Chromosome 3
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Human Chromosome‐Specific Aneuploidy Is Influenced by DNA
Article Human chromosome-specific aneuploidy is influenced by DNA-dependent centromeric features Marie Dumont1,†, Riccardo Gamba1,†, Pierre Gestraud1,2,3, Sjoerd Klaasen4, Joseph T Worrall5, Sippe G De Vries6, Vincent Boudreau7, Catalina Salinas-Luypaert1, Paul S Maddox7, Susanne MA Lens6, Geert JPL Kops4 , Sarah E McClelland5, Karen H Miga8 & Daniele Fachinetti1,* Abstract Introduction Intrinsic genomic features of individual chromosomes can contri- Defects during cell division can lead to loss or gain of chromosomes bute to chromosome-specific aneuploidy. Centromeres are key in the daughter cells, a phenomenon called aneuploidy. This alters elements for the maintenance of chromosome segregation fidelity gene copy number and cell homeostasis, leading to genomic instabil- via a specialized chromatin marked by CENP-A wrapped by repeti- ity and pathological conditions including genetic diseases and various tive DNA. These long stretches of repetitive DNA vary in length types of cancers (Gordon et al, 2012; Santaguida & Amon, 2015). among human chromosomes. Using CENP-A genetic inactivation in While it is known that selection is a key process in maintaining aneu- human cells, we directly interrogate if differences in the centro- ploidy in cancer, a preceding mis-segregation event is required. It was mere length reflect the heterogeneity of centromeric DNA-depen- shown that chromosome-specific aneuploidy occurs under conditions dent features and whether this, in turn, affects the genesis of that compromise genome stability, such as treatments with micro- chromosome-specific aneuploidy. Using three distinct approaches, tubule poisons (Caria et al, 1996; Worrall et al, 2018), heterochro- we show that mis-segregation rates vary among different chromo- matin hypomethylation (Fauth & Scherthan, 1998), or following somes under conditions that compromise centromere function. -

3 Chromosome Chapter
Chromosome 3 ©Chromosome Disorder Outreach Inc. (CDO) Technical genetic content provided by Dr. Iosif Lurie, M.D. Ph.D Medical Geneticist and CDO Medical Consultant/Advisor. Ideogram courtesy of the University of Washington Department of Pathology: ©1994 David Adler.hum_03.gif Introduction The size of chromosome 3 is ~200 Mb. Within this chromosome, there are thousands of genes, many of which are necessary for normal intellectual development or involved in the formation of body organs. Deletions of Chromosome 3 The length of the short arm of chromosome 3 is ~90 Mb. Most known deletions of 3p are caused by the loss of its distal 15 Mb segment (3p25–pter). Deletions of the more proximal segments are relatively rare; there are only ~50 reports on such patients. Therefore, it would be premature to talk about any syndrome related to deletions of the proximal part of 3p. The location of the breakpoints, size of deletion, and reported abnormalities are different in most described patients. However, recurrent aortal stenosis in patients with deletion 3p11p14.2, abnormal lung lobation in patients with deletion 3p12p14.2, agenesis or hypoplasia of the corpus callosum in patients with deletion 3p13, microphthalmia and coloboma in patients with deletion 3p13p21.1, choanal atresia and absent gallbladder in patients with deletion 3p13p21, and hearing loss in patients with deletion 3p14 are all indicators that the above–mentioned segments likely contain genes involved in the formation of these systems. Deletions of 3p Deletion of 3p25–pter The most distal segment of the short arm of chromosome 3 is 3p26 and spans ~8 Mb. -

Detailed Genetic and Physical Map of the 3P Chromosome Region Surrounding the Familial Renal Cell Carcinoma Chromosome Translocation, T(3;8)(Pl4.2;Q24.1)1
[CANCER RESEARCH 53. 3118-3124. July I. 1993] Detailed Genetic and Physical Map of the 3p Chromosome Region Surrounding the Familial Renal Cell Carcinoma Chromosome Translocation, t(3;8)(pl4.2;q24.1)1 Sal LaForgia,2 Jerzy Lasota, Parida Latif, Leslie Boghosian-Sell, Kumar Kastury, Masataka Olita, Teresa Druck, Lakshmi Atchison, Linda A. Cannizzaro, Gilad Barnea, Joseph Schlessinger, William Modi, Igor Kuzmin, Kaiman Tory, Berton Zbar, Carlo M. Croce, Michael Lerman, and Kay Huebner3 Jefferson Cancer Institute. Thomas Jefferson Medical College. Philadelphia, Pennsylvania 19107 (S. L. J. L. L B-S.. K. K.. M. O.. T. D.. L A. C.. C. M. C.. K. H.I: Laboratory of Immunobiology. National Cancer Institute. Frederick Cancer Research and Development Center. Frederick. Maryland 21701 (F. L, l. K.. K. T.. B. Z.. M. L): Biological Carcinogenesis and Development Program. Program Resources Inc./Dyn Corp.. Frederick Cancer Research and Development Center. Frederick. Maryland 21701 1W. M.Õ: Chestnut Hill College. Philadelphia. Pennsylvania 19118 (L A.): and Department oj Pharmacology. New York University. New York. New York 10012 (G. B., J. S.I ABSTRACT location of the critical 3p region(s) harboring the target gene(s) had been hampered by the paucity of well-localized, widely available Extensive studies of loss of heterozygosity of 3p markers in renal cell molecular probes. Recently, efforts to isolate and localize large num carcinomas (RCCs) have established that there are at least three regions bers of 3p molecular probes have been undertaken (25-28). As the critical in kidney tumorigenesis, one most likely coincident with the von Hippel-Lindau gene at 3p25.3, one in 3p21 which may also be critical in probe density on 3p increased, in parallel with recent LOH studies, it small cell lung carcinomas, and one in 3pl3-pl4.2, a region which includes became clear that multiple independent loci on 3p were involved the 3p chromosome translocation break of familial RCC with the t(3;8)- (summarized in Refs. -

De Novo Centromere Formation on a Chromosome Fragment in Maize
De novo centromere formation on a chromosome fragment in maize Shulan Fua,1, Zhenling Lva,1, Zhi Gaob,1, Huajun Wuc, Junling Pangc, Bing Zhanga, Qianhua Donga, Xiang Guoa, Xiu-Jie Wangc, James A. Birchlerb,2, and Fangpu Hana,2 aState Key Laboratory of Plant Cell and Chromosome Engineering, Institute of Genetics and Developmental Biology, Chinese Academy of Sciences, Beijing 100101, China; bDivision of Biological Science, University of Missouri, Columbia, MO 65211-7400; and cState Key Laboratory of Plant Genomics, Institute of Genetics and Developmental Biology, Chinese Academy of Sciences, Beijing 100101, China Contributed by James A. Birchler, March 1, 2013 (sent for review February 3, 2013) The centromere is the part of the chromosome that organizes the been described for a barley chromosome (14) and a maize chro- kinetochore, which mediates chromosome movement during mito- mosome fragment in oat (15). Here we provide evidence that a sis and meiosis. A small fragment from chromosome 3, named small chromosome fragment in maize, which was recovered from Duplication 3a (Dp3a), was described from UV-irradiated materials UV irradiation decades ago, lacks CentC and CRM sequences but by Stadler and Roman in the 1940s [Stadler LJ, Roman H (1948) Ge- nevertheless has a de novo site with centromere function through- netics 33(3):273–303]. The genetic behavior of Dp3a is reminiscent of out the life cycle. This small chromosome fragment has a 350-kb a ring chromosome, but fluoresecent in situ hybridization detected CENH3-binding region, which is involved in new centromere for- telomeres at both ends, suggesting a linear structure. -

Arabidopsis MZT1 Homologs GIP1 and GIP2 Are Essential for Centromere Architecture
Arabidopsis MZT1 homologs GIP1 and GIP2 are essential for centromere architecture Morgane Batzenschlagera, Inna Lermontovab, Veit Schubertb, Jörg Fuchsb, Alexandre Berra, Maria A. Koinic, Guy Houlnéa, Etienne Herzoga, Twan Ruttenb, Abdelmalek Aliouaa, Paul Franszc, Anne-Catherine Schmita, and Marie-Edith Chaboutéa,1 aInstitut de biologie moléculaire des plantes, CNRS, Université de Strasbourg, 67000 Strasbourg, France; bLeibniz Institute of Plant Genetics and Crop Plant Research OT Gatersleben, D-06466 Stadt Seeland, Germany; and cSwammerdam Institute for Life Sciences, University of Amsterdam, 1098 XH, Amsterdam, The Netherlands Edited by James A. Birchler, University of Missouri, Columbia, MO, and approved May 12, 2015 (received for review April 2, 2015) Centromeres play a pivotal role in maintaining genome integrity Previously, we characterized the γ-tubulin complex protein 3- by facilitating the recruitment of kinetochore and sister-chromatid interacting proteins (GIPs), GIP1 and GIP2 (Table S1), as es- cohesion proteins, both required for correct chromosome segre- sential for the recruitment of γ-tubulin complexes at microtubule gation. Centromeres are epigenetically specified by the presence (MT) organizing centers in Arabidopsis (7, 8). This function seems of the histone H3 variant (CENH3). In this study, we investigate the conserved in the human and Schizosaccharomyces pombe GIP role of the highly conserved γ-tubulin complex protein 3-interact- homologs named mitotic spindle organizing protein 1 (MZT1) ing proteins (GIPs) in Arabidopsis centromere regulation. We show (9–11). More recently, we localized GIPs at the nucleoplasm pe- that GIPs form a complex with CENH3 in cycling cells. GIP depletion riphery, close to chromocenters, where they modulate the nuclear in the gip1gip2 knockdown mutant leads to a decreased CENH3 architecture (12, 13). -

Mutagen Exposures and Chromosome 3 Aberrations in Acute Myelocytic Leukemia R Lindquist1, AM Forsblom1,ÅO¨ St2 and G Gahrton1
Leukemia (2000) 14, 112–118 2000 Macmillan Publishers Ltd All rights reserved 0887-6924/00 $15.00 www.nature.com/leu Mutagen exposures and chromosome 3 aberrations in acute myelocytic leukemia R Lindquist1, AM Forsblom1,ÅO¨ st2 and G Gahrton1 1Department of Hematology, Karolinska Institutet at Huddinge University Hospital, Huddinge; and 2Department of Pathology and Cytology, Karolinska Institutet and Medilab, Stockholm, Sweden Thirteen patients with acute myelocytic leukemia (AML) and have had such exposures or other exposures of mutagens such with clonal aberrations involving chromosome 3 were studied. as previous therapies with antineoplastic agents and/or Three patients had monosomy 3, four had trisomy 3, and six had structural aberrations of chromosome 3. In the majority of radiation. cases chromosome 3 aberrations were parts of complex kary- otypes, but in two patients, the abnormalities appeared as sin- gle aberrations, one as an interstitial deletion del(3)(p13p21) Materials and methods and the other as monosomy 3. All breakpoints of chromosome 3 were found in the fragile site regions 3p14.2, 3q21 and 3q26– Patients 27. All patients with monosomy 3 or structural aberrations of chromosome 3 and one of the four patients with trisomy 3 had been exposed to mutagens, such as occupational exposures Thirteen patients with AML and chromosome 3 aberrations to organic solvents and/or petroleum products or treatments were investigated; six males and seven females. The median with irradiation or antineoplastic agents. The association age was 62 years (range 16–84) (Table 1). The patients partici- among mutagen exposure, structural chromosome 3 aber- pated in prospective treatment studies within the Leukemia rations and fragile sites in AML may indicate that targeting of Group of Middle Sweden (LGMS).19,20 Almost all patients with the mutagens to these sites is of importance for the etiology of the disease. -

Molecular Aspects of Y-Chromosome Degeneration in Drosophila
Downloaded from genome.cshlp.org on September 30, 2021 - Published by Cold Spring Harbor Laboratory Press Letter Sex chromosome evolution: Molecular aspects of Y-chromosome degeneration in Drosophila Doris Bachtrog Department of Ecology, Behavior and Evolution, University of California, San Diego, La Jolla, California 92093, USA Ancient Y-chromosomes of various organisms contain few active genes and an abundance of repetitive DNA. The neo-Y chromosome of Drosophila miranda is in transition from an ordinary autosome to a genetically inert Y-chromosome, while its homolog, the neo-X chromosome, is evolving partial dosage compensation. Here, I compare four large genomic regions located on the neo-sex chromosomes that contain a total of 12 homologous genes. In addition, I investigate the partial coding sequence for 56 more homologous gene pairs from the neo-sex chromosomes. Little modification has occurred on the neo-X chromosome, and genes are highly constrained at the protein level. In contrast, a diverse array of molecular changes is contributing to the observed degeneration of the neo-Y chromosome. In particular, the four large regions surveyed on the neo-Y chromosome harbor several transposable element insertions, large deletions, and a large structural rearrangement. About one-third of all neo-Y-linked genes are nonfunctional, containing either premature stop codons and/or frameshift mutations. Intact genes on the neo-Y are accumulating amino acid and unpreferred codon changes. In addition, both 5Ј- and 3Ј-flanking regions of genes and intron sequences are less constrained on the neo-Y relative to the neo-X. Despite heterogeneity in levels of dosage compensation along the neo-X chromosomeofD. -

Receptor Signaling Through Osteoclast-Associated Monocyte
Downloaded from http://www.jimmunol.org/ by guest on September 29, 2021 is online at: average * The Journal of Immunology The Journal of Immunology , 20 of which you can access for free at: 2015; 194:3169-3179; Prepublished online 27 from submission to initial decision 4 weeks from acceptance to publication February 2015; doi: 10.4049/jimmunol.1402800 http://www.jimmunol.org/content/194/7/3169 Collagen Induces Maturation of Human Monocyte-Derived Dendritic Cells by Signaling through Osteoclast-Associated Receptor Heidi S. Schultz, Louise M. Nitze, Louise H. Zeuthen, Pernille Keller, Albrecht Gruhler, Jesper Pass, Jianhe Chen, Li Guo, Andrew J. Fleetwood, John A. Hamilton, Martin W. Berchtold and Svetlana Panina J Immunol cites 43 articles Submit online. Every submission reviewed by practicing scientists ? is published twice each month by Submit copyright permission requests at: http://www.aai.org/About/Publications/JI/copyright.html Author Choice option Receive free email-alerts when new articles cite this article. Sign up at: http://jimmunol.org/alerts http://jimmunol.org/subscription Freely available online through http://www.jimmunol.org/content/suppl/2015/02/27/jimmunol.140280 0.DCSupplemental This article http://www.jimmunol.org/content/194/7/3169.full#ref-list-1 Information about subscribing to The JI No Triage! Fast Publication! Rapid Reviews! 30 days* Why • • • Material References Permissions Email Alerts Subscription Author Choice Supplementary The Journal of Immunology The American Association of Immunologists, Inc., 1451 Rockville Pike, Suite 650, Rockville, MD 20852 Copyright © 2015 by The American Association of Immunologists, Inc. All rights reserved. Print ISSN: 0022-1767 Online ISSN: 1550-6606. -

3P26 Deletions FTNW
3p26 deletions rarechromo.org Genes and chromosomes Our bodies are made up of trillions of cells. Most of the cells contain a set of around 20,000 different genes; this genetic information tells the body how to develop, grow and function. Genes are carried on structures called chromosomes. Chromosomes usually come in pairs, one chromosome from p arm each parent. Of the 46 chromosomes, two are a pair of sex chromosomes: (two Xs for a girl and an X and a Y for a boy). The remaining 44 chromosomes are grouped into 22 pairs and are numbered 1 to 22, approximately from largest to smallest. These are called autosomes. Each chromosome has a short (p) arm (from petit, the French for small) and a long (q) arm (see diagram, right). People with a 3p26 deletion have lost DNA from the area of the diagram marked in red. Chromosome Deletions A sperm cell from the father and an egg cell from the mother each carries just one copy of each chromosome. q arm When they join together they form a single cell that now carries two copies of each chromosome. This cell must make many copies of itself (and all the chromosomes and genetic material) in order to make all of the many cells that form during human growth and development. Sometimes during the formation of the egg or sperm cells or during this complicated copying and replication process, parts of the Chromosome 3 chromosomes can break off or become arranged differently from usual. People with a 3p26 deletion have one intact chromosome 3, but a piece from the short arm of the other copy is missing. -

3P Duplications
3p duplications rarechromo.org Duplication of 3p A duplication of 3p is a rare genetic condition caused by an extra part of one of the body’s 46 chromosomes – chromosome 3. For a healthy development, chromosomes should contain just the right amount of genetic material – not too much and not too little. A 3p duplication can result in developmental delay and congenital (heart) malformations. What are chromosomes? Chromosomes are made up mostly of DNA and are the structures in each of the body’s cells that carry the genetic information in the form of genes that tells the body how to develop, grow and function. Chromosomes usually come in pairs, with one chromosome from each pair coming from the father and one from the mother. Of the 46 chromosomes, two are a pair of sex chromosomes, XX (two X chromosomes) in females and XY (one X and one Y chromosome) in males. The remaining 44 chromosomes are grouped in 22 pairs, numbered 1 to 22 from largest to smallest. Chromosomes have a short arm, named p (shown at the top in the figure), and a long arm, named q (shown at the bottom in the figure). The two arms of a chromosome meet at a point called the centromere. q arm centromere p arm Looking at chromosome 3p You can’t see chromosomes with the naked eye, but if you stain them and magnify them many hundreds of times under a microscope, you can see that each one has a distinctive pattern of light and dark bands. Looking at chromosomes under a microscope, it may be possible to see the genetic material that is missing or extra, if the piece is large enough. -

Chromosomal Instability Is Associated with Higher Expression of Genes
Volume 10 Number 11 November 2008 pp. 1222–1230 1222 www.neoplasia.com Anna V. Roschke*, Oleg K. Glebov*, Chromosomal Instability Is † Samir Lababidi , Kristen S. Gehlhaus*, † Associated with Higher John N. Weinstein and Ilan R. Kirsch* Expression of Genes Implicated *Genetics Branch, Center for Cancer Research, National Cancer Institute, Bethesda, MD 20889-5105, USA; in Epithelial-Mesenchymal † Laboratory of Molecular Pharmacology, Center for Transition, Cancer Invasiveness, Cancer Research, National Cancer Institute, Bethesda, and Metastasis and with MD 20889-5105, USA Lower Expression of Genes Involved in Cell Cycle Checkpoints, DNA Repair, and Chromatin Maintenance1 Abstract Chromosomal instability—a hallmark of epithelial cancers—is an ongoing process that results in aneuploidy and karyotypic heterogeneity of a cancer cell population. Previously, we stratified cancer cell lines in the NCI-60 drug discovery panel based on their karyotypic complexity and heterogeneity. Using this stratification in conjunction with drug response data for the cell lines allowed us to identify classes of chemical compounds whose growth- inhibitory activity correlates with karyotypic complexity and chromosomal instability. In this article, we asked the question: What are the biological processes, pathways, or genes associated with chromosomal instability of can- cer cells? We found that increased instability of the chromosomal content in a cancer cell population, particularly, persistent gains and losses of chromosomes, is associated with elevated expression of genes involved with aggressive cellular behavior, including invasion- and metastasis-associated changes in cell communication, ad- hesion, motility, and migration. These same karyotypic features are negatively correlated with the expression of genes involved in cell cycle checkpoints, DNA repair, and chromatin maintenance. -
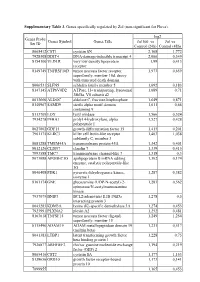
Supplementary Table 3. Genes Specifically Regulated by Zol (Non-Significant for Fluva)
Supplementary Table 3. Genes specifically regulated by Zol (non-significant for Fluva). log2 Genes Probe Genes Symbol Genes Title Zol100 vs Zol vs Set ID Control (24h) Control (48h) 8065412 CST1 cystatin SN 2,168 1,772 7928308 DDIT4 DNA-damage-inducible transcript 4 2,066 0,349 8154100 VLDLR very low density lipoprotein 1,99 0,413 receptor 8149749 TNFRSF10D tumor necrosis factor receptor 1,973 0,659 superfamily, member 10d, decoy with truncated death domain 8006531 SLFN5 schlafen family member 5 1,692 0,183 8147145 ATP6V0D2 ATPase, H+ transporting, lysosomal 1,689 0,71 38kDa, V0 subunit d2 8013660 ALDOC aldolase C, fructose-bisphosphate 1,649 0,871 8140967 SAMD9 sterile alpha motif domain 1,611 0,66 containing 9 8113709 LOX lysyl oxidase 1,566 0,524 7934278 P4HA1 prolyl 4-hydroxylase, alpha 1,527 0,428 polypeptide I 8027002 GDF15 growth differentiation factor 15 1,415 0,201 7961175 KLRC3 killer cell lectin-like receptor 1,403 1,038 subfamily C, member 3 8081288 TMEM45A transmembrane protein 45A 1,342 0,401 8012126 CLDN7 claudin 7 1,339 0,415 7993588 TMC7 transmembrane channel-like 7 1,318 0,3 8073088 APOBEC3G apolipoprotein B mRNA editing 1,302 0,174 enzyme, catalytic polypeptide-like 3G 8046408 PDK1 pyruvate dehydrogenase kinase, 1,287 0,382 isozyme 1 8161174 GNE glucosamine (UDP-N-acetyl)-2- 1,283 0,562 epimerase/N-acetylmannosamine kinase 7937079 BNIP3 BCL2/adenovirus E1B 19kDa 1,278 0,5 interacting protein 3 8043283 KDM3A lysine (K)-specific demethylase 3A 1,274 0,453 7923991 PLXNA2 plexin A2 1,252 0,481 8163618 TNFSF15 tumor necrosis