Chapter 2, Probability Theory
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

The Probability Set-Up.Pdf
CHAPTER 2 The probability set-up 2.1. Basic theory of probability We will have a sample space, denoted by S (sometimes Ω) that consists of all possible outcomes. For example, if we roll two dice, the sample space would be all possible pairs made up of the numbers one through six. An event is a subset of S. Another example is to toss a coin 2 times, and let S = fHH;HT;TH;TT g; or to let S be the possible orders in which 5 horses nish in a horse race; or S the possible prices of some stock at closing time today; or S = [0; 1); the age at which someone dies; or S the points in a circle, the possible places a dart can hit. We should also keep in mind that the same setting can be described using dierent sample set. For example, in two solutions in Example 1.30 we used two dierent sample sets. 2.1.1. Sets. We start by describing elementary operations on sets. By a set we mean a collection of distinct objects called elements of the set, and we consider a set as an object in its own right. Set operations Suppose S is a set. We say that A ⊂ S, that is, A is a subset of S if every element in A is contained in S; A [ B is the union of sets A ⊂ S and B ⊂ S and denotes the points of S that are in A or B or both; A \ B is the intersection of sets A ⊂ S and B ⊂ S and is the set of points that are in both A and B; ; denotes the empty set; Ac is the complement of A, that is, the points in S that are not in A. -

3.2.3 Binomial Distribution
3.2.3 Binomial Distribution The binomial distribution is based on the idea of a Bernoulli trial. A Bernoulli trail is an experiment with two, and only two, possible outcomes. A random variable X has a Bernoulli(p) distribution if 8 > <1 with probability p X = > :0 with probability 1 − p, where 0 ≤ p ≤ 1. The value X = 1 is often termed a “success” and X = 0 is termed a “failure”. The mean and variance of a Bernoulli(p) random variable are easily seen to be EX = (1)(p) + (0)(1 − p) = p and VarX = (1 − p)2p + (0 − p)2(1 − p) = p(1 − p). In a sequence of n identical, independent Bernoulli trials, each with success probability p, define the random variables X1,...,Xn by 8 > <1 with probability p X = i > :0 with probability 1 − p. The random variable Xn Y = Xi i=1 has the binomial distribution and it the number of sucesses among n independent trials. The probability mass function of Y is µ ¶ ¡ ¢ n ¡ ¢ P Y = y = py 1 − p n−y. y For this distribution, t n EX = np, Var(X) = np(1 − p),MX (t) = [pe + (1 − p)] . 1 Theorem 3.2.2 (Binomial theorem) For any real numbers x and y and integer n ≥ 0, µ ¶ Xn n (x + y)n = xiyn−i. i i=0 If we take x = p and y = 1 − p, we get µ ¶ Xn n 1 = (p + (1 − p))n = pi(1 − p)n−i. i i=0 Example 3.2.2 (Dice probabilities) Suppose we are interested in finding the probability of obtaining at least one 6 in four rolls of a fair die. -

Determinism, Indeterminism and the Statistical Postulate
Tipicality, Explanation, The Statistical Postulate, and GRW Valia Allori Northern Illinois University [email protected] www.valiaallori.com Rutgers, October 24-26, 2019 1 Overview Context: Explanation of the macroscopic laws of thermodynamics in the Boltzmannian approach Among the ingredients: the statistical postulate (connected with the notion of probability) In this presentation: typicality Def: P is a typical property of X-type object/phenomena iff the vast majority of objects/phenomena of type X possesses P Part I: typicality is sufficient to explain macroscopic laws – explanatory schema based on typicality: you explain P if you explain that P is typical Part II: the statistical postulate as derivable from the dynamics Part III: if so, no preference for indeterministic theories in the quantum domain 2 Summary of Boltzmann- 1 Aim: ‘derive’ macroscopic laws of thermodynamics in terms of the microscopic Newtonian dynamics Problems: Technical ones: There are to many particles to do exact calculations Solution: statistical methods - if 푁 is big enough, one can use suitable mathematical technique to obtain information of macro systems even without having the exact solution Conceptual ones: Macro processes are irreversible, while micro processes are not Boltzmann 3 Summary of Boltzmann- 2 Postulate 1: the microscopic dynamics microstate 푋 = 푟1, … . 푟푁, 푣1, … . 푣푁 in phase space Partition of phase space into Macrostates Macrostate 푀(푋): set of macroscopically indistinguishable microstates Macroscopic view Many 푋 for a given 푀 given 푀, which 푋 is unknown Macro Properties (e.g. temperature): they slowly vary on the Macro scale There is a particular Macrostate which is incredibly bigger than the others There are many ways more to have, for instance, uniform temperature than not equilibrium (Macro)state 4 Summary of Boltzmann- 3 Entropy: (def) proportional to the size of the Macrostate in phase space (i.e. -

Topic 1: Basic Probability Definition of Sets
Topic 1: Basic probability ² Review of sets ² Sample space and probability measure ² Probability axioms ² Basic probability laws ² Conditional probability ² Bayes' rules ² Independence ² Counting ES150 { Harvard SEAS 1 De¯nition of Sets ² A set S is a collection of objects, which are the elements of the set. { The number of elements in a set S can be ¯nite S = fx1; x2; : : : ; xng or in¯nite but countable S = fx1; x2; : : :g or uncountably in¯nite. { S can also contain elements with a certain property S = fx j x satis¯es P g ² S is a subset of T if every element of S also belongs to T S ½ T or T S If S ½ T and T ½ S then S = T . ² The universal set is the set of all objects within a context. We then consider all sets S ½ . ES150 { Harvard SEAS 2 Set Operations and Properties ² Set operations { Complement Ac: set of all elements not in A { Union A \ B: set of all elements in A or B or both { Intersection A [ B: set of all elements common in both A and B { Di®erence A ¡ B: set containing all elements in A but not in B. ² Properties of set operations { Commutative: A \ B = B \ A and A [ B = B [ A. (But A ¡ B 6= B ¡ A). { Associative: (A \ B) \ C = A \ (B \ C) = A \ B \ C. (also for [) { Distributive: A \ (B [ C) = (A \ B) [ (A \ C) A [ (B \ C) = (A [ B) \ (A [ C) { DeMorgan's laws: (A \ B)c = Ac [ Bc (A [ B)c = Ac \ Bc ES150 { Harvard SEAS 3 Elements of probability theory A probabilistic model includes ² The sample space of an experiment { set of all possible outcomes { ¯nite or in¯nite { discrete or continuous { possibly multi-dimensional ² An event A is a set of outcomes { a subset of the sample space, A ½ . -

5 Stochastic Processes
5 Stochastic Processes Contents 5.1. The Bernoulli Process ...................p.3 5.2. The Poisson Process .................. p.15 1 2 Stochastic Processes Chap. 5 A stochastic process is a mathematical model of a probabilistic experiment that evolves in time and generates a sequence of numerical values. For example, a stochastic process can be used to model: (a) the sequence of daily prices of a stock; (b) the sequence of scores in a football game; (c) the sequence of failure times of a machine; (d) the sequence of hourly traffic loads at a node of a communication network; (e) the sequence of radar measurements of the position of an airplane. Each numerical value in the sequence is modeled by a random variable, so a stochastic process is simply a (finite or infinite) sequence of random variables and does not represent a major conceptual departure from our basic framework. We are still dealing with a single basic experiment that involves outcomes gov- erned by a probability law, and random variables that inherit their probabilistic † properties from that law. However, stochastic processes involve some change in emphasis over our earlier models. In particular: (a) We tend to focus on the dependencies in the sequence of values generated by the process. For example, how do future prices of a stock depend on past values? (b) We are often interested in long-term averages,involving the entire se- quence of generated values. For example, what is the fraction of time that a machine is idle? (c) We sometimes wish to characterize the likelihood or frequency of certain boundary events. -

1 Normal Distribution
1 Normal Distribution. 1.1 Introduction A Bernoulli trial is simple random experiment that ends in success or failure. A Bernoulli trial can be used to make a new random experiment by repeating the Bernoulli trial and recording the number of successes. Now repeating a Bernoulli trial a large number of times has an irritating side e¤ect. Suppose we take a tossed die and look for a 3 to come up, but we do this 6000 times. 1 This is a Bernoulli trial with a probability of success of 6 repeated 6000 times. What is the probability that we will see exactly 1000 success? This is de…nitely the most likely possible outcome, 1000 successes out of 6000 tries. But it is still very unlikely that an particular experiment like this will turn out so exactly. In fact, if 6000 tosses did produce exactly 1000 successes, that would be rather suspicious. The probability of exactly 1000 successes in 6000 tries almost does not need to be calculated. whatever the probability of this, it will be very close to zero. It is probably too small a probability to be of any practical use. It turns out that it is not all that bad, 0:014. Still this is small enough that it means that have even a chance of seeing it actually happen, we would need to repeat the full experiment as many as 100 times. All told, 600,000 tosses of a die. When we repeat a Bernoulli trial a large number of times, it is unlikely that we will be interested in a speci…c number of successes, and much more likely that we will be interested in the event that the number of successes lies within a range of possibilities. -

Bernoulli Random Forests: Closing the Gap Between Theoretical Consistency and Empirical Soundness
Proceedings of the Twenty-Fifth International Joint Conference on Artificial Intelligence (IJCAI-16) Bernoulli Random Forests: Closing the Gap between Theoretical Consistency and Empirical Soundness , , , ? Yisen Wang† ‡, Qingtao Tang† ‡, Shu-Tao Xia† ‡, Jia Wu , Xingquan Zhu ⇧ † Dept. of Computer Science and Technology, Tsinghua University, China ‡ Graduate School at Shenzhen, Tsinghua University, China ? Quantum Computation & Intelligent Systems Centre, University of Technology Sydney, Australia ⇧ Dept. of Computer & Electrical Engineering and Computer Science, Florida Atlantic University, USA wangys14, tqt15 @mails.tsinghua.edu.cn; [email protected]; [email protected]; [email protected] { } Traditional Bernoulli Trial Controlled Abstract Tree Node Splitting Tree Node Splitting Random forests are one type of the most effective ensemble learning methods. In spite of their sound Random Attribute Bagging Bernoulli Trial Controlled empirical performance, the study on their theoreti- Attribute Bagging cal properties has been left far behind. Recently, several random forests variants with nice theoreti- Random Structure/Estimation cal basis have been proposed, but they all suffer Random Bootstrap Sampling Points Splitting from poor empirical performance. In this paper, we (a) Breiman RF (b) BRF propose a Bernoulli random forests model (BRF), which intends to close the gap between the theoreti- Figure 1: Comparisons between Breiman RF (left panel) vs. cal consistency and the empirical soundness of ran- the proposed BRF (right panel). The tree node splitting of dom forests classification. Compared to Breiman’s Breiman RF is deterministic, so the final trees are highly data- original random forests, BRF makes two simplifi- dependent. Instead, BRF employs two Bernoulli distributions cations in tree construction by using two indepen- to control the tree construction. -

Probability Theory
Probability Theory Course Notes — Harvard University — 2011 C. McMullen March 29, 2021 Contents I TheSampleSpace ........................ 2 II Elements of Combinatorial Analysis . 5 III RandomWalks .......................... 15 IV CombinationsofEvents . 24 V ConditionalProbability . 29 VI The Binomial and Poisson Distributions . 37 VII NormalApproximation. 44 VIII Unlimited Sequences of Bernoulli Trials . 55 IX Random Variables and Expectation . 60 X LawofLargeNumbers...................... 68 XI Integral–Valued Variables. Generating Functions . 70 XIV RandomWalkandRuinProblems . 70 I The Exponential and the Uniform Density . 75 II Special Densities. Randomization . 94 These course notes accompany Feller, An Introduction to Probability Theory and Its Applications, Wiley, 1950. I The Sample Space Some sources and uses of randomness, and philosophical conundrums. 1. Flipped coin. 2. The interrupted game of chance (Fermat). 3. The last roll of the game in backgammon (splitting the stakes at Monte Carlo). 4. Large numbers: elections, gases, lottery. 5. True randomness? Quantum theory. 6. Randomness as a model (in reality only one thing happens). Paradox: what if a coin keeps coming up heads? 7. Statistics: testing a drug. When is an event good evidence rather than a random artifact? 8. Significance: among 1000 coins, if one comes up heads 10 times in a row, is it likely to be a 2-headed coin? Applications to economics, investment and hiring. 9. Randomness as a tool: graph theory; scheduling; internet routing. We begin with some previews. Coin flips. What are the chances of 10 heads in a row? The probability is 1/1024, less than 0.1%. Implicit assumptions: no biases and independence. 10 What are the chance of heads 5 out of ten times? ( 5 = 252, so 252/1024 = 25%). -
Discrete Distributions: Empirical, Bernoulli, Binomial, Poisson
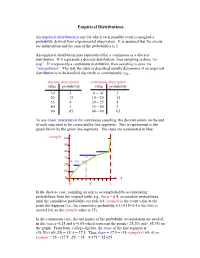
Empirical Distributions An empirical distribution is one for which each possible event is assigned a probability derived from experimental observation. It is assumed that the events are independent and the sum of the probabilities is 1. An empirical distribution may represent either a continuous or a discrete distribution. If it represents a discrete distribution, then sampling is done “on step”. If it represents a continuous distribution, then sampling is done via “interpolation”. The way the table is described usually determines if an empirical distribution is to be handled discretely or continuously; e.g., discrete description continuous description value probability value probability 10 .1 0 – 10- .1 20 .15 10 – 20- .15 35 .4 20 – 35- .4 40 .3 35 – 40- .3 60 .05 40 – 60- .05 To use linear interpolation for continuous sampling, the discrete points on the end of each step need to be connected by line segments. This is represented in the graph below by the green line segments. The steps are represented in blue: rsample 60 50 40 30 20 10 0 x 0 .5 1 In the discrete case, sampling on step is accomplished by accumulating probabilities from the original table; e.g., for x = 0.4, accumulate probabilities until the cumulative probability exceeds 0.4; rsample is the event value at the point this happens (i.e., the cumulative probability 0.1+0.15+0.4 is the first to exceed 0.4, so the rsample value is 35). In the continuous case, the end points of the probability accumulation are needed, in this case x=0.25 and x=0.65 which represent the points (.25,20) and (.65,35) on the graph. -

Randomization-Based Inference for Bernoulli Trial Experiments And
Article Statistical Methods in Medical Research 2019, Vol. 28(5) 1378–1398 ! The Author(s) 2018 Randomization-based inference for Article reuse guidelines: sagepub.com/journals-permissions Bernoulli trial experiments and DOI: 10.1177/0962280218756689 implications for observational studies journals.sagepub.com/home/smm Zach Branson and Marie-Abe`le Bind Abstract We present a randomization-based inferential framework for experiments characterized by a strongly ignorable assignment mechanism where units have independent probabilities of receiving treatment. Previous works on randomization tests often assume these probabilities are equal within blocks of units. We consider the general case where they differ across units and show how to perform randomization tests and obtain point estimates and confidence intervals. Furthermore, we develop rejection-sampling and importance-sampling approaches for conducting randomization-based inference conditional on any statistic of interest, such as the number of treated units or forms of covariate balance. We establish that our randomization tests are valid tests, and through simulation we demonstrate how the rejection-sampling and importance-sampling approaches can yield powerful randomization tests and thus precise inference. Our work also has implications for observational studies, which commonly assume a strongly ignorable assignment mechanism. Most methodologies for observational studies make additional modeling or asymptotic assumptions, while our framework only assumes the strongly ignorable assignment -

Non-Archimedean Probability (NAP) Theory
Non-Archimedean Probability Vieri Benci∗ Leon Horsten† Sylvia Wenmackers‡ October 29, 2018 Abstract We propose an alternative approach to probability theory closely re- lated to the framework of numerosity theory: non-Archimedean prob- ability (NAP). In our approach, unlike in classical probability theory, all subsets of an infinite sample space are measurable and zero- and unit-probability events pose no particular epistemological problems. We use a non-Archimedean field as the range of the probability func- tion. As a result, the property of countable additivity in Kolmogorov’s axiomatization of probability is replaced by a different type of infinite additivity. Mathematics subject classification. 60A05, 03H05 Keywords. Probability, Axioms of Kolmogorov, Nonstandard models, De Finetti lottery, Non-Archimedean fields. Contents 1 Introduction 2 1.1 Somenotation .......................... 3 2 Kolmogorov’s probability theory 4 2.1 Kolmogorov’saxioms.. .. .. .. .. .. .. 4 2.2 Problems with Kolmogorov’s axioms . 6 3 Non-Archimedean Probability 7 arXiv:1106.1524v1 [math.PR] 8 Jun 2011 3.1 The axioms of Non-Archimedean Probability . 7 3.2 Analysisofthefourthaxiom . 10 3.3 First consequences of the axioms . 11 3.4 Infinitesums ........................... 13 ∗Dipartimento di Matematica Applicata, Universit`adegli Studi di Pisa, Via F. Buonar- roti 1/c, Pisa, ITALY and Department of Mathematics, College of Science, King Saud University, Riyadh, 11451, SAUDI ARABIA. e-mail: [email protected] †Department of Philosophy, University of Bristol, 43 Woodland Rd, BS81UU Bristol, UNITED KINGDOM. e-mail: [email protected] ‡Faculty of Philosophy, University of Groningen, Oude Boteringestraat 52, 9712 GL Groningen, THE NETHERLANDS. e-mail: [email protected] 1 4 NAP-spaces and Λ-limits 14 4.1 Fineideals............................ -

Some Basic Probabilistic Processes
CHAPTER FOUR some basic probabilistic processes This chapter presents a few simple probabilistic processes and develops family relationships among the PMF's and PDF's associated with these processes. Although we shall encounter many of the most common PMF's and PDF's here, it is not our purpose to develop a general catalogue. A listing of the most frequently occurring PRfF's and PDF's and some of their properties appears as an appendix at the end of this book. 4-1 The 8er~ouil1Process A single Bernoulli trial generates an experimental value of discrete random variable x, described by the PMF to expand pkT(z) in a power series and then note the coefficient of zko in this expansion, recalling that any z transform may be written in the form pkT(~)= pk(0) + zpk(1) + z2pk(2) + ' ' This leads to the result known as the binomial PMF, 1-P xo-0 OlPll xo= 1 otherwise where the notation is the common Random variable x, as described above, is known as a Bernoulli random variable, and we note that its PMF has the z transform discussed in Sec. 1-9. Another way to derive the binomial PMF would be to work in a sequential sample space for an experiment which consists of n independ- The sample space for each Bernoulli trial is of the form ent Bernoulli trials, We have used the notation t success (;'') (;'') = Ifailure1 on the nth trial Either by use of the transform or by direct calculation we find We refer to the outcome of a Bernoulli trial as a success when the ex- Each sample point which represents an outcome of exactly ko suc- perimental value of x is unity and as a failure when the experimental cesses in the n trials would have a probability assignment equal to value of x is zero.