The NKG2A–HLA-E Axis As a Novel Checkpoint in the Tumor Microenvironment Linda Borst, Sjoerd H
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Human and Mouse CD Marker Handbook Human and Mouse CD Marker Key Markers - Human Key Markers - Mouse
Welcome to More Choice CD Marker Handbook For more information, please visit: Human bdbiosciences.com/eu/go/humancdmarkers Mouse bdbiosciences.com/eu/go/mousecdmarkers Human and Mouse CD Marker Handbook Human and Mouse CD Marker Key Markers - Human Key Markers - Mouse CD3 CD3 CD (cluster of differentiation) molecules are cell surface markers T Cell CD4 CD4 useful for the identification and characterization of leukocytes. The CD CD8 CD8 nomenclature was developed and is maintained through the HLDA (Human Leukocyte Differentiation Antigens) workshop started in 1982. CD45R/B220 CD19 CD19 The goal is to provide standardization of monoclonal antibodies to B Cell CD20 CD22 (B cell activation marker) human antigens across laboratories. To characterize or “workshop” the antibodies, multiple laboratories carry out blind analyses of antibodies. These results independently validate antibody specificity. CD11c CD11c Dendritic Cell CD123 CD123 While the CD nomenclature has been developed for use with human antigens, it is applied to corresponding mouse antigens as well as antigens from other species. However, the mouse and other species NK Cell CD56 CD335 (NKp46) antibodies are not tested by HLDA. Human CD markers were reviewed by the HLDA. New CD markers Stem Cell/ CD34 CD34 were established at the HLDA9 meeting held in Barcelona in 2010. For Precursor hematopoetic stem cell only hematopoetic stem cell only additional information and CD markers please visit www.hcdm.org. Macrophage/ CD14 CD11b/ Mac-1 Monocyte CD33 Ly-71 (F4/80) CD66b Granulocyte CD66b Gr-1/Ly6G Ly6C CD41 CD41 CD61 (Integrin b3) CD61 Platelet CD9 CD62 CD62P (activated platelets) CD235a CD235a Erythrocyte Ter-119 CD146 MECA-32 CD106 CD146 Endothelial Cell CD31 CD62E (activated endothelial cells) Epithelial Cell CD236 CD326 (EPCAM1) For Research Use Only. -

Early Detection of Peripheral Blood Cell Signature in Children Developing B-Cell Autoimmunity at a Young Age
2024 Diabetes Volume 68, October 2019 Early Detection of Peripheral Blood Cell Signature in Children Developing b-Cell Autoimmunity at a Young Age Henna Kallionpää,1 Juhi Somani,2 Soile Tuomela,1 Ubaid Ullah,1 Rafael de Albuquerque,1 Tapio Lönnberg,1 Elina Komsi,1 Heli Siljander,3,4 Jarno Honkanen,3,4 Taina Härkönen,3,4 Aleksandr Peet,5,6 Vallo Tillmann,5,6 Vikash Chandra,3,7 Mahesh Kumar Anagandula,8 Gun Frisk,8 Timo Otonkoski,3,7 Omid Rasool,1 Riikka Lund,1 Harri Lähdesmäki,2 Mikael Knip,3,4,9,10 and Riitta Lahesmaa1 Diabetes 2019;68:2024–2034 | https://doi.org/10.2337/db19-0287 The appearance of type 1 diabetes (T1D)-associated function before T1D and suggest a potential role for IL32 autoantibodies is the first and only measurable param- in the pathogenesis of T1D. eter to predict progression toward T1D in genetically susceptible individuals. However, autoantibodies indi- cate an active autoimmune reaction, wherein the im- Family and sibling studies in type 1 diabetes (T1D) have mune tolerance is already broken. Therefore, there is implicated a firm genetic predisposition to a locus con- a clear and urgent need for new biomarkers that predict taining HLA class I and class II genes on chromosome the onset of the autoimmune reaction preceding auto- 6 suggesting a role for CD4+ as well as CD8+ T cells in T1D fl antibody positivity or re ect progressive b-cell destruc- pathogenesis (1–3). As much as 30–50% of the genetic risk – tion. Here we report the mRNA sequencing based is conferred by HLA class II molecules, which are crucial in analysis of 306 samples including fractionated samples antigen presentation to CD4+ T cells. -

Single Cell Transcriptome Atlas of Immune Cells in Human Small
bioRxiv preprint doi: https://doi.org/10.1101/721258; this version posted August 1, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. 1 Single cell transcriptome atlas of immune cells in human small 2 intestine and in celiac disease 3 4 Nader Atlasy1,a,4, Anna Bujko2,4, Peter B Brazda1,a, Eva Janssen-Megens1,a , Espen S. 5 Bækkevold2, Jørgen Jahnsen3, Frode L. Jahnsen2, Hendrik G. Stunnenberg1,a,* 6 7 8 1Department of Molecular Biology, Science Faculty, Radboud University, Nijmegen, The 9 Netherlands. 10 2 Department of Pathology, University of Oslo and Oslo University Hospital, Rikshospitalet, 11 Oslo, Norway 12 3 Department of Gastroenterology, Akershus University Hospital and University of Oslo, 13 Oslo, Norway. 14 15 acurrent address: Princess Maxima Centre for Pediatric Oncology, Heidelberglaan 25, 3584 16 CS Utrecht 17 18 19 4These authors contributed equally to this study 20 21 22 *Corresponding author: [email protected] 23 24 25 26 Celiac disease (CeD) is an autoimmune disorder in which ingestion of dietary gluten 27 triggers an immune reaction in the small intestine1,2. The CeD lesion is characterized by 28 crypt hyperplasia, villous atrophy and chronic inflammation with accumulation of 29 leukocytes both in the lamina propria (LP) and in the epithelium3, which eventually 30 leads to destruction of the intestinal epithelium1 and subsequent digestive complications 31 and higher risk of non-hodgkin lymphoma4. A lifetime gluten-free diet is currently the 32 only available treatment5. Gluten-specific LP CD4 T cells and cytotoxic intraepithelial 33 CD8+ T cells are thought to be central in disease pathology1,6-8, however, CeD is a 34 complex immune-mediated disorder and to date the findings are mostly based on 35 analysis of heterogeneous cell populations and on animal models. -
Single-Cell Profiling Identifies Impaired Adaptive NK Cells Expanded After HCMV Reactivation in Haploidentical HSCT
Single-cell profiling identifies impaired adaptive NK cells expanded after HCMV reactivation in haploidentical HSCT Elisa Zaghi, … , Enrico Lugli, Domenico Mavilio JCI Insight. 2021;6(12):e146973. https://doi.org/10.1172/jci.insight.146973. Research Article Hematology Immunology Graphical abstract Find the latest version: https://jci.me/146973/pdf RESEARCH ARTICLE Single-cell profiling identifies impaired adaptive NK cells expanded after HCMV reactivation in haploidentical HSCT Elisa Zaghi,1 Michela Calvi,1,2 Simone Puccio,3 Gianmarco Spata,1 Sara Terzoli,1 Clelia Peano,4 Alessandra Roberto,3 Federica De Paoli,3 Jasper J.P. van Beek,3 Jacopo Mariotti,5 Chiara De Philippis,5 Barbara Sarina,5 Rossana Mineri,6 Stefania Bramanti,5 Armando Santoro,5 Vu Thuy Khanh Le-Trilling,7 Mirko Trilling,7 Emanuela Marcenaro,8 Luca Castagna,5 Clara Di Vito,1,2 Enrico Lugli,3,9 and Domenico Mavilio1,2 1Unit of Clinical and Experimental Immunology, IRCCS Humanitas Research Hospital, Rozzano, Milan, Italy. 2BIOMETRA, Università degli Studi di Milano, Milan, Italy. 3Laboratory of Translational Immunology, 4Institute of Genetic and Biomedical Research, UoS Milan, National Research Council, and Genomic Unit, 5Bone Marrow Transplant Unit, and 6Molecular Biology Section, Clinical Investigation Laboratory, IRCCS Humanitas Research Hospital, Milan, Italy. 7Institute for Virology, University Hospital Essen, University Duisburg-Essen, Essen, Germany. 8Department of Experimental Medicine, University of Genoa, Genoa, Italy. 9Flow Cytometry Core, IRCCS Humanitas Research Hospital, Milan, Italy. Haploidentical hematopoietic stem cell transplantation (h-HSCT) represents an efficient curative approach for patients affected by hematologic malignancies in which the reduced intensity conditioning induces a state of immunologic tolerance between donor and recipient. -

Single-Cell RNA Sequencing Demonstrates the Molecular and Cellular Reprogramming of Metastatic Lung Adenocarcinoma
ARTICLE https://doi.org/10.1038/s41467-020-16164-1 OPEN Single-cell RNA sequencing demonstrates the molecular and cellular reprogramming of metastatic lung adenocarcinoma Nayoung Kim 1,2,3,13, Hong Kwan Kim4,13, Kyungjong Lee 5,13, Yourae Hong 1,6, Jong Ho Cho4, Jung Won Choi7, Jung-Il Lee7, Yeon-Lim Suh8,BoMiKu9, Hye Hyeon Eum 1,2,3, Soyean Choi 1, Yoon-La Choi6,10,11, Je-Gun Joung1, Woong-Yang Park 1,2,6, Hyun Ae Jung12, Jong-Mu Sun12, Se-Hoon Lee12, ✉ ✉ Jin Seok Ahn12, Keunchil Park12, Myung-Ju Ahn 12 & Hae-Ock Lee 1,2,3,6 1234567890():,; Advanced metastatic cancer poses utmost clinical challenges and may present molecular and cellular features distinct from an early-stage cancer. Herein, we present single-cell tran- scriptome profiling of metastatic lung adenocarcinoma, the most prevalent histological lung cancer type diagnosed at stage IV in over 40% of all cases. From 208,506 cells populating the normal tissues or early to metastatic stage cancer in 44 patients, we identify a cancer cell subtype deviating from the normal differentiation trajectory and dominating the metastatic stage. In all stages, the stromal and immune cell dynamics reveal ontological and functional changes that create a pro-tumoral and immunosuppressive microenvironment. Normal resident myeloid cell populations are gradually replaced with monocyte-derived macrophages and dendritic cells, along with T-cell exhaustion. This extensive single-cell analysis enhances our understanding of molecular and cellular dynamics in metastatic lung cancer and reveals potential diagnostic and therapeutic targets in cancer-microenvironment interactions. 1 Samsung Genome Institute, Samsung Medical Center, Seoul 06351, Korea. -

Supplementary Table 1: Adhesion Genes Data Set
Supplementary Table 1: Adhesion genes data set PROBE Entrez Gene ID Celera Gene ID Gene_Symbol Gene_Name 160832 1 hCG201364.3 A1BG alpha-1-B glycoprotein 223658 1 hCG201364.3 A1BG alpha-1-B glycoprotein 212988 102 hCG40040.3 ADAM10 ADAM metallopeptidase domain 10 133411 4185 hCG28232.2 ADAM11 ADAM metallopeptidase domain 11 110695 8038 hCG40937.4 ADAM12 ADAM metallopeptidase domain 12 (meltrin alpha) 195222 8038 hCG40937.4 ADAM12 ADAM metallopeptidase domain 12 (meltrin alpha) 165344 8751 hCG20021.3 ADAM15 ADAM metallopeptidase domain 15 (metargidin) 189065 6868 null ADAM17 ADAM metallopeptidase domain 17 (tumor necrosis factor, alpha, converting enzyme) 108119 8728 hCG15398.4 ADAM19 ADAM metallopeptidase domain 19 (meltrin beta) 117763 8748 hCG20675.3 ADAM20 ADAM metallopeptidase domain 20 126448 8747 hCG1785634.2 ADAM21 ADAM metallopeptidase domain 21 208981 8747 hCG1785634.2|hCG2042897 ADAM21 ADAM metallopeptidase domain 21 180903 53616 hCG17212.4 ADAM22 ADAM metallopeptidase domain 22 177272 8745 hCG1811623.1 ADAM23 ADAM metallopeptidase domain 23 102384 10863 hCG1818505.1 ADAM28 ADAM metallopeptidase domain 28 119968 11086 hCG1786734.2 ADAM29 ADAM metallopeptidase domain 29 205542 11085 hCG1997196.1 ADAM30 ADAM metallopeptidase domain 30 148417 80332 hCG39255.4 ADAM33 ADAM metallopeptidase domain 33 140492 8756 hCG1789002.2 ADAM7 ADAM metallopeptidase domain 7 122603 101 hCG1816947.1 ADAM8 ADAM metallopeptidase domain 8 183965 8754 hCG1996391 ADAM9 ADAM metallopeptidase domain 9 (meltrin gamma) 129974 27299 hCG15447.3 ADAMDEC1 ADAM-like, -

Supplementary Table S5. Differentially Expressed Gene Lists of PD-1High CD39+ CD8 Tils According to 4-1BB Expression Compared to PD-1+ CD39- CD8 Tils
BMJ Publishing Group Limited (BMJ) disclaims all liability and responsibility arising from any reliance Supplemental material placed on this supplemental material which has been supplied by the author(s) J Immunother Cancer Supplementary Table S5. Differentially expressed gene lists of PD-1high CD39+ CD8 TILs according to 4-1BB expression compared to PD-1+ CD39- CD8 TILs Up- or down- regulated genes in Up- or down- regulated genes Up- or down- regulated genes only PD-1high CD39+ CD8 TILs only in 4-1BBneg PD-1high CD39+ in 4-1BBpos PD-1high CD39+ CD8 compared to PD-1+ CD39- CD8 CD8 TILs compared to PD-1+ TILs compared to PD-1+ CD39- TILs CD39- CD8 TILs CD8 TILs IL7R KLRG1 TNFSF4 ENTPD1 DHRS3 LEF1 ITGA5 MKI67 PZP KLF3 RYR2 SIK1B ANK3 LYST PPP1R3B ETV1 ADAM28 H2AC13 CCR7 GFOD1 RASGRP2 ITGAX MAST4 RAD51AP1 MYO1E CLCF1 NEBL S1PR5 VCL MPP7 MS4A6A PHLDB1 GFPT2 TNF RPL3 SPRY4 VCAM1 B4GALT5 TIPARP TNS3 PDCD1 POLQ AKAP5 IL6ST LY9 PLXND1 PLEKHA1 NEU1 DGKH SPRY2 PLEKHG3 IKZF4 MTX3 PARK7 ATP8B4 SYT11 PTGER4 SORL1 RAB11FIP5 BRCA1 MAP4K3 NCR1 CCR4 S1PR1 PDE8A IFIT2 EPHA4 ARHGEF12 PAICS PELI2 LAT2 GPRASP1 TTN RPLP0 IL4I1 AUTS2 RPS3 CDCA3 NHS LONRF2 CDC42EP3 SLCO3A1 RRM2 ADAMTSL4 INPP5F ARHGAP31 ESCO2 ADRB2 CSF1 WDHD1 GOLIM4 CDK5RAP1 CD69 GLUL HJURP SHC4 GNLY TTC9 HELLS DPP4 IL23A PITPNC1 TOX ARHGEF9 EXO1 SLC4A4 CKAP4 CARMIL3 NHSL2 DZIP3 GINS1 FUT8 UBASH3B CDCA5 PDE7B SOGA1 CDC45 NR3C2 TRIB1 KIF14 TRAF5 LIMS1 PPP1R2C TNFRSF9 KLRC2 POLA1 CD80 ATP10D CDCA8 SETD7 IER2 PATL2 CCDC141 CD84 HSPA6 CYB561 MPHOSPH9 CLSPN KLRC1 PTMS SCML4 ZBTB10 CCL3 CA5B PIP5K1B WNT9A CCNH GEM IL18RAP GGH SARDH B3GNT7 C13orf46 SBF2 IKZF3 ZMAT1 TCF7 NECTIN1 H3C7 FOS PAG1 HECA SLC4A10 SLC35G2 PER1 P2RY1 NFKBIA WDR76 PLAUR KDM1A H1-5 TSHZ2 FAM102B HMMR GPR132 CCRL2 PARP8 A2M ST8SIA1 NUF2 IL5RA RBPMS UBE2T USP53 EEF1A1 PLAC8 LGR6 TMEM123 NEK2 SNAP47 PTGIS SH2B3 P2RY8 S100PBP PLEKHA7 CLNK CRIM1 MGAT5 YBX3 TP53INP1 DTL CFH FEZ1 MYB FRMD4B TSPAN5 STIL ITGA2 GOLGA6L10 MYBL2 AHI1 CAND2 GZMB RBPJ PELI1 HSPA1B KCNK5 GOLGA6L9 TICRR TPRG1 UBE2C AURKA Leem G, et al. -

Supporting Info
advances.sciencemag.org/cgi/content/full/6/29/eaba9589/DC1 Supplementary Materials for Cell invasion in digital microfluidic microgel systems Bingyu B. Li, Erica Y. Scott, M. Dean Chamberlain, Bill T. V. Duong, Shuailong Zhang, Susan J. Done, Aaron R. Wheeler* *Corresponding author. Email: [email protected] Published 15 July 2020, Sci. Adv. 6, eaba9589 (2020) DOI: 10.1126/sciadv.aba9589 This PDF file includes: Figs. S1 to S8 Tables S1 and S2 References Supplementary Figure S1 CIMMS microgels and cell viability. For (a-c) core-shell microgels were formed from 2.4 mg/mL collagen I and 10% BM extract. (a) Cartoon indicating the orientation of the microgel shown in the images. SEM images of (b) the collagen I core (scale bar = 20 µm) and (c) the edge of microgel with core on the "top" and the shell on the "bottom" (scale bar = 100 µm). The pore size of the collagen I core was calculated to be 3.1 m by the method reported by Blackmon et al. (53), which is nearly identical to the pore size reported for the same concentration of collagen I cast in well plates by Lang et al. (54). For (d-f) MDA-MB-231 cells were seeded onto simple collagen I microgels (2.4 mg/mL). (d) Cell count per droplet with seeding density at 200,000 (lt. pink), 400,000 (fuscia) or 600,000 cells/mL (burgundy). Error bars represent ± 1 std. dev. from n = 4 microgels per condition. (e) 3D confocal fluorescent microscopy image of cells stained on day 4 with calcein-AM (live-green) and propidium iodide (dead-red). -

Human Lectins, Their Carbohydrate Affinities and Where to Find Them
biomolecules Review Human Lectins, Their Carbohydrate Affinities and Where to Review HumanFind Them Lectins, Their Carbohydrate Affinities and Where to FindCláudia ThemD. Raposo 1,*, André B. Canelas 2 and M. Teresa Barros 1 1, 2 1 Cláudia D. Raposo * , Andr1 é LAQVB. Canelas‐Requimte,and Department M. Teresa of Chemistry, Barros NOVA School of Science and Technology, Universidade NOVA de Lisboa, 2829‐516 Caparica, Portugal; [email protected] 12 GlanbiaLAQV-Requimte,‐AgriChemWhey, Department Lisheen of Chemistry, Mine, Killoran, NOVA Moyne, School E41 of ScienceR622 Co. and Tipperary, Technology, Ireland; canelas‐ [email protected] NOVA de Lisboa, 2829-516 Caparica, Portugal; [email protected] 2* Correspondence:Glanbia-AgriChemWhey, [email protected]; Lisheen Mine, Tel.: Killoran, +351‐212948550 Moyne, E41 R622 Tipperary, Ireland; [email protected] * Correspondence: [email protected]; Tel.: +351-212948550 Abstract: Lectins are a class of proteins responsible for several biological roles such as cell‐cell in‐ Abstract:teractions,Lectins signaling are pathways, a class of and proteins several responsible innate immune for several responses biological against roles pathogens. such as Since cell-cell lec‐ interactions,tins are able signalingto bind to pathways, carbohydrates, and several they can innate be a immuneviable target responses for targeted against drug pathogens. delivery Since sys‐ lectinstems. In are fact, able several to bind lectins to carbohydrates, were approved they by canFood be and a viable Drug targetAdministration for targeted for drugthat purpose. delivery systems.Information In fact, about several specific lectins carbohydrate were approved recognition by Food by andlectin Drug receptors Administration was gathered for that herein, purpose. plus Informationthe specific organs about specific where those carbohydrate lectins can recognition be found by within lectin the receptors human was body. -

Single-Cell Analysis of Crohn's Disease Lesions Identifies
bioRxiv preprint doi: https://doi.org/10.1101/503102; this version posted December 20, 2018. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Single-cell analysis of Crohn’s disease lesions identifies a pathogenic cellular module associated with resistance to anti-TNF therapy JC Martin1,2,3, G Boschetti1,2,3, C Chang1,2,3, R Ungaro4, M Giri5, LS Chuang5, S Nayar5, A Greenstein6, M. Dubinsky7, L Walker1,2,5,8, A Leader1,2,3, JS Fine9, CE Whitehurst9, L Mbow9, S Kugathasan10, L.A. Denson11, J.Hyams12, JR Friedman13, P Desai13, HM Ko14, I Laface1,2,8, Guray Akturk1,2,8, EE Schadt15,16, S Gnjatic1,2,8, A Rahman1,2,5,8, , M Merad1,2,3,8,17,18*, JH Cho5,17,*, E Kenigsberg1,15,16,17* 1 Precision Immunology Institute, Icahn School of Medicine at Mount Sinai, New York, NY 10029, USA. 2 Tisch Cancer Institute, Icahn School of Medicine at Mount Sinai, New York, NY 10029, USA. 3 Department of Oncological Sciences, Icahn School of Medicine at Mount Sinai, New York, NY 10029, USA. 4 The Dr. Henry D. Janowitz Division of Gastroenterology, Icahn School of Medicine at Mount Sinai, New York City, NY 10029, USA. 5 Charles Bronfman Institute for Personalized Medicine, Icahn School of Medicine at Mount Sinai, New York, NY 10029, USA. 6 Department of Colorectal Surgery, Icahn School of Medicine at Mount Sinai, New York, NY 10029, USA 7 Department of Pediatrics, Susan and Leonard Feinstein IBD Clinical Center, Icahn School of Medicine at Mount Sinai, New York, NY 10029, USA. -

High-Dimensional Analysis Reveals a Distinct Population of Adaptive-Like Tissue
bioRxiv preprint doi: https://doi.org/10.1101/2019.12.20.883785; this version posted December 20, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. 1 High-dimensional analysis reveals a distinct population of adaptive-like tissue- 2 resident NK cells in human lung 3 4 Nicole Marquardt1*, Marlena Scharenberg1, Jeffrey E. Mold2, Joanna Hård2, Eliisa 5 Kekäläinen3,4, Marcus Buggert1, Son Nguyen5,6, Jennifer N. Wilson1, Mamdoh Al- 6 Ameri7, Hans-Gustaf Ljunggren1, Jakob Michaëlsson1 7 8 Running title: Adaptive-like human lung tissue-resident NK cells 9 10 Affiliations: 11 1Center for Infectious Medicine, Department of Medicine, Karolinska Institutet, 12 Stockholm, Sweden 13 2Department of Cell and Molecular Biology, Karolinska Institutet, Stockholm, Sweden 14 3Translational Immunology Research Program & Department of Bacteriology and 15 Immunology, University of Helsinki, Finland 16 4HUSLAB, Division of Clinical Microbiology, Helsinki University Hospital, Helsinki, 17 Finland, 18 5Department of Microbiology, Perelman School of Medicine, University of 19 Pennsylvania, Philadelphia, PA, USA 20 6Institute for Immunology, Perelman School of Medicine, University of Pennsylvania, 21 Philadelphia, PA, USA 22 7Thoracic Surgery, Department of Molecular Medicine and Surgery, Karolinska 23 Institutet, Karolinska University Hospital, Stockholm, Sweden 24 1 bioRxiv preprint doi: https://doi.org/10.1101/2019.12.20.883785; this version posted December 20, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. -
Type of the Paper (Article
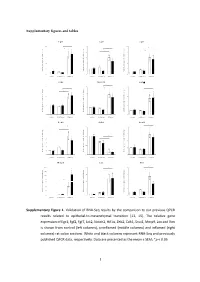
Supplementary figures and tables E g r 1 F g f2 F g f7 1 0 * 5 1 0 * * e e e * g g g * n n n * a a a 8 4 * 8 h h h * c c c d d d * l l l o o o * f f f * n n n o o o 6 3 6 i i i s s s s s s e e e r r r p p p x x x e e e 4 2 4 e e e n n n e e e g g g e e e v v v i i i t t t 2 1 2 a a a l l l e e e R R R 0 0 0 c o n tro l u n in fla m e d in fla m e d c o n tro l u n in fla m e d in fla m e d c o n tro l u n in fla m e d in fla m e d J a k 2 N o tc h 2 H if1 * 3 4 6 * * * e e e g g g n n n a a * * a * h h * h c c c 3 * d d * d l l l * o o o f f 2 f 4 n n n o o o i i i s s s s s s e e e r r 2 r p p p x x x e e e e e e n n n e e 1 e 2 g g g e e 1 e v v v i i i t t t a a a l l l e e e R R R 0 0 0 c o n tro l u n in fla m e d in fla m e d c o n tro l u n in fla m e d in fla m e d c o n tro l u n in fla m e d in fla m e d Z e b 2 C d h 1 S n a i1 * * 7 1 .5 4 * * e e e g g g 6 n n n * a a a * h h h c c c 3 * d d d l l l 5 o o o f f f 1 .0 * n n n * o o o i i i 4 * s s s s s s e e e r r r 2 p p p x x x 3 e e e e e e n n n e e e 0 .5 g g g 2 e e e 1 v v v i i i t t t a a a * l l l e e e 1 * R R R 0 0 .0 0 c o n tro l u n in fla m e d in fla m e d c o n tro l u n in fla m e d in fla m e d c o n tro l u n in fla m e d in fla m e d M m p 9 L o x V im 2 0 0 2 0 8 * * * e e e * g g g 1 5 0 * n n n * a a a * h h h * c c c 1 5 * 6 d d d l l l 1 0 0 o o o f f f n n n o o o i i i 5 0 s s s s s s * e e e r r r 1 0 4 3 0 p p p * x x x e e e * e e e n n n e e e 2 0 g g g e e e 5 2 v v v i i i t t t a a a l l l 1 0 e e e R R R 0 0 0 c o n tro l u n in fla m e d in fla m e d c o n tro l u n in fla m e d in fla m e d c o n tro l u n in fla m e d in fla m e d Supplementary Figure 1.