Exon Shuffling Played a Decisive Role in the Evolution of the Genetic Toolkit for the Multicellular Body Plan of Metazoa
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Predicted Glycosyltransferases Promote Development and Prevent Spurious Cell Clumping in the Choanoflagellate S
RESEARCH ADVANCE Predicted glycosyltransferases promote development and prevent spurious cell clumping in the choanoflagellate S. rosetta Laura A Wetzel1,2, Tera C Levin1,2, Ryan E Hulett1,2, Daniel Chan1,2, Grant A King1,2, Reef Aldayafleh1,2, David S Booth1,2, Monika Abedin Sigg1,2, Nicole King1,2* 1Department of Molecular and Cell Biology, University of California, Berkeley, Berkeley, United States; 2Howard Hughes Medical Institute, University of California, Berkeley, Berkeley, United States Abstract In a previous study we established forward genetics in the choanoflagellate Salpingoeca rosetta and found that a C-type lectin gene is required for rosette development (Levin et al., 2014). Here we report on critical improvements to genetic screens in S. rosetta while also investigating the genetic basis for rosette defect mutants in which single cells fail to develop into orderly rosettes and instead aggregate promiscuously into amorphous clumps of cells. Two of the mutants, Jumble and Couscous, mapped to lesions in genes encoding two different predicted glycosyltransferases and displayed aberrant glycosylation patterns in the basal extracellular matrix (ECM). In animals, glycosyltransferases sculpt the polysaccharide-rich ECM, regulate integrin and cadherin activity, and, when disrupted, contribute to tumorigenesis. The finding that predicted glycosyltransferases promote proper rosette development and prevent cell aggregation in S. rosetta suggests a pre-metazoan role for glycosyltransferases in regulating development and preventing abnormal tumor-like multicellularity. *For correspondence: DOI: https://doi.org/10.7554/eLife.41482.001 [email protected] Competing interests: The authors declare that no competing interests exist. Introduction Funding: See page 23 The transition to multicellularity was essential for the evolution of animals from their single celled Received: 05 September 2018 ancestors (Szathma´ry and Smith, 1995). -

The Architecture of Cell Differentiation in Choanoflagellates And
bioRxiv preprint doi: https://doi.org/10.1101/452185; this version posted October 29, 2018. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. 1 The architecture of cell differentiation in choanoflagellates 2 and sponge choanocytes 3 Davis Laundon1,2,6, Ben Larson3, Kent McDonald3, Nicole King3,4 and Pawel 4 Burkhardt1,5* 5 1 Marine Biological Association of the United Kingdom, The Laboratory, Citadel Hill, 6 Plymouth, PL1 2PB, United Kingdom 7 2 Plymouth University, Drake Circus, Plymouth, PL4 8AA, United Kingdom 8 3 Department of Molecular and Cell Biology, University of California, Berkeley, USA 9 4 Howard Hughes Medical Institute 10 5 Sars International Centre for Molecular Marine Biology, University of Bergen, 11 Thormohlensgate 55, 5020 Bergen, Norway 12 6 Current Affiliation: University of East Anglia, Norwich, NR4 7TJ, United Kingdom 13 14 *Correspondence [email protected] 15 16 17 18 19 20 21 22 23 24 25 bioRxiv preprint doi: https://doi.org/10.1101/452185; this version posted October 29, 2018. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. 26 SUMMARY 27 Collar cells are ancient animal cell types which are conserved across the animal 28 kingdom [1] and their closest relatives, the choanoflagellates [2]. -

Mechanisms and Impact of Post-Transcriptional Exon Shuffling
Mechanisms and impact of Post-Transcriptional Exon Shuffling (PTES) Ginikachukwu Osagie Izuogu Doctor of Philosophy Institute of Genetic Medicine Newcastle University September 2016 i ABSTRACT Most eukaryotic genes undergo splicing to remove introns and join exons sequentially to produce protein-coding or non-coding transcripts. Post-transcriptional Exon Shuffling (PTES) describes a new class of RNA molecules, characterized by exon order different from the underlying genomic context. PTES can result in linear and circular RNA (circRNA) molecules and enhance the complexity of transcriptomes. Prior to my studies, I developed PTESFinder, a computational tool for PTES identification from high-throughput RNAseq data. As various sources of artefacts (including pseudogenes, template-switching and others) can confound PTES identification, I first assessed the effectiveness of filters within PTESFinder devised to systematically exclude artefacts. When compared to 4 published methods, PTESFinder achieves the highest specificity (~0.99) and comparable sensitivity (~0.85). To define sub-cellular distribution of PTES, I performed in silico analyses of data from various cellular compartments and revealed diverse populations of PTES in nuclei and enrichment in cytosol of various cell lines. Identification of PTES from chromatin-associated RNAseq data and an assessment of co-transcriptional splicing, established that PTES may occur during transcription. To assess if PTES contribute to the proteome, I analyzed sucrose-gradient fractionated data from HEK293, treated with arsenite to induce translational arrest and dislodge ribosomes. My results showed no effect of arsenite treatment on ribosome occupancy within PTES transcripts, indicating that these transcripts are not generally bound by polysomes and do not contribute to the proteome. -

1 Lessons from Simple Marine Models on the Bacterial Regulation
bioRxiv preprint doi: https://doi.org/10.1101/211797; this version posted October 31, 2017. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. Lessons from simple marine models on the bacterial regulation of eukaryotic development Arielle Woznicaa and Nicole Kinga,1 a Howard Hughes Medical Institute and Department of Molecular and Cell Biology, University of California, Berkeley, California 94720, United States 1 Corresponding author, [email protected]. 1 bioRxiv preprint doi: https://doi.org/10.1101/211797; this version posted October 31, 2017. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. 1 Highlights 2 - Cues from environmental bacteria influence the development of many marine 3 eukaryotes 4 5 - The molecular cues produced by environmental bacteria are structurally diverse 6 7 - Eukaryotes can respond to many different environmental bacteria 8 9 - Some environmental bacteria act as “information hubs” for diverse eukaryotes 10 11 - Experimentally tractable systems, like the choanoflagellate S. rosetta, promise to 12 reveal molecular mechanisms underlying these interactions 13 14 Abstract 15 Molecular cues from environmental bacteria influence important developmental 16 decisions in diverse marine eukaryotes. Yet, relatively little is understood about the 17 mechanisms underlying these interactions, in part because marine ecosystems are 18 dynamic and complex. -

A Six-Gene Phylogeny Provides New Insights Into Choanoflagellate Evolution Martin Carr, Daniel J
A six-gene phylogeny provides new insights into choanoflagellate evolution Martin Carr, Daniel J. Richter, Parinaz Fozouni, Timothy J. Smith, Alexandra Jeuck, Barry S.C. Leadbeater, Frank Nitsche To cite this version: Martin Carr, Daniel J. Richter, Parinaz Fozouni, Timothy J. Smith, Alexandra Jeuck, et al.. A six- gene phylogeny provides new insights into choanoflagellate evolution. Molecular Phylogenetics and Evolution, Elsevier, 2017, 107, pp.166 - 178. 10.1016/j.ympev.2016.10.011. hal-01393449 HAL Id: hal-01393449 https://hal.archives-ouvertes.fr/hal-01393449 Submitted on 7 Nov 2016 HAL is a multi-disciplinary open access L’archive ouverte pluridisciplinaire HAL, est archive for the deposit and dissemination of sci- destinée au dépôt et à la diffusion de documents entific research documents, whether they are pub- scientifiques de niveau recherche, publiés ou non, lished or not. The documents may come from émanant des établissements d’enseignement et de teaching and research institutions in France or recherche français ou étrangers, des laboratoires abroad, or from public or private research centers. publics ou privés. Distributed under a Creative Commons Attribution| 4.0 International License Molecular Phylogenetics and Evolution 107 (2017) 166–178 Contents lists available at ScienceDirect Molecular Phylogenetics and Evolution journal homepage: www.elsevier.com/locate/ympev A six-gene phylogeny provides new insights into choanoflagellate evolution ⇑ Martin Carr a, ,1, Daniel J. Richter b,1,2, Parinaz Fozouni b,3, Timothy J. Smith a, Alexandra Jeuck c, Barry S.C. Leadbeater d, Frank Nitsche c a School of Applied Sciences, University of Huddersfield, Huddersfield HD1 3DH, UK b Department of Molecular and Cell Biology, University of California, Berkeley, CA 94720-3200, USA c University of Cologne, Biocentre, General Ecology, Zuelpicher Str. -

"Evolutionary Emergence of Genes Through Retrotransposition"
Evolutionary Emergence of Advanced article Genes Through Article Contents . Introduction Retrotransposition . Gene Alteration Following Retrotransposon Insertion . Retrotransposon Recruitment by Host Genome . Retrotransposon-mediated Gene Duplication Richard Cordaux, University of Poitiers, Poitiers, France . Conclusion Mark A Batzer, Department of Biological Sciences, Louisiana State University, Baton Rouge, doi: 10.1002/9780470015902.a0020783 Louisiana, USA Variation in the number of genes among species indicates that new genes are continuously generated over evolutionary times. Evidence is accumulating that transposable elements, including retrotransposons (which account for about 90% of all transposable elements inserted in primate genomes), are potent mediators of new gene origination. Retrotransposons have fostered genetic innovation during human and primate evolution through: (i) alteration of structure and/or expression of pre-existing genes following their insertion, (ii) recruitment (or domestication) of their coding sequence by the host genome and (iii) their ability to mediate gene duplication via ectopic recombination, sequence transduction and gene retrotransposition. Introduction genes, respectively, and de novo origination from previ- ously noncoding genomic sequence. Genome sequencing Variation in the number of genes among species indicates projects have also highlighted that new gene structures can that new genes are continuously generated over evolution- arise as a result of the activity of transposable elements ary times. Although the emergence of new genes and (TEs), which are mobile genetic units or ‘jumping genes’ functions is of central importance to the evolution of that have been bombarding the genomes of most species species, studies on the formation of genetic innovations during evolution. For example, there are over three million have only recently become possible. -
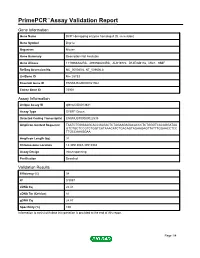
Primepcr™Assay Validation Report
PrimePCR™Assay Validation Report Gene Information Gene Name DCP1 decapping enzyme homolog A (S. cerevisiae) Gene Symbol Dcp1a Organism Mouse Gene Summary Description Not Available Gene Aliases 1110066A22Rik, 4930568L04Rik, AU019772, D14Ertd817e, Mitc1, SMIF RefSeq Accession No. NC_000080.6, NT_039606.8 UniGene ID Mm.28733 Ensembl Gene ID ENSMUSG00000021962 Entrez Gene ID 75901 Assay Information Unique Assay ID qMmuCID0013841 Assay Type SYBR® Green Detected Coding Transcript(s) ENSMUST00000022535 Amplicon Context Sequence TAATCTGGGAAGCACCGAGACTCTAGAAGAGACACCCTCTGGGTCACAGGATAA GTCTGCTCCGTCTGGTCATAAACATCTGACAGTAGAAGAGTTATTTGGAACCTCC TTGCCAAAGGAA Amplicon Length (bp) 91 Chromosome Location 14:30513043-30518984 Assay Design Intron-spanning Purification Desalted Validation Results Efficiency (%) 98 R2 0.9997 cDNA Cq 22.41 cDNA Tm (Celsius) 81 gDNA Cq 24.87 Specificity (%) 100 Information to assist with data interpretation is provided at the end of this report. Page 1/4 PrimePCR™Assay Validation Report Dcp1a, Mouse Amplification Plot Amplification of cDNA generated from 25 ng of universal reference RNA Melt Peak Melt curve analysis of above amplification Standard Curve Standard curve generated using 20 million copies of template diluted 10-fold to 20 copies Page 2/4 PrimePCR™Assay Validation Report Products used to generate validation data Real-Time PCR Instrument CFX384 Real-Time PCR Detection System Reverse Transcription Reagent iScript™ Advanced cDNA Synthesis Kit for RT-qPCR Real-Time PCR Supermix SsoAdvanced™ SYBR® Green Supermix Experimental Sample qPCR Mouse Reference Total RNA Data Interpretation Unique Assay ID This is a unique identifier that can be used to identify the assay in the literature and online. Detected Coding Transcript(s) This is a list of the Ensembl transcript ID(s) that this assay will detect. -

Exon Amplification: a Strategy to Isolate Mammalian Genes Based on RNA Splicing (Gene Cloning/Polymerase Chain Reaction) ALAN J
Proc. Natl. Acad. Sci. USA Vol. 88, pp. 4005-4009, May 1991 Genetics Exon amplification: A strategy to isolate mammalian genes based on RNA splicing (gene cloning/polymerase chain reaction) ALAN J. BUCKLER*, DAVID D. CHANG, SHARON L. GRAw, J. DAVID BROOK, DANIEL A. HABER, PHILLIP A. SHARP, AND DAVID E. HOUSMAN Center for Cancer Research, Department of Biology, Massachusetts Institute of Technology, Cambridge, MA 02139 Contributed by Phillip A. Sharp, January 25, 1991 ABSTRACT We have developed a method, exon amplifi- (7, 8). Thus, this method may be generally applicable for the cation, for fast and efficient isolation of coding sequences from selection of exon sequences from any gene. The method is complex mammalian genomic DNA. This method is based on also both rapid and easily adapted to large scale experiments. the selection of RNA sequences, exons, which are flanked by A series of cloned genomic DNA fragments can be screened functional 5' and 3' splice sites. Fragments of cloned genomic within 1-2 weeks. The sensitivity of this method is high. DNA are inserted into an intron, which is flanked by 5' and 3' Genomic DNA segments of 20 kilobases (kb) or more can be splice sites of the human immunodeficiency virus 1 tat gene successfully screened in a single transfection by using a set contained within the plasmid pSPL1. COS-7 cells are trans- of pooled subclones. This method thus allows the rapid fected with these constructs, and the resulting RNA transcripts identification of exons in mammalian genomic DNA and are processed in vivo. Splice sites of exons contained within the should facilitate the isolation of a wide spectrum of genes of inserted genomic fragment are paired with splice sites of the significance in physiology and development. -

Mrna Turnover Philip Mitchell* and David Tollervey†
320 mRNA turnover Philip Mitchell* and David Tollervey† Nuclear RNA-binding proteins can record pre-mRNA are cotransported to the cytoplasm with the mRNP. These processing events in the structure of messenger proteins may preserve a record of the nuclear history of the ribonucleoprotein particles (mRNPs). During initial rounds of pre-mRNA in the cytoplasmic mRNP structure. This infor- translation, the mature mRNP structure is established and is mation can strongly influence the cytoplasmic fate of the monitored by mRNA surveillance systems. Competition for the mRNA and is used by mRNA surveillance systems that act cap structure links translation and subsequent mRNA as a checkpoint of mRNP integrity, particularly in the identi- degradation, which may also involve multiple deadenylases. fication of premature translation termination codons (PTCs). Addresses Cotransport of nuclear mRNA-binding proteins with mRNA Wellcome Trust Centre for Cell Biology, ICMB, University of Edinburgh, from the nucleus to the cytoplasm (nucleocytoplasmic shut- Kings’ Buildings, Edinburgh EH9 3JR, UK tling) was first observed for the heterogeneous nuclear *e-mail: [email protected] ribonucleoprotein (hnRNP) proteins. Some hnRNP proteins †e-mail: [email protected] are stripped from the mRNA at export [1], but hnRNP A1, Current Opinion in Cell Biology 2001, 13:320–325 A2, E, I and K are all exported (see [2]). Although roles for 0955-0674/01/$ — see front matter these hnRNP proteins in transport and translation have been © 2001 Elsevier Science Ltd. All rights reserved. reported [3•,4•], their affects on mRNA stability have been little studied. More is known about hnRNP D/AUF1 and Abbreviations AREs AU-rich sequence elements another nuclear RNA-binding protein, HuR, which act CBC cap-binding complex antagonistically to modulate the stability of a range of DAN deadenylating nuclease mRNAs containing AU-rich sequence elements (AREs) DSEs downstream sequence elements (reviewed in [2]). -

Supplemental Information
Supplemental information Dissection of the genomic structure of the miR-183/96/182 gene. Previously, we showed that the miR-183/96/182 cluster is an intergenic miRNA cluster, located in a ~60-kb interval between the genes encoding nuclear respiratory factor-1 (Nrf1) and ubiquitin-conjugating enzyme E2H (Ube2h) on mouse chr6qA3.3 (1). To start to uncover the genomic structure of the miR- 183/96/182 gene, we first studied genomic features around miR-183/96/182 in the UCSC genome browser (http://genome.UCSC.edu/), and identified two CpG islands 3.4-6.5 kb 5’ of pre-miR-183, the most 5’ miRNA of the cluster (Fig. 1A; Fig. S1 and Seq. S1). A cDNA clone, AK044220, located at 3.2-4.6 kb 5’ to pre-miR-183, encompasses the second CpG island (Fig. 1A; Fig. S1). We hypothesized that this cDNA clone was derived from 5’ exon(s) of the primary transcript of the miR-183/96/182 gene, as CpG islands are often associated with promoters (2). Supporting this hypothesis, multiple expressed sequences detected by gene-trap clones, including clone D016D06 (3, 4), were co-localized with the cDNA clone AK044220 (Fig. 1A; Fig. S1). Clone D016D06, deposited by the German GeneTrap Consortium (GGTC) (http://tikus.gsf.de) (3, 4), was derived from insertion of a retroviral construct, rFlpROSAβgeo in 129S2 ES cells (Fig. 1A and C). The rFlpROSAβgeo construct carries a promoterless reporter gene, the β−geo cassette - an in-frame fusion of the β-galactosidase and neomycin resistance (Neor) gene (5), with a splicing acceptor (SA) immediately upstream, and a polyA signal downstream of the β−geo cassette (Fig. -

Sequences at the Exon-Intron Boundaries* (Split Gene/Mrna Splicing/Eukaryotic Gene Structure) R
Proc. Nati. Acad. Sci. USA Vol. 75, No. 10, pp. 4853-4857, October 1978 Biochemistry Ovalbumin gene: Evidence for a leader sequence in mRNA and DNA sequences at the exon-intron boundaries* (split gene/mRNA splicing/eukaryotic gene structure) R. BREATHNACH, C. BENOIST, K. O'HARE, F. GANNON, AND P. CHAMBON Laboratoire de Genetique Mol6culaire des Eucaryotes du Centre National de la Recherche Scientifique, Unite 44 de l'Institut National de la Sant6 et de la Recherche MWdicale, Institut de Chimie Biologique, Facult6 de Melecine, Strasbourg 67085, France Communicated by A. Frey-Wyssling, July 31, 1978 ABSTRACT Selected regions of cloned EcoRI fragments the 5' end of ov-mRNA and have revealed some interesting of the chicken ovalbumin gene have been sequenced. The po- features in the DNA sequences at exon-intron boundaries. sitions where the sequences coding for ovalbumin mRNA (ov- mRNA) are interrupted in the genome have been determined, and a previously unreported interruption in the DNA sequences MATERIALS AND METHODS coding for the 5' nontranslated region of the messenger has been discovered. Because directly repeated sequences are found at Plasmid pCR1 ov 2.1 containing the ov-ds-cDNA insert was exon-intron boundaries, the nucleotide sequence alone cannot prepared as described (9). EcoRI fragments "b," "c," and "d" define unique excision-ligation points for the processing of a previously cloned in X vectors (3) were transferred to the plas- possible ov-mRNA precursor. However, the sequences in these mid pBR 322. An EcoRI/HindIII of the EcoRI boundary regions share common features; this leads to the fragment proposal that there are, in fact, unique excision-ligation points fragment "a" containing the entirety of exon 7 (Fig. -

Nuclear PTEN Safeguards Pre-Mrna Splicing to Link Golgi Apparatus for Its Tumor Suppressive Role
ARTICLE DOI: 10.1038/s41467-018-04760-1 OPEN Nuclear PTEN safeguards pre-mRNA splicing to link Golgi apparatus for its tumor suppressive role Shao-Ming Shen1, Yan Ji2, Cheng Zhang1, Shuang-Shu Dong2, Shuo Yang1, Zhong Xiong1, Meng-Kai Ge1, Yun Yu1, Li Xia1, Meng Guo1, Jin-Ke Cheng3, Jun-Ling Liu1,3, Jian-Xiu Yu1,3 & Guo-Qiang Chen1 Dysregulation of pre-mRNA alternative splicing (AS) is closely associated with cancers. However, the relationships between the AS and classic oncogenes/tumor suppressors are 1234567890():,; largely unknown. Here we show that the deletion of tumor suppressor PTEN alters pre-mRNA splicing in a phosphatase-independent manner, and identify 262 PTEN-regulated AS events in 293T cells by RNA sequencing, which are associated with significant worse outcome of cancer patients. Based on these findings, we report that nuclear PTEN interacts with the splicing machinery, spliceosome, to regulate its assembly and pre-mRNA splicing. We also identify a new exon 2b in GOLGA2 transcript and the exon exclusion contributes to PTEN knockdown-induced tumorigenesis by promoting dramatic Golgi extension and secretion, and PTEN depletion significantly sensitizes cancer cells to secretion inhibitors brefeldin A and golgicide A. Our results suggest that Golgi secretion inhibitors alone or in combination with PI3K/Akt kinase inhibitors may be therapeutically useful for PTEN-deficient cancers. 1 Department of Pathophysiology, Key Laboratory of Cell Differentiation and Apoptosis of Chinese Ministry of Education, Shanghai Jiao Tong University School of Medicine (SJTU-SM), Shanghai 200025, China. 2 Institute of Health Sciences, Shanghai Institutes for Biological Sciences of Chinese Academy of Sciences and SJTU-SM, Shanghai 200025, China.