RNA-Seq Reveals Conservation of Function Among the Yolk Sacs Of
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

3 Embryology and Development
BIOL 6505 − INTRODUCTION TO FETAL MEDICINE 3. EMBRYOLOGY AND DEVELOPMENT Arlet G. Kurkchubasche, M.D. INTRODUCTION Embryology – the field of study that pertains to the developing organism/human Basic embryology –usually taught in the chronologic sequence of events. These events are the basis for understanding the congenital anomalies that we encounter in the fetus, and help explain the relationships to other organ system concerns. Below is a synopsis of some of the critical steps in embryogenesis from the anatomic rather than molecular basis. These concepts will be more intuitive and evident in conjunction with diagrams and animated sequences. This text is a synopsis of material provided in Langman’s Medical Embryology, 9th ed. First week – ovulation to fertilization to implantation Fertilization restores 1) the diploid number of chromosomes, 2) determines the chromosomal sex and 3) initiates cleavage. Cleavage of the fertilized ovum results in mitotic divisions generating blastomeres that form a 16-cell morula. The dense morula develops a central cavity and now forms the blastocyst, which restructures into 2 components. The inner cell mass forms the embryoblast and outer cell mass the trophoblast. Consequences for fetal management: Variances in cleavage, i.e. splitting of the zygote at various stages/locations - leads to monozygotic twinning with various relationships of the fetal membranes. Cleavage at later weeks will lead to conjoined twinning. Second week: the week of twos – marked by bilaminar germ disc formation. Commences with blastocyst partially embedded in endometrial stroma Trophoblast forms – 1) cytotrophoblast – mitotic cells that coalesce to form 2) syncytiotrophoblast – erodes into maternal tissues, forms lacunae which are critical to development of the uteroplacental circulation. -

Meta-Analysis and Inding from Seoul Breast Cancer Study (SEBCS)
The Pharmacogenomics Journal https://doi.org/10.1038/s41397-018-0016-6 ARTICLE Associations between genetic polymorphisms of membrane transporter genes and prognosis after chemotherapy: meta-analysis and finding from Seoul Breast Cancer Study (SEBCS) 1 1 1 1 2 3 4 Ji-Eun Kim ● Jaesung Choi ● JooYong Park ● Chulbum Park ● Se Mi Lee ● Seong Eun Park ● Nan Song ● 5 6 4,7 8 1,4,9 10 Seokang Chung ● Hyuna Sung ● Wonshik Han ● Jong Won Lee ● Sue K. Park ● Mi Kyung Kim ● 4,7 9,11 1,4,9,12 1,4,9 Dong-Young Noh ● Keun-Young Yoo ● Daehee Kang ● Ji-Yeob Choi Received: 7 June 2017 / Revised: 13 October 2017 / Accepted: 4 December 2017 © Macmillan Publishers Limited, part of Springer Nature 2018 Abstract Membrane transporters can be major determinants of the pharmacokinetic profiles of anticancer drugs. The associations between genetic variations of ATP-binding cassette (ABC) and solute carrier (SLC) genes and cancer survival were investigated through a meta-analysis and an association study in the Seoul Breast Cancer Study (SEBCS). Including the SEBCS, the meta-analysis was conducted among 38 studies of genetic variations of transporters on various cancer survivors. 1234567890();,: The population of SEBCS consisted of 1 338 breast cancer patients who had been treated with adjuvant chemotherapy. A total of 7 750 SNPs were selected from 453 ABC and/or SLC genes typed by an Affymetrix 6.0 chip. ABCB1 rs1045642 was associated with poor progression-free survival in a meta-analysis (HR = 1.33, 95% CI: 1.07–1.64). ABCB1, SLC8A1, and SLC12A8 were associated with breast cancer survival in SEBCS (Pgene < 0.05). -

Screening and Identification of Key Biomarkers in Clear Cell Renal Cell Carcinoma Based on Bioinformatics Analysis
bioRxiv preprint doi: https://doi.org/10.1101/2020.12.21.423889; this version posted December 23, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Screening and identification of key biomarkers in clear cell renal cell carcinoma based on bioinformatics analysis Basavaraj Vastrad1, Chanabasayya Vastrad*2 , Iranna Kotturshetti 1. Department of Biochemistry, Basaveshwar College of Pharmacy, Gadag, Karnataka 582103, India. 2. Biostatistics and Bioinformatics, Chanabasava Nilaya, Bharthinagar, Dharwad 580001, Karanataka, India. 3. Department of Ayurveda, Rajiv Gandhi Education Society`s Ayurvedic Medical College, Ron, Karnataka 562209, India. * Chanabasayya Vastrad [email protected] Ph: +919480073398 Chanabasava Nilaya, Bharthinagar, Dharwad 580001 , Karanataka, India bioRxiv preprint doi: https://doi.org/10.1101/2020.12.21.423889; this version posted December 23, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Abstract Clear cell renal cell carcinoma (ccRCC) is one of the most common types of malignancy of the urinary system. The pathogenesis and effective diagnosis of ccRCC have become popular topics for research in the previous decade. In the current study, an integrated bioinformatics analysis was performed to identify core genes associated in ccRCC. An expression dataset (GSE105261) was downloaded from the Gene Expression Omnibus database, and included 26 ccRCC and 9 normal kideny samples. Assessment of the microarray dataset led to the recognition of differentially expressed genes (DEGs), which was subsequently used for pathway and gene ontology (GO) enrichment analysis. -

New Products JULY 2013
R&D Systems Tools for Cell Biology Research™ New Products JULY 2013 GMP-grade Recombinant Proteins Contents R&D Systems now offers GMP-grade cytokines and growth factors for research and further manu- facturing applications where current Good Manufacturing Practices (GMP) are required. GMP-grade Recombinant Proteins 2 proteins are manufactured in our ISO-certified facility in compliance with relevant guidelines1 and are produced with extensive documentation at every stage of development from cell culture to final fill and Quantikine® ELISA Kits 3 formulation. GMP-grade Recombinant Human IL-6 and Recombinant Human TNF-a have recently been added to our line of GMP-grade proteins. Additional GMP-grade proteins will be available in the next several months. For an up-to-date product listing or additional information, please visit our website at Luminex® Screening Assays 4 www.RnDSystems.com/GMP. Luminex® Performance Assays 4-5 Features New GMP Proteins ✓ Extensive documentation at every stage of development ProtEin SOURCE Catalog # SIZE Polyclonal Antibodies 6-7 ✓ Documentation of lot-to-lot consistency and traceability Human IL-6 E. coli 206-GMP-010 10 µg Monoclonal Antibodies 7-8 of materials used 206-GMP-050 50 µg ✓ Rigorous quality control using stringent analytical 206-GMP-01M 1 mg processes Biotinylated Antibodies 8 Human TNF-a/ E. coli 210-GMP-010 10 µg ✓ Proven formulations to ensure consistent reconstitution TNFSF1A and results 210-GMP-050 50 µg Antibody Controls 8 210-GMP-01M 1 mg ELISpot Kits & Development Modules 8 30000 60 Fluorokine® Flow Cytometry Kits 8 25000 17348 ) 2 50 20000 Fluorochrome-labeled Antibodies 9 40 8673 DuoSet® ELISA and DuoSet IC ELISA 15000 Development Systems 10 30 Peak Intensity Peak 10000 20 Cell-Based ELISA Assay Kits 10 5000 17565 (Mean RFU x10 Viability Cell 8780 10 Parameter Assay Kits 10 0 0 6000 9000 12000 15000 18000 21000 24000 -3 -2 -1 0 1 10 10 10 10 10 Apoptosis Detection 10 Mass Charge Ratio Recombinant Human TNF-α GMP (ng/mL) MALDI-TOF Analysis of GMP-grade Recombinant Human TNF-a. -

Small Cell Ovarian Carcinoma: Genomic Stability and Responsiveness to Therapeutics
Gamwell et al. Orphanet Journal of Rare Diseases 2013, 8:33 http://www.ojrd.com/content/8/1/33 RESEARCH Open Access Small cell ovarian carcinoma: genomic stability and responsiveness to therapeutics Lisa F Gamwell1,2, Karen Gambaro3, Maria Merziotis2, Colleen Crane2, Suzanna L Arcand4, Valerie Bourada1,2, Christopher Davis2, Jeremy A Squire6, David G Huntsman7,8, Patricia N Tonin3,4,5 and Barbara C Vanderhyden1,2* Abstract Background: The biology of small cell ovarian carcinoma of the hypercalcemic type (SCCOHT), which is a rare and aggressive form of ovarian cancer, is poorly understood. Tumourigenicity, in vitro growth characteristics, genetic and genomic anomalies, and sensitivity to standard and novel chemotherapeutic treatments were investigated in the unique SCCOHT cell line, BIN-67, to provide further insight in the biology of this rare type of ovarian cancer. Method: The tumourigenic potential of BIN-67 cells was determined and the tumours formed in a xenograft model was compared to human SCCOHT. DNA sequencing, spectral karyotyping and high density SNP array analysis was performed. The sensitivity of the BIN-67 cells to standard chemotherapeutic agents and to vesicular stomatitis virus (VSV) and the JX-594 vaccinia virus was tested. Results: BIN-67 cells were capable of forming spheroids in hanging drop cultures. When xenografted into immunodeficient mice, BIN-67 cells developed into tumours that reflected the hypercalcemia and histology of human SCCOHT, notably intense expression of WT-1 and vimentin, and lack of expression of inhibin. Somatic mutations in TP53 and the most common activating mutations in KRAS and BRAF were not found in BIN-67 cells by DNA sequencing. -

Iron Depletion Reduces Abce1 Transcripts While Inducing The
Preprints (www.preprints.org) | NOT PEER-REVIEWED | Posted: 22 October 2019 doi:10.20944/preprints201910.0252.v1 1 Research Article 2 Iron depletion Reduces Abce1 Transcripts While 3 Inducing the Mitophagy Factors Pink1 and Parkin 4 Jana Key 1,2, Nesli Ece Sen 1, Aleksandar Arsovic 1, Stella Krämer 1, Robert Hülse 1, Suzana 5 Gispert-Sanchez 1 and Georg Auburger 1,* 6 1 Experimental Neurology, Goethe University Medical School, 60590 Frankfurt am Main; 7 2 Faculty of Biosciences, Goethe-University Frankfurt am Main, Germany 8 * Correspondence: [email protected] 9 10 Abstract: Lifespan extension was recently achieved in Caenorhabditis elegans nematodes by 11 mitochondrial stress and mitophagy, triggered via iron depletion. Conversely in man, deficient 12 mitophagy due to Pink1/Parkin mutations triggers iron accumulation in patient brain and limits 13 survival. We now aimed to identify murine fibroblast factors, which adapt their mRNA expression 14 to acute iron manipulation, relate to mitochondrial dysfunction and may influence survival. After 15 iron depletion, expression of the plasma membrane receptor Tfrc with its activator Ireb2, the 16 mitochondrial membrane transporter Abcb10, the heme-release factor Pgrmc1, the heme- 17 degradation enzyme Hmox1, the heme-binding cholesterol metabolizer Cyp46a1, as well as the 18 mitophagy regulators Pink1 and Parkin showed a negative correlation to iron levels. After iron 19 overload, these factors did not change expression. Conversely, a positive correlation of mRNA levels 20 with both conditions of iron availability was observed for the endosomal factors Slc11a2 and Steap2, 21 as well as for the iron-sulfur-cluster (ISC)-containing factors Ppat, Bdh2 and Nthl1. -

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

Protein Identities in Evs Isolated from U87-MG GBM Cells As Determined by NG LC-MS/MS
Protein identities in EVs isolated from U87-MG GBM cells as determined by NG LC-MS/MS. No. Accession Description Σ Coverage Σ# Proteins Σ# Unique Peptides Σ# Peptides Σ# PSMs # AAs MW [kDa] calc. pI 1 A8MS94 Putative golgin subfamily A member 2-like protein 5 OS=Homo sapiens PE=5 SV=2 - [GG2L5_HUMAN] 100 1 1 7 88 110 12,03704523 5,681152344 2 P60660 Myosin light polypeptide 6 OS=Homo sapiens GN=MYL6 PE=1 SV=2 - [MYL6_HUMAN] 100 3 5 17 173 151 16,91913397 4,652832031 3 Q6ZYL4 General transcription factor IIH subunit 5 OS=Homo sapiens GN=GTF2H5 PE=1 SV=1 - [TF2H5_HUMAN] 98,59 1 1 4 13 71 8,048185945 4,652832031 4 P60709 Actin, cytoplasmic 1 OS=Homo sapiens GN=ACTB PE=1 SV=1 - [ACTB_HUMAN] 97,6 5 5 35 917 375 41,70973209 5,478027344 5 P13489 Ribonuclease inhibitor OS=Homo sapiens GN=RNH1 PE=1 SV=2 - [RINI_HUMAN] 96,75 1 12 37 173 461 49,94108966 4,817871094 6 P09382 Galectin-1 OS=Homo sapiens GN=LGALS1 PE=1 SV=2 - [LEG1_HUMAN] 96,3 1 7 14 283 135 14,70620005 5,503417969 7 P60174 Triosephosphate isomerase OS=Homo sapiens GN=TPI1 PE=1 SV=3 - [TPIS_HUMAN] 95,1 3 16 25 375 286 30,77169764 5,922363281 8 P04406 Glyceraldehyde-3-phosphate dehydrogenase OS=Homo sapiens GN=GAPDH PE=1 SV=3 - [G3P_HUMAN] 94,63 2 13 31 509 335 36,03039959 8,455566406 9 Q15185 Prostaglandin E synthase 3 OS=Homo sapiens GN=PTGES3 PE=1 SV=1 - [TEBP_HUMAN] 93,13 1 5 12 74 160 18,68541938 4,538574219 10 P09417 Dihydropteridine reductase OS=Homo sapiens GN=QDPR PE=1 SV=2 - [DHPR_HUMAN] 93,03 1 1 17 69 244 25,77302971 7,371582031 11 P01911 HLA class II histocompatibility antigen, -

ABCG1 (ABC8), the Human Homolog of the Drosophila White Gene, Is a Regulator of Macrophage Cholesterol and Phospholipid Transport
ABCG1 (ABC8), the human homolog of the Drosophila white gene, is a regulator of macrophage cholesterol and phospholipid transport Jochen Klucken*, Christa Bu¨ chler*, Evelyn Orso´ *, Wolfgang E. Kaminski*, Mustafa Porsch-Ozcu¨ ¨ ru¨ mez*, Gerhard Liebisch*, Michael Kapinsky*, Wendy Diederich*, Wolfgang Drobnik*, Michael Dean†, Rando Allikmets‡, and Gerd Schmitz*§ *Institute for Clinical Chemistry and Laboratory Medicine, University of Regensburg, 93042 Regensburg, Germany; †National Cancer Institute, Laboratory of Genomic Diversity, Frederick, MD 21702-1201; and ‡Departments of Ophthalmology and Pathology, Columbia University, Eye Research Addition, New York, NY 10032 Edited by Jan L. Breslow, The Rockefeller University, New York, NY, and approved November 3, 1999 (received for review June 14, 1999) Excessive uptake of atherogenic lipoproteins such as modified low- lesterol transport. Although several effector molecules have been density lipoprotein complexes by vascular macrophages leads to proposed to participate in macrophage cholesterol efflux (6, 9), foam cell formation, a critical step in atherogenesis. Cholesterol efflux including endogenous apolipoprotein E (10) and the cholesteryl mediated by high-density lipoproteins (HDL) constitutes a protective ester transfer protein (11), the detailed molecular mechanisms mechanism against macrophage lipid overloading. The molecular underlying cholesterol export in these cells have not yet been mechanisms underlying this reverse cholesterol transport process are characterized. currently not fully understood. To identify effector proteins that are Recently, mutations of the ATP-binding cassette (ABC) trans- involved in macrophage lipid uptake and release, we searched for porter ABCA1 gene have been causatively linked to familial HDL genes that are regulated during lipid influx and efflux in human deficiency and Tangier disease (12–14). -

A Comprehensive Mapping of the Structure and Gene Organisation in the Sheep MHC Class I Region N
Siva Subramaniam et al. BMC Genomics (2015) 16:810 DOI 10.1186/s12864-015-1992-4 RESEARCH ARTICLE Open Access A comprehensive mapping of the structure and gene organisation in the sheep MHC class I region N. Siva Subramaniam1, EF Morgan1, JD Wetherall1, MJ Stear2,3* and DM Groth1 Abstract Background: The major histocompatibility complex (MHC) is a chromosomal region that regulates immune responsiveness in vertebrates. This region is one of the most important for disease resistance because it has been associated with resistance or susceptibility to a wide variety of diseases and because the MHC often accounts for more of the variance than other loci. Selective breeding for disease resistance is becoming increasingly common in livestock industries, and it is important to determine how this will influence MHC polymorphism and resistance to diseases that are not targeted for selection. However, in sheep the order and sequence of the protein coding genes is controversial. Yet this information is needed to determine precisely how the MHC influences resistance and susceptibility to disease. Methods: CHORI bacterial artificial chromosomes (BACs) known to contain sequences from the sheep MHC class I region were sub-cloned, and the clones partially sequenced. The resulting sequences were analysed and re-assembled to identify gene content and organisation within each BAC. The low resolution MHC class I physical map was then compared to the cattle reference genome, the Chinese Merino sheep MHC map published by Gao, et al. (2010) and the recently available sheep reference genome. Results: Immune related class I genes are clustered into 3 blocks; beta, kappa and a novel block not previously identified in other organisms. -

Human Pluripotent Stem Cells As a Model of Trophoblast Differentiation in Both Normal Development and Disease
Human pluripotent stem cells as a model of trophoblast differentiation in both normal development and disease Mariko Horiia,b,1, Yingchun Lia,b,1, Anna K. Wakelanda,b,1, Donald P. Pizzoa, Katharine K. Nelsona,b, Karen Sabatinib,c, Louise Chang Laurentb,c, Ying Liud,e,f, and Mana M. Parasta,b,2 aDepartment of Pathology, University of California, San Diego, La Jolla, CA 92093; bSanford Consortium for Regenerative Medicine, University of California, San Diego, La Jolla, CA 92093; cDepartment of Reproductive Medicine, University of California, San Diego, La Jolla, CA 92093; dDepartment of Neurosurgery, Center for Stem Cell and Regenerative Medicine, University of Texas Health Sciences Center, Houston, TX 77030; eThe Senator Lloyd and B. A. Bentsen Center for Stroke Research, University of Texas Health Sciences Center, Houston, TX 77030; and fThe Brown Foundation Institute of Molecular Medicine for the Prevention of Human Diseases, University of Texas Health Sciences Center, Houston, TX 77030 Edited by R. Michael Roberts, University of Missouri–Columbia, Columbia, MO, and approved May 25, 2016 (received for review March 24, 2016) Trophoblast is the primary epithelial cell type in the placenta, a Elf5 (Ets domain transcription factor) and Eomes (Eomeso- transient organ required for proper fetal growth and develop- dermin), also have been shown to be required for maintenance of ment. Different trophoblast subtypes are responsible for gas/nutrient the TSC fate in the mouse (8, 9). exchange (syncytiotrophoblasts, STBs) and invasion and maternal Significantly less is known about TE specification and the TSC vascular remodeling (extravillous trophoblasts, EVTs). Studies of niche in the human embryo (10, 11). -
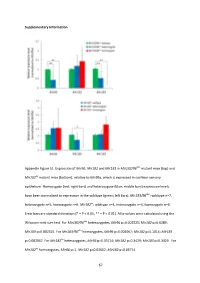
And Mir183 in Mir183/96 Dko Mutant Mice (Top) And
Supplementary Information Appendix Figure S1. Expression of Mir96 , Mir182 and Mir183 in Mir183/96 dko mutant mice (top) and Mir182 ko mutant mice (bottom), relative to Mir99a , which is expressed in cochlear sensory epithelium. Homozygote (red; right bars) and heterozygote (blue; middle bars) expression levels have been normalised to expression in the wildtype (green; left bars). Mir183/96 dko : wildtype n=7, heterozygote n=5, homozygote n=6. Mir182 ko : wildtype n=4, heterozygote n=4, homozygote n=4. Error bars are standard deviation (* = P < 0.05, ** = P < 0.01). All p-values were calculated using the Wilcoxon rank sum test. For Mir183/96 dko heterozygotes, Mir96 p=0.002525; Mir182 p=0.6389; Mir183 p=0.002525. For Mir183/96 dko homozygotes, Mir96 p=0.002067; Mir182 p=0.1014; Mir183 p=0.002067. For Mir182 ko heterozygotes, Mir96 p=0.05714; Mir182 p=0.3429; Mir183 p=0.3429. For Mir182 ko homozygotes, Mir96 p=1; Mir182 p=0.02652; Mir183 p=0.05714. 67 68 Appendix Figure S2. Individual ABR thresholds of wildtype, heterozygous and homozygous Mir183/96 dko mice at all ages tested. Number of mice of each genotype tested at each age is shown on the threshold plot. 69 70 Appendix Figure S3. Individual ABR thresholds of wildtype, heterozygous and homozygous Mir182 ko mice at all ages tested. Number of mice of each genotype tested at each age is shown on the threshold plot. 71 Appendix Figure S4. Mean ABR waveforms at 12kHz, shown at 20dB (top) and 50dB (bottom) above threshold (sensation level, SL) ± standard deviation, at four weeks old.