RAD Sequencing Data Reveal a Radiation of Willow Species
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

New Insight Into the Phylogeographic Pattern Of
New insight into the phylogeographic pattern of Liriodendron chinense (Magnoliaceae) revealed by chloroplast DNA: east–west lineage split and genetic mixture within western subtropical China Aihong Yang, Yongda Zhong, Shujuan Liu, Lipan Liu, Tengyun Liu, Yanqiang Li and Faxin Yu The Key Laboratory of Horticultural Plant Genetic and Improvement of Jiangxi, Institute of Biological Resources, Jiangxi Academy of Sciences, Nanchang, Jiangxi, China ABSTRACT Background: Subtropical China is a global center of biodiversity and one of the most important refugia worldwide. Mountains play an important role in conserving the genetic resources of species. Liriodendron chinense is a Tertiary relict tree largely endemic to subtropical China. In this study, we aimed to achieve a better understanding of the phylogeographical pattern of L. chinense andtoexploretheroleofmountainsintheconservationofL. chinense genetic resources. Methods: Three chloroplast regions (psbJ-petA, rpl32-ndhF, and trnK5’-matK) were sequenced in 40 populations of L. chinense for phylogeographical analyses. Relationships among chloroplast DNA (cpDNA) haplotypes were determined using median-joining networks, and genetic structure was examined by spatial analysis of molecular variance (SAMOVA). The ancestral area of the species was reconstructed using the Bayesian binary Markov Chain Monte Carlo (BBM) method according to its geographic distribution and a maximum parsimony (MP) tree based on Bayesian methods. Results: Obvious phylogeographic structure was found in L. chinense. SAMOVA Submitted 13 September 2018 revealed seven groups matching the major landscape features of the L. chinense Accepted 26 December 2018 Published 1 February 2019 distribution area. The haplotype network showed three clades distributed in the eastern, southwestern, and northwestern regions. Separate northern and southern Corresponding author Faxin Yu, [email protected] refugia were found in the Wu Mountains and Yungui Plateau, with genetic admixture in the Dalou Mountains and Wuling Mountains. -

Quantifying Trends of Land Change in Qinghai-Tibet Plateau During 2001–2015
remote sensing Article Quantifying Trends of Land Change in Qinghai-Tibet Plateau during 2001–2015 Chao Wang, Qiong Gao and Mei Yu * Department of Environmental Sciences, University of Puerto Rico, Rio Piedras, San Juan, PR 00936, USA; [email protected] (C.W.); [email protected] (Q.G.) * Correspondence: [email protected]; Tel.: +1-787-764-0000 Received: 1 September 2019; Accepted: 17 October 2019; Published: 20 October 2019 Abstract: The Qinghai-Tibet Plateau (QTP) is among the most sensitive ecosystems to changes in global climate and human activities, and quantifying its consequent change in land-cover land-use (LCLU) is vital for assessing the responses and feedbacks of alpine ecosystems to global climate changes. In this study, we first classified annual LCLU maps from 2001–2015 in QTP from MODIS satellite images, then analyzed the patterns of regional hotspots with significant land changes across QTP, and finally, associated these trends in land change with climate forcing and human activities. The pattern of land changes suggested that forests and closed shrublands experienced substantial expansions in the southeastern mountainous region during 2001–2015 with the expansion of massive meadow loss. Agricultural land abandonment and the conversion by conservation policies existed in QTP, and the newly-reclaimed agricultural land partially offset the loss with the resulting net change of 5.1%. Although the urban area only expanded 586 km2, mainly at the expense of agricultural − land, its rate of change was the largest (41.2%). Surface water exhibited a large expansion of 5866 km2 (10.2%) in the endorheic basins, while mountain glaciers retreated 8894 km2 ( 3.4%) mainly in the − southern and southeastern QTP. -

Beta Diversity Patterns of Fish and Conservation Implications in The
A peer-reviewed open-access journal ZooKeys 817: 73–93 (2019)Beta diversity patterns of fish and conservation implications in... 73 doi: 10.3897/zookeys.817.29337 RESEARCH ARTICLE http://zookeys.pensoft.net Launched to accelerate biodiversity research Beta diversity patterns of fish and conservation implications in the Luoxiao Mountains, China Jiajun Qin1,*, Xiongjun Liu2,3,*, Yang Xu1, Xiaoping Wu1,2,3, Shan Ouyang1 1 School of Life Sciences, Nanchang University, Nanchang 330031, China 2 Key Laboratory of Poyang Lake Environment and Resource Utilization, Ministry of Education, School of Environmental and Chemical Engi- neering, Nanchang University, Nanchang 330031, China 3 School of Resource, Environment and Chemical Engineering, Nanchang University, Nanchang 330031, China Corresponding author: Shan Ouyang ([email protected]); Xiaoping Wu ([email protected]) Academic editor: M.E. Bichuette | Received 27 August 2018 | Accepted 20 December 2018 | Published 15 January 2019 http://zoobank.org/9691CDA3-F24B-4CE6-BBE9-88195385A2E3 Citation: Qin J, Liu X, Xu Y, Wu X, Ouyang S (2019) Beta diversity patterns of fish and conservation implications in the Luoxiao Mountains, China. ZooKeys 817: 73–93. https://doi.org/10.3897/zookeys.817.29337 Abstract The Luoxiao Mountains play an important role in maintaining and supplementing the fish diversity of the Yangtze River Basin, which is also a biodiversity hotspot in China. However, fish biodiversity has declined rapidly in this area as the result of human activities and the consequent environmental changes. Beta diversity was a key concept for understanding the ecosystem function and biodiversity conservation. Beta diversity patterns are evaluated and important information provided for protection and management of fish biodiversity in the Luoxiao Mountains. -

Name Group Description Biennial Biennial These Are
Name Group Description Price Pot Size Nursery Biennial Biennial These are short lived plants that overwinter and flower in their 2.95 9cm SEND second or third year, and should then self seed around the garden in a suitable location..normally several plants per pot for 'pricking out'. Cacti/Succulents Cacti/Succulents We have a collection of varieties in small quantities. For hardy 8.95 1ltr SEND sedums and semperviviums , see 'Rock Plants' , Overwinter in dry frost free shed or greenhouse. House Plants Tender Plants SEND This section includes many half hardy plants for the house, conservervatory , or sheltered position outside in mild areas. We grow these in Kent, but can supply them in Staffs to order. We try and grow many of the old favorites that can now be hard to find. See Annual/biennial for 'patio plants' and 'cacti and succulents' section. Rock Plants Rock Plants Low growing perennials and dwarf shrubs, suited to the front of SEND borders, shady corners etc, where they will not get smothered or hidden by larger perennials and shrubs. Water Plants Water Plants A range of plants that require to grow in wet soil or shallow SEND water. Other moisture loving plants are listed under perennials , ferns and grasses. ABELIA Chinensis Shrub A small shrub with fragrant white rose tinted fls July - Aug 8.95 3 lt MMuc ABELIA gr. "Edward Goucher" Shrub Small semi-evergreen shrub, lilac pink flowers in late 8.95 3lt SEND summer.PF ABELIA Grandiflora (white) Shrub syn 'Lake Maggiore' AGM Evergreen shrub with white flws. 8.95 3lt SEND Likes shelter from winter wind,sun or pt shade Fls. -

Article-Lanzhou
13 by He Yuanqing, Yao Tandong, Cheng Guodong, and Yang Meixue Climatic records in a firn core from an Alpine temperate glacier on Mt. Yulong, southeastern part of the Tibetan Plateau Cold and Arid Regions Environmental and Engineering Research Institute, Chinese Academy of Sciences, Lanzhou 73000, China. Mt. Yulong is the southernmost glacier-covered area in Asian southwestern monsoon climate. Their total area is 11.61 km2. Eurasia, including China. There are 19 sub-tropical The glaciers resemble a group of flying dragons, giving Mt. Yulong temperate glaciers on the mountain, controlled by the (white dragon) as its name. Many explorers, tourists, poets and scientists have described southwestern monsoon climate. In the summer of 1999, Mt. Yulong from different points of view (Ward, 1924; Wissmann, a firn core, 10.10 m long, extending down to glacier ice, 1937). However, as they were unable to cross the extremely steep was recovered in the accumulation area of the largest mountain slopes and the forested area to the glacier above 4000 m a.s.l., some reported data were not correct. Most of the literature has glacier, Baishui No.1. Periodic variations of climatic focussed on the alpine landscape and the snow scenery, and there are signals above 7.8 m depth were apparent, and net accu- few accounts of the existing glaciers. Ren et al (1957) and Luo and mulation of four years was identified by the annual Yang (1963) first reported the distribution of these glaciers and of the late Pleistocene glacial landforms. To clarify the scale of glacia- oscillations of isotopic and ionic composition. -

Diversity and Geographical Pattern of Altitudinal Belts in the Hengduan Mountains in China
J. Mt. Sci. (2010) 7: 123–132 DOI: 10.1007/s11629-010-1011-9 Diversity and Geographical Pattern of Altitudinal Belts in the Hengduan Mountains in China YAO Yonghui, ZHANG Baiping*, HAN Fang, and PANG Yu Institute of Geographic Sciences and Natural Resources Research, Chinese Academy of Sciences, Beijing 100101, China. E-mail: [email protected] * Corresponding author, e-mail: [email protected] © Science Press and Institute of Mountain Hazards and Environment, CAS and Springer-Verlag Berlin Heidelberg 2010 Abstract: This paper analyses the diversity and effect needs further study. In addition, the data spatial pattern of the altitudinal belts in the quality and data accuracy at present also affect to Hengduan Mountains in China. A total of 7 types of some extent the result of quantitative modeling and base belts and 26 types of altitudinal belts are should be improved with RS/GIS in the future. identified in the study region. The main altitudinal belt lines, such as forest line, the upper limit of dark Keywords: Hengduan Mountains; Altitudinal belt coniferous forest and snow line, have similar spectra; Exposure effect; Quadratic model latitudinal and longitudinal spatial patterns, namely, arched quadratic curve model with latitudes and concave quadratic curve model along longitudinal Introduction direction. These patterns can be together called as “Hyperbolic-paraboloid model”, revealing the complexity and speciality of the environment and Globally and generally, the upper and lower ecology in the study region. This result further limits of altitudinal belts vary (increase) from high validates the hypnosis of a common quadratic model latitudes to low latitudes, from continental for spatial pattern of mountain altitudinal belts peripheries to inland areas, and from very humid proposed by the authors. -

Number 3, Spring 1998 Director’S Letter
Planning and planting for a better world Friends of the JC Raulston Arboretum Newsletter Number 3, Spring 1998 Director’s Letter Spring greetings from the JC Raulston Arboretum! This garden- ing season is in full swing, and the Arboretum is the place to be. Emergence is the word! Flowers and foliage are emerging every- where. We had a magnificent late winter and early spring. The Cornus mas ‘Spring Glow’ located in the paradise garden was exquisite this year. The bright yellow flowers are bright and persistent, and the Students from a Wake Tech Community College Photography Class find exfoliating bark and attractive habit plenty to photograph on a February day in the Arboretum. make it a winner. It’s no wonder that JC was so excited about this done soon. Make sure you check of themselves than is expected to seedling selection from the field out many of the special gardens in keep things moving forward. I, for nursery. We are looking to propa- the Arboretum. Our volunteer one, am thankful for each and every gate numerous plants this spring in curators are busy planting and one of them. hopes of getting it into the trade. preparing those gardens for The magnolias were looking another season. Many thanks to all Lastly, when you visit the garden I fantastic until we had three days in our volunteers who work so very would challenge you to find the a row of temperatures in the low hard in the garden. It shows! Euscaphis japonicus. We had a twenties. There was plenty of Another reminder — from April to beautiful seven-foot specimen tree damage to open flowers, but the October, on Sunday’s at 2:00 p.m. -

Essd-2020-71.Pdf
Discussions https://doi.org/10.5194/essd-2020-71 Earth System Preprint. Discussion started: 7 September 2020 Science c Author(s) 2020. CC BY 4.0 License. Open Access Open Data 1 Consolidating the Randolph Glacier Inventory and the Glacier 2 Inventory of China over the Qinghai-Tibetan Plateau and th 3 Investigating Glacier Changes Since the mid-20 Century 4 Xiaowan Liu1,2,3, Zongxue Xu1,2, Hong Yang3,4, Xiuping Li5, Dingzhi Peng1,2 5 1College of Water Sciences, Beijing Normal University, Beijing, 100875, China 6 2Beijiing Key Laboratory of Urban Hydrological Cycle and Sponge City Technology, Beijing, 100875, China 7 3Eawag, Swiss Federal Institute of Aquatic Science and Technology, 8600 Dübendorf, Switzerland 8 4Department of Environmental Science, MGU University of Basel, Petersplatz 1, 4001, Switzerland 9 5Institute of Tibetan Plateau Research, Chinese Academy of Sciences, Beijing, 100101, China 10 Correspondence to: Zongxue Xu ([email protected]) 11 Abstract. Glacier retreat in the Qinghai-Tibetan Plateau (QTP), the ‘third pole of the world’, has attracted the 12 attention of researchers worldwide. Glacier inventories in the 1970s and the 2000s provide valuable information 13 to infer changes in individual glaciers. However, individual glacier volumes are either missing, incomplete or have 14 large errors in these inventories, and thus, the use of these datasets to investigate changes in glaciers in QTP in the 15 past few decades has become a challenge, particularly in the context of climate change. In this study, individual 16 glacier volume data in the Randolph Glacier Inventory version 4.0 (RGI 4.0, 1970s) and the second Glacier 17 Inventory of China (GIC-Ⅱ, 2000s) are recalculated and consolidated using a slope-dependent algorithm based on 18 elevation datasets for the QTP. -

Downloaded from Brill.Com10/07/2021 12:04:04PM Via Free Access © Koninklijke Brill NV, Leiden, 2017
Amphibia-Reptilia 38 (2017): 517-532 A new moth-preying alpine pit viper species from Qinghai-Tibetan Plateau (Viperidae, Crotalinae) Jingsong Shi1,2,∗, Gang Wang3, Xi’er Chen4, Yihao Fang5,LiDing6, Song Huang7,MianHou8,9, Jun Liu1,2, Pipeng Li9 Abstract. The Sanjiangyuan region of Qinghai-Tibetan Plateau is recognized as a biodiversity hotspot of alpine mammals but a barren area in terms of amphibians and reptiles. Here, we describe a new pit viper species, Gloydius rubromaculatus sp. n. Shi, Li and Liu, 2017 that was discovered in this region, with a brief taxonomic revision of the genus Gloydius.The new species can be distinguished from the other congeneric species by the following characteristics: cardinal crossbands on the back, indistinct canthus rostralis, glossy dorsal scales, colubrid-like oval head shape, irregular small black spots on the head scales, black eyes and high altitude distribution (3300-4770 m above sea level). The mitochondrial phylogenetic reconstruction supported the validity of the new species and furthermore reaffirms that G. intermedius changdaoensis, G. halys cognatus, G. h. caraganus and G. h. stejnegeri should be elevated as full species. Gloydius rubromaculatus sp. n. was found to be insectivorous: preying on moths (Lepidoptera, Noctuidae, Sideridis sp.) in the wild. This unusual diet may be one of the key factors to the survival of this species in such a harsh alpine environment. Keywords: Gloydius rubromaculatus sp. n., insectivorous, new species, Sanjiangyuan region. Introduction leopards (Uncia uncia), wild yaks (Bos grun- niens) and Tibetan antelopes (Pantholops hodg- The Sanjiangyuan region (the Source of Three sonii) (Shen and Tan, 2012). -
R Graphics Output
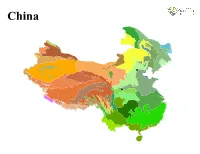
China China LEGEND Previously sampled Malaise trap site Ecoregion Alashan Plateau semi−desert North Tibetan Plateau−Kunlun Mountains alpine desert Altai alpine meadow and tundra Northeast China Plain deciduous forests Altai montane forest and forest steppe Northeast Himalayan subalpine conifer forests Altai steppe and semi−desert Northern Indochina subtropical forests Amur meadow steppe Northern Triangle subtropical forests Bohai Sea saline meadow Northwestern Himalayan alpine shrub and meadows Central China Loess Plateau mixed forests Nujiang Langcang Gorge alpine conifer and mixed forests Central Tibetan Plateau alpine steppe Ordos Plateau steppe Changbai Mountains mixed forests Pamir alpine desert and tundra Changjiang Plain evergreen forests Qaidam Basin semi−desert Da Hinggan−Dzhagdy Mountains conifer forests Qilian Mountains conifer forests Daba Mountains evergreen forests Qilian Mountains subalpine meadows Daurian forest steppe Qin Ling Mountains deciduous forests East Siberian taiga Qionglai−Minshan conifer forests Eastern Gobi desert steppe Rock and Ice Eastern Himalayan alpine shrub and meadows Sichuan Basin evergreen broadleaf forests Eastern Himalayan broadleaf forests South China−Vietnam subtropical evergreen forests Eastern Himalayan subalpine conifer forests Southeast Tibet shrublands and meadows Emin Valley steppe Southern Annamites montane rain forests Guizhou Plateau broadleaf and mixed forests Suiphun−Khanka meadows and forest meadows Hainan Island monsoon rain forests Taklimakan desert Helanshan montane conifer forests -

Vandusen Botanical Garden Plant Sale Catalogue 2016
Welcome to VanDusen Botanical Gardens’ 38th Annual Plant Sale. This catalogue will guide you through the thousands of wonderful plants that we have available for your purchase. We are proud to present the largest plant sale in the lower mainland. All the plants have been carefully selected for you by our many knowledgeable plant sale volunteers and gardening experts. In recognition of the increasing number of people who are gardening in smaller spaces and containers, we are featuring plants suitable for planters, pots and patios. We want to help gardeners explore this whole new world of inspiring and endless design featuring stunning colours, style and impact. Heartfelt thanks to the over 400 volunteers who work long, hard hours and contribute their vast collective gardening knowledge to make this plant sale such a success and therefore an important financial contribution to VanDusen Botanical Garden. Thank you for your support! Margaret Lord, Plant Sale Chair 2016 We wish to thank our sponsors and vendors for their support: Alouette Nursery, B.C. Greenhouse Builders Ltd., Budget Printing, Canadian Springs Co. Ltd., Creperie La Boheme, DeVry Greenhouses, Erica Enterprises Ltd., GardenWorks, Harvest Power, Inline Nurseries, Los Beans Coffee Roasting Co., Mangal Kiss Street Food Services, MedTech EMS Doug House, Oriental Orchids Ltd., Pepsi Cola Canada Ltd., Pops Predatory Plants, Salmon’s Rentals Ltd., Scouts Canada, Snow Mountain Organic Orchards, Solodko Ukrainian Bakery, Southlands Nursery, Taisuco Canada, Sunflower Creperie La Boheme, -

Changes of Major Terrestrial Ecosystems in China Since 1960
Global and Planetary Change 48 (2005) 287–302 www.elsevier.com/locate/gloplacha Changes of major terrestrial ecosystems in China since 1960 Tian Xiang YueT, Ze Meng Fan, Ji Yuan Liu Institute of Geographical Sciences and Natural Resources Research, Chinese Academy of Sciences, Jia No. 11, Datun, Anwai, 100101 Beijing, China Abstract Daily temperature and precipitation data since 1960 are selected from 735 weather stations that are scattered over China. After comparatively analyzing relative interpolation methods, gradient-plus-inverse distance squared (GIDS) is selected to create temperature surfaces and Kriging interpolation method is selected to create precipitation surfaces. Digital elevation model of China is combined into Holdridge Life Zone (HLZ) model on the basis of simulating relationships between temperature and elevation in different regions of China. HLZ model is operated on the created temperature and precipitation surfaces in ARC/ INFO environment. Spatial pattern of major terrestrial ecosystems in China and its change in the four decades of 1960s, 1970s, 1980s and 1990s are analyzed in terms of results from operating HLZ model. The results show that HLZ spatial pattern in China has had a great change since 1960. For instance, nival area and subtropical thorn woodland had a rapid decrease on an average and they might disappear in 159 years and 96 years, respectively, if their areas would decrease at present rate. Alpine dry tundra and cool temperate scrub continuously increased in the four decades and the decadal increase rates are, respectively, 13.1% and 3.4%. HLZ patch connectivity has a continuous increase trend and HLZ diversity has a continuous decrease trend on the average.