishing Switzerland 2015 This work is subject to copyright. -

Signature of Author
An Analysis of Representations for Protein Structure Prediction by Barbara K. Moore Bryant S.B., S.M., Massachusetts Institute of Technology (1986) Submitted to the Department of Electrical Engineering and Computer Science in partial fulfillment of the requirements for the degree of Doctor of Philosophy at the MASSACHUSETTS INSTITUTE OF TECHNOLOGY September 1994 ( Massachusetts Institute of Technology 1994 SignatureofAuthor ............. ,...... ........-.. .. Department of Electrical Engineering and Computer Science September 2, 1994 Certified by... ..... -- Patrick. H. Winston Professor, Electrical Engineering and Computer Science This Supervisor %-"ULfrtifioAvL.Lv--k &y ho, . Tomas o) no-Perez and Computer Science L! Thp.i. .,nPrvi.qnr Accepted by .. ... , a. ,- · F' drY................... / Frederic~) R. Morgenthaler Departmental /Committee on Graduate Students Iv:r tfF:r _ ;: ! --.. t.8r'(itr i-7etFr An Analysis of Representations for Protein Structure Prediction by Barbara K. Moore Bryant Submitted to the Department of Electrical Engineering and Computer Science on September 2, 1994, in partial fulfillment of the requirements for the degree of Doctor of Philosophy Abstract Protein structure prediction is a grand challenge in' the fields of biology and computer science. Being able to quickly determine the structure of a protein from its amino acid sequence would be extremely useful to biologists interested in elucidating the mechanisms of life and finding ways to cure disease. In spite of a wealth of knowl- edge about proteins and their structure, the structure prediction problem has gone unsolved in the nearly forty years since the first determination of a protein structure by X-ray crystallography. In this thesis, I discuss issues in the representation of protein structure and se- quence for algorithms which perform structure prediction. -

Graphical Pangenomics
Graphical pangenomics Erik Peter Garrison Wellcome Sanger Institute University of Cambridge This dissertation is submitted for the degree of Doctor of Philosophy Fitzwilliam College September 2018 for E2 & E3 Declaration I hereby declare that except where specific reference is made to the work of others, the contents of this dissertation are original and have not been submitted in whole or in part for consideration for any other degree or qualification in this, or any other university. This dissertation is my own work and contains nothing which is the outcome of work done in collaboration with others, except as specified in the text and Acknowledgements. This dissertation contains fewer than 65,000 words including appendices, bibliography, footnotes, tables and equations and has fewer than 150 figures. Erik Peter Garrison September 2018 Graphical pangenomics Erik Peter Garrison Completely sequencing genomes is expensive, and to save costs we often analyze new genomic data in the context of a reference genome. This approach distorts our image of the inferred genome, an effect which we describe as reference bias. To mitigate reference bias, I repurpose graphical models previously used in genome assembly and alignment to serve as a reference system in resequencing. To do so I formalize the concept of a variation graph to link genomes to a graphical model of their mutual alignment that is capable of representing any kind of genomic variation, both small and large. As this model combines both sequence and variation information in one structure it serves as a natural basis for resequencing. By indexing the topology, sequence space, and haplotype space of these graphs and developing generalizations of sequence alignment suitable to them, I am able to use them as reference systems in the analysis of a wide array of genomic systems, from large vertebrate genomes to microbial pangenomes. -

Lecture Notes in Bioinformatics 10847
Lecture Notes in Bioinformatics 10847 Subseries of Lecture Notes in Computer Science LNBI Series Editors Sorin Istrail Brown University, Providence, RI, USA Pavel Pevzner University of California, San Diego, CA, USA Michael Waterman University of Southern California, Los Angeles, CA, USA LNBI Editorial Board Søren Brunak Technical University of Denmark, Kongens Lyngby, Denmark Mikhail S. Gelfand IITP, Research and Training Center on Bioinformatics, Moscow, Russia Thomas Lengauer Max Planck Institute for Informatics, Saarbrücken, Germany Satoru Miyano University of Tokyo, Tokyo, Japan Eugene Myers Max Planck Institute of Molecular Cell Biology and Genetics, Dresden, Germany Marie-France Sagot Université Lyon 1, Villeurbanne, France David Sankoff University of Ottawa, Ottawa, Canada Ron Shamir Tel Aviv University, Ramat Aviv, Tel Aviv, Israel Terry Speed Walter and Eliza Hall Institute of Medical Research, Melbourne, VIC, Australia Martin Vingron Max Planck Institute for Molecular Genetics, Berlin, Germany W. Eric Wong University of Texas at Dallas, Richardson, TX, USA More information about this series at http://www.springer.com/series/5381 Fa Zhang • Zhipeng Cai Pavel Skums • Shihua Zhang (Eds.) Bioinformatics Research and Applications 14th International Symposium, ISBRA 2018 Beijing, China, June 8–11, 2018 Proceedings 123 Editors Fa Zhang Pavel Skums Chinese Academy of Sciences Georgia State University Beijing Atlanta, GA China USA Zhipeng Cai Shihua Zhang Georgia State University Chinese Academy of Sciences Atlanta, GA Beijing USA China ISSN 0302-9743 ISSN 1611-3349 (electronic) Lecture Notes in Bioinformatics ISBN 978-3-319-94967-3 ISBN 978-3-319-94968-0 (eBook) https://doi.org/10.1007/978-3-319-94968-0 Library of Congress Control Number: 2018947451 LNCS Sublibrary: SL8 – Bioinformatics © Springer International Publishing AG, part of Springer Nature 2018 This work is subject to copyright. -
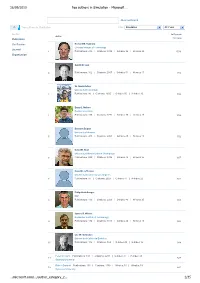
Top Authors in Simulation - Microsoft …
26/09/2010 Top authors in Simulation - Microsoft … Advanced Search Top authors in Simulation Filter: Simulation All Years Author In Domain Author Publication Citations Conference Richard M. Fujimoto Georgia Institute of Technology Journal 1 Publications: 210 | Citations: 3939 | G-Index: 56 | H-Index: 29 1578 Organization Averill M. Law 2 Publications: 102 | Citations: 2657 | G-Index: 51 | H-Index: 17 942 W. David Kelton University of Cincinnati 3 Publications: 86 | Citations: 2605 | G-Index: 50 | H-Index: 15 896 Barry L. Nelson Purdue University 4 Publications: 158 | Citations: 1772 | G-Index: 38 | H-Index: 20 852 Bernard Zeigler University of Arizona 5 Publications: 200 | Citations: 2401 | G-Index: 45 | H-Index: 18 792 David M. Nicol University of Illinois Urbana Champaign 6 Publications: 229 | Citations: 2706 | G-Index: 43 | H-Index: 25 697 David R. Jefferson University of California Los Angeles 7 Publications: 54 | Citations: 2024 | G-Index: 44 | H-Index: 22 681 Philip Heidelberger IBM 8 Publications: 116 | Citations: 2201 | G-Index: 43 | H-Index: 25 603 James R. Wilson Rochester Institute of Technology 9 Publications: 194 | Citations: 1113 | G-Index: 24 | H-Index: 18 586 Lee W. Schruben University of California Berkeley 10 Publications: 102 | Citations: 862 | G-Index: 25 | H-Index: 16 544 Peter W. Glynn Publications: 183 | Citations: 2291 | G-Index: 41 | H-Index: 23 11 527 Stanford University Robert Sargent Publications: 101 | Citations: 1136 | G-Index: 30 | H-Index: 19 12 521 Syracuse University …microsoft.com/…/author_category_2… 1/35 26/09/2010 Top authors in Simulation - Microsoft … Osman Balci Publications: 78 | Citations: 782 | G-Index: 23 | H-Index: 16 13 454 Virginia Polytechnic Institute and State University Jayadev Misra Publications: 136 | Citations: 3325 | G-Index: 56 | H-Index: 26 14 453 University of Texas Austin Rajive Bagrodia Publications: 175 | Citations: 3160 | G-Index: 51 | H-Index: 24 15 435 University of California Los Angeles Richard E. -

Contents U U U
Contents u u u ACM Awards Reception and Banquet, June 2018 .................................................. 2 Introduction ......................................................................................................................... 3 A.M. Turing Award .............................................................................................................. 4 ACM Prize in Computing ................................................................................................. 5 ACM Charles P. “Chuck” Thacker Breakthrough in Computing Award ............. 6 ACM – AAAI Allen Newell Award .................................................................................. 7 Software System Award ................................................................................................... 8 Grace Murray Hopper Award ......................................................................................... 9 Paris Kanellakis Theory and Practice Award ...........................................................10 Karl V. Karlstrom Outstanding Educator Award .....................................................11 Eugene L. Lawler Award for Humanitarian Contributions within Computer Science and Informatics ..........................................................12 Distinguished Service Award .......................................................................................13 ACM Athena Lecturer Award ........................................................................................14 Outstanding Contribution -

Ismb 2010 Organization
Vol. 26 ISMB 2010, pages i2–i6 BIOINFORMATICS doi:10.1093/bioinformatics/btq246 ISMB 2010 ORGANIZATION PROCEEDINGS COMMITTEE CHAIRS Gene Regulation and Transcriptomics Mona Singh, Proceedings Chair, Princeton University, USA Hanah Margalit, The Hebrew University of Jerusalem, Israel Joel S. Bader, Proceedings Co-Chair, Johns Hopkins University, Eric Xing, Carnegie Mellon University, Pittsburgh, USA Baltimore, USA Population Genomics Eleazar Eskin, University of California, Los Angeles, USA Eran Halperin, Tel-Aviv University, Israel PROCEEDINGS AREA CHAIRS Protein Interactions and Molecular Networks Bioimaging Alfonso Valencia, Spanish National Cancer Research Centre, Gene Myers, Howard Hughes Medical Institute, Ashburn, Madrid, Spain USA Roded Sharan, Tel-Aviv University, Israel Robert F. Murphy, Carnegie Mellon University, Pittsburgh, Protein Structure and Function USA Bonnie Berger, Massachusetts Institute of Technology, Cambridge, Databases and Ontologies USA Alex Bateman, Wellcome Trust Sanger Institute, Hinxton, UK Nir Ben-Tal, Tel-Aviv University, Israel Suzanna Lewis, Lawrence Berkeley National Labs, Berkeley, Sequence Analysis USA Michael Brudno, University of Toronto, Canada Disease Models and Epidemiology Cenk Sahinalp, Simon Fraser University, Vancouver, Canada Thomas Lengauer, Max-Planck Institute for Informatics, Text Mining Saarbruken, Germany Andrey Rzhetsky, University of Chicago, USA Yves Moreau, Catholic University of Leuven, Belgium Hagit Shatkay, Queen’s University, Kingston, Canada Evolution and Comparative Genomics