Close Encounters of the Third Domain: the Emerging Genomic View of Archaeal Diversity and Evolution
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Revised Glossary for AQA GCSE Biology Student Book
Biology Glossary amino acids small molecules from which proteins are A built abiotic factor physical or non-living conditions amylase a digestive enzyme (carbohydrase) that that affect the distribution of a population in an breaks down starch ecosystem, such as light, temperature, soil pH anaerobic respiration respiration without using absorption the process by which soluble products oxygen of digestion move into the blood from the small intestine antibacterial chemicals chemicals produced by plants as a defence mechanism; the amount abstinence method of contraception whereby the produced will increase if the plant is under attack couple refrains from intercourse, particularly when an egg might be in the oviduct antibiotic e.g. penicillin; medicines that work inside the body to kill bacterial pathogens accommodation ability of the eyes to change focus antibody protein normally present in the body acid rain rain water which is made more acidic by or produced in response to an antigen, which it pollutant gases neutralises, thus producing an immune response active site the place on an enzyme where the antimicrobial resistance (AMR) an increasing substrate molecule binds problem in the twenty-first century whereby active transport in active transport, cells use energy bacteria have evolved to develop resistance against to transport substances through cell membranes antibiotics due to their overuse against a concentration gradient antiretroviral drugs drugs used to treat HIV adaptation features that organisms have to help infections; they -

Marsarchaeota Are an Aerobic Archaeal Lineage Abundant in Geothermal Iron Oxide Microbial Mats
Marsarchaeota are an aerobic archaeal lineage abundant in geothermal iron oxide microbial mats Authors: Zackary J. Jay, Jacob P. Beam, Mansur Dlakic, Douglas B. Rusch, Mark A. Kozubal, and William P. Inskeep This is a postprint of an article that originally appeared in Nature Microbiology on May 14, 2018. The final version can be found at https://dx.doi.org/10.1038/s41564-018-0163-1. Jay, Zackary J. , Jacob P. Beam, Mensur Dlakic, Douglas B. Rusch, Mark A. Kozubal, and William P. Inskeep. "Marsarchaeota are an aerobic archaeal lineage abundant in geothermal iron oxide microbial mats." Nature Microbiology 3, no. 6 (May 2018): 732-740. DOI: 10.1038/ s41564-018-0163-1. Made available through Montana State University’s ScholarWorks scholarworks.montana.edu Marsarchaeota are an aerobic archaeal lineage abundant in geothermal iron oxide microbial mats Zackary J. Jay1,4,7, Jacob P. Beam1,5,7, Mensur Dlakić2, Douglas B. Rusch3, Mark A. Kozubal1,6 and William P. Inskeep 1* The discovery of archaeal lineages is critical to our understanding of the universal tree of life and evolutionary history of the Earth. Geochemically diverse thermal environments in Yellowstone National Park provide unprecedented opportunities for studying archaea in habitats that may represent analogues of early Earth. Here, we report the discovery and character- ization of a phylum-level archaeal lineage proposed and herein referred to as the ‘Marsarchaeota’, after the red planet. The Marsarchaeota contains at least two major subgroups prevalent in acidic, microaerobic geothermal Fe(III) oxide microbial mats across a temperature range from ~50–80 °C. Metagenomics, single-cell sequencing, enrichment culturing and in situ transcrip- tional analyses reveal their biogeochemical role as facultative aerobic chemoorganotrophs that may also mediate the reduction of Fe(III). -

Metagenomic Insights Into the Uncultured Diversity and Physiology of Microbes in Four Hypersaline Soda Lake Brines
Lawrence Berkeley National Laboratory Recent Work Title Metagenomic Insights into the Uncultured Diversity and Physiology of Microbes in Four Hypersaline Soda Lake Brines. Permalink https://escholarship.org/uc/item/9xc5s0v5 Journal Frontiers in microbiology, 7(FEB) ISSN 1664-302X Authors Vavourakis, Charlotte D Ghai, Rohit Rodriguez-Valera, Francisco et al. Publication Date 2016 DOI 10.3389/fmicb.2016.00211 Peer reviewed eScholarship.org Powered by the California Digital Library University of California ORIGINAL RESEARCH published: 25 February 2016 doi: 10.3389/fmicb.2016.00211 Metagenomic Insights into the Uncultured Diversity and Physiology of Microbes in Four Hypersaline Soda Lake Brines Charlotte D. Vavourakis 1, Rohit Ghai 2, 3, Francisco Rodriguez-Valera 2, Dimitry Y. Sorokin 4, 5, Susannah G. Tringe 6, Philip Hugenholtz 7 and Gerard Muyzer 1* 1 Microbial Systems Ecology, Department of Aquatic Microbiology, Institute for Biodiversity and Ecosystem Dynamics, University of Amsterdam, Amsterdam, Netherlands, 2 Evolutionary Genomics Group, Departamento de Producción Vegetal y Microbiología, Universidad Miguel Hernández, San Juan de Alicante, Spain, 3 Department of Aquatic Microbial Ecology, Biology Centre of the Czech Academy of Sciences, Institute of Hydrobiology, Ceskéˇ Budejovice,ˇ Czech Republic, 4 Research Centre of Biotechnology, Winogradsky Institute of Microbiology, Russian Academy of Sciences, Moscow, Russia, 5 Department of Biotechnology, Delft University of Technology, Delft, Netherlands, 6 The Department of Energy Joint Genome Institute, Walnut Creek, CA, USA, 7 Australian Centre for Ecogenomics, School of Chemistry and Molecular Biosciences and Institute for Molecular Bioscience, The University of Queensland, Brisbane, QLD, Australia Soda lakes are salt lakes with a naturally alkaline pH due to evaporative concentration Edited by: of sodium carbonates in the absence of major divalent cations. -

Direct Charging of Trnacua with Pyrrolysine in Vitro and in Vivo
letters to nature .............................................................. gene product (see Supplementary Fig. S1). The tRNA pool extracted from Methanosarcina acetivorans or tRNACUA transcribed in vitro Direct charging of tRNACUA with was used in charging experiments. Charged and uncharged tRNA species were separated by electrophoresis in a denaturing acid-urea pyrrolysine in vitro and in vivo 10,11 polyacrylamide gel and tRNACUA was specifically detected by northern blotting with an oligonucleotide probe. The oligonucleo- Sherry K. Blight1*, Ross C. Larue1*, Anirban Mahapatra1*, tide complementary to tRNA could hybridize to a tRNA in the David G. Longstaff1, Edward Chang1, Gang Zhao2†, Patrick T. Kang4, CUA Kari B. Green-Church5, Michael K. Chan2,3,4 & Joseph A. Krzycki1,4 pool of tRNAs isolated from wild-type M. acetivorans but not to the tRNA pool from a pylT deletion mutant of M. acetivorans (A.M., 1Department of Microbiology, 484 West 12th Avenue, 2Department of Chemistry, A. Patel, J. Soares, R.L. and J.A.K., unpublished observations). 3 100 West 18th Avenue, Department of Biochemistry, 484 West 12th Avenue, Both tRNACUA and aminoacyl-tRNACUA were detectable in the The Ohio State University, Columbus, Ohio 43210, USA isolated cellular tRNA pool (Fig. 1). Alkaline hydrolysis deacylated 4Ohio State University Biochemistry Program, 484 West 12th Avenue, The Ohio the cellular charged species, but subsequent incubation with pyrro- State University, Columbus, Ohio 43210, USA lysine, ATP and PylS-His6 resulted in maximal conversion of 50% of 5CCIC/Mass Spectrometry and Proteomics Facility, The Ohio State University, deacylated tRNACUA to a species that migrated with the same 116 W 19th Ave, Columbus, Ohio 43210, USA electrophoretic mobility as the aminoacyl-tRNACUA present in the * These authors contributed equally to this work. -
18.4 Bacteria and Archaea Kingdom Eubacteria Domain Bacteria
18.4 Bacteria and Archaea Kingdom Eubacteria Domain Bacteria 18.4 Bacteria and Archaea Description Bacteria are single-celled prokaryotes. 18.4 Bacteria and Archaea Where do they live? Prokaryotes are widespread on Earth. ( Est. over 1 billion types of bacteria, and over 1030 individual prokaryote cells on earth.) Found in all land and ocean environments, even inside other organisms! 18.4 Bacteria and Archaea Common Examples • E. Coli • Tetanus bacteria • Salmonella bacteria • Tuberculosis bacteria • Staphylococcus • Streptococcus 18.4 Bacteria and Archaea Modes Of Nutrition • Bacteria may be heterotrophs or autotrophs 18.4 Bacteria and Archaea Bacteria Reproduce How? • by binary fission. • exchange genes during conjugation= conjugation bridge increases diversity. • May survive by forming endospores = specialized cell with thick protective cell wall. TEM; magnification 6000x • Can survive for centuries until environment improves. Have been found in mummies! 18.4 Bacteria and Archaea • Bacteria Diagram – plasmid = small piece of genetic material, can replicate independently of the chromosome – flagellum = different than in eukaryotes, but for movement – pili = used to stick the bacteria to eachpili other or surfaces plasma membrance flagellum chromosome cell wall plasmid This diagram shows the typical structure of a prokaryote. Archaea and bacteria look very similar, although they have important molecular differences. 18.4 Bacteria and Archaea • Classified by: their need for oxygen, how they gram stain, and their shapes 18.4 Bacteria and Archaea Main Groups by Shapes – rod-shaped, called bacilli – spiral, called spirilla or spirochetes – spherical, called cocci Spirochaeta: spiral Lactobacilli: rod-shaped Enterococci: spherical 18.4 Bacteria and Archaea • Main Groups by their need for oxygen. -
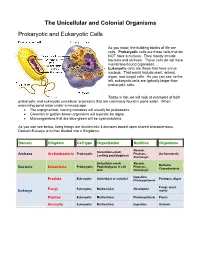
The Unicellular and Colonial Organisms Prokaryotic And
The Unicellular and Colonial Organisms Prokaryotic and Eukaryotic Cells As you know, the building blocks of life are cells. Prokaryotic cells are those cells that do NOT have a nucleus. They mostly include bacteria and archaea. These cells do not have membrane-bound organelles. Eukaryotic cells are those that have a true nucleus. That would include plant, animal, algae, and fungal cells. As you can see, to the left, eukaryotic cells are typically larger than prokaryotic cells. Today in lab, we will look at examples of both prokaryotic and eukaryotic unicellular organisms that are commonly found in pond water. When examining pond water under a microscope… The unpigmented, moving microbes will usually be protozoans. Greenish or golden-brown organisms will typically be algae. Microorganisms that are blue-green will be cyanobacteria. As you can see below, living things are divided into 3 domains based upon shared characteristics. Domain Eukarya is further divided into 4 Kingdoms. Domain Kingdom Cell type Organization Nutrition Organisms Absorb, Unicellular-small; Prokaryotic Photsyn., Archaeacteria Archaea Archaebacteria Lacking peptidoglycan Chemosyn. Unicellular-small; Absorb, Bacteria, Prokaryotic Peptidoglycan in cell Photsyn., Bacteria Eubacteria Cyanobacteria wall Chemosyn. Ingestion, Eukaryotic Unicellular or colonial Protozoa, Algae Protista Photosynthesis Fungi, yeast, Fungi Eukaryotic Multicellular Absorption Eukarya molds Plantae Eukaryotic Multicellular Photosynthesis Plants Animalia Eukaryotic Multicellular Ingestion Animals Prokaryotic Organisms – the archaea, non-photosynthetic bacteria, and cyanobacteria Archaea - Microorganisms that resemble bacteria, but are different from them in certain aspects. Archaea cell walls do not include the macromolecule peptidoglycan, which is always found in the cell walls of bacteria. Archaea usually live in extreme, often very hot or salty environments, such as hot mineral springs or deep-sea hydrothermal vents. -

Biology Chapter 19 Kingdom Protista Domain Eukarya Description Kingdom Protista Is the Most Diverse of All the Kingdoms
Biology Chapter 19 Kingdom Protista Domain Eukarya Description Kingdom Protista is the most diverse of all the kingdoms. Protists are eukaryotes that are not animals, plants, or fungi. Some unicellular, some multicellular. Some autotrophs, some heterotrophs. Some with cell walls, some without. Didinium protist devouring a Paramecium protist that is longer than it is! Read about it on p. 573! Where Do They Live? • Because of their diversity, we find protists in almost every habitat where there is water or at least moisture! Common Examples • Ameba • Algae • Paramecia • Water molds • Slime molds • Kelp (Sea weed) Classified By: (DON’T WRITE THIS DOWN YET!!! • Mode of nutrition • Cell walls present or not • Unicellular or multicellular Protists can be placed in 3 groups: animal-like, plantlike, or funguslike. Didinium, is a specialist, only feeding on Paramecia. They roll into a ball and form cysts when there is are no Paramecia to eat. Paramecia, on the other hand are generalists in their feeding habits. Mode of Nutrition Depends on type of protist (see Groups) Main Groups How they Help man How they Hurt man Ecosystem Roles KEY CONCEPT Animal-like protists = PROTOZOA, are single- celled heterotrophs that can move. Oxytricha Reproduce How? • Animal like • Unicellular – by asexual reproduction – Paramecium – does conjugation to exchange genetic material Animal-like protists Classified by how they move. macronucleus contractile vacuole food vacuole oral groove micronucleus cilia • Protozoa with flagella are zooflagellates. – flagella help zooflagellates swim – more than 2000 zooflagellates • Some protists move with pseudopods = “false feet”. – change shape as they move –Ex. amoebas • Some protists move with pseudopods. -

Hydrogenases of Methanogens
ANRV413-BI79-18 ARI 27 April 2010 21:0 Hydrogenases from Methanogenic Archaea, Nickel, a Novel Cofactor, and H2 Storage Rudolf K. Thauer, Anne-Kristin Kaster, Meike Goenrich, Michael Schick, Takeshi Hiromoto, and Seigo Shima Max Planck Institute for Terrestrial Microbiology, D-35043 Marburg, Germany; email: [email protected] Annu. Rev. Biochem. 2010. 79:507–36 Key Words First published online as a Review in Advance on H2 activation, energy-converting hydrogenase, complex I of the March 17, 2010 respiratory chain, chemiosmotic coupling, electron bifurcation, The Annual Review of Biochemistry is online at reversed electron transfer biochem.annualreviews.org This article’s doi: Abstract 10.1146/annurev.biochem.030508.152103 Most methanogenic archaea reduce CO2 with H2 to CH4. For the Copyright c 2010 by Annual Reviews. activation of H2, they use different [NiFe]-hydrogenases, namely All rights reserved energy-converting [NiFe]-hydrogenases, heterodisulfide reductase- 0066-4154/10/0707-0507$20.00 associated [NiFe]-hydrogenase or methanophenazine-reducing by University of Texas - Austin on 06/10/13. For personal use only. [NiFe]-hydrogenase, and F420-reducing [NiFe]-hydrogenase. The energy-converting [NiFe]-hydrogenases are phylogenetically related Annu. Rev. Biochem. 2010.79:507-536. Downloaded from www.annualreviews.org to complex I of the respiratory chain. Under conditions of nickel limitation, some methanogens synthesize a nickel-independent [Fe]- hydrogenase (instead of F420-reducing [NiFe]-hydrogenase) and by that reduce their nickel requirement. The [Fe]-hydrogenase harbors a unique iron-guanylylpyridinol cofactor (FeGP cofactor), in which a low-spin iron is ligated by two CO, one C(O)CH2-, one S-CH2-, and a sp2-hybridized pyridinol nitrogen. -

Structural Biology of the C-Terminal Domain Of
STRUCTURAL BIOLOGY OF THE C-TERMINAL DOMAIN OF EUKARYOTIC REPLICATION FACTOR MCM10 By Patrick David Robertson Dissertation Submitted to the Faculty of the Graduate School of Vanderbilt University in partial fulfillment of the requirements for the degree of DOCTOR OF PHILOSOPHY in Biological Sciences August, 2010 Nashville, Tennessee Approved: Brandt F. Eichman Walter J. Chazin James G. Patton Hassane Mchaourab To my wife Sabrina, thank you for your enduring love and support ii ACKNOWLEDGMENTS I would like to begin by expressing my sincerest gratitude to my mentor and Ph.D. advisor, Dr. Brandt Eichman. In addition to your excellent guidance and training, your passion for science has been a source of encouragement and inspiration over the past five years. I consider working with you to be a great privilege and I am grateful for the opportunity. I would also like to thank the members of my thesis committee: Drs. Walter Chazin, Ellen Fanning, James Patton, and Hassane Mchaourab. My research and training would not have been possible without your insight, advice and intellectual contributions. I would like to acknowledge all of the members of the Eichman laboratory, past and present, for their technical assistance and camaraderie over the years. I would especially like to thank Dr. Eric Warren for his contributions to the research presented here, as well for his friendship of the years. I would also like to thank Drs. Benjamin Chagot and Sivaraja Vaithiyalingam from the Chazin laboratory for their expert assistance and NMR training. I would especially like to thank my parents, Joyce and David, my brother Jeff, and the rest of my family for their love and encouragement throughout my life. -

BIRDS AS MARINE ORGANISMS: a REVIEW Calcofi Rep., Vol
AINLEY BIRDS AS MARINE ORGANISMS: A REVIEW CalCOFI Rep., Vol. XXI, 1980 BIRDS AS MARINE ORGANISMS: A REVIEW DAVID G. AINLEY Point Reyes Bird Observatory Stinson Beach, CA 94970 ABSTRACT asociadas con esos peces. Se indica que el estudio de las Only 9 of 156 avian families are specialized as sea- aves marinas podria contribuir a comprender mejor la birds. These birds are involved in marine energy cycles dinamica de las poblaciones de peces anterior a la sobre- during all aspects of their lives except for the 10% of time explotacion por el hombre. they spend in some nesting activities. As marine organ- isms their occurrence and distribution are directly affected BIRDS AS MARINE ORGANISMS: A REVIEW by properties of their oceanic habitat, such as water temp- As pointed out by Sanger(1972) and Ainley and erature, salinity, and turbidity. In their trophic relation- Sanger (1979), otherwise comprehensive reviews of bio- ships, almost all are secondary or tertiary carnivores. As logical oceanography have said little or nothing about a group within specific ecosystems, estimates of their seabirds in spite of the fact that they are the most visible feeding rates range between 20 and 35% of annual prey part of the marine biota. The reasons for this oversight are production. Their usual prey are abundant, schooling or- no doubt complex, but there are perhaps two major ones. ganisms such as euphausiids and squid (invertebrates) First, because seabirds have not been commercially har- and clupeids, engraulids, and exoccetids (fish). Their high vested to any significant degree, fisheries research, which rates of feeding and metabolism, and the large amounts of supplies most of our knowledge about marine ecosys- nutrients they return to the marine environment, indicate tems, has ignored them. -

A Knowledge-Centric E-Research Platform for Marine Life and Oceanographic Research
A KNOWLEDGE-CENTRIC E-RESEARCH PLATFORM FOR MARINE LIFE AND OCEANOGRAPHIC RESEARCH Ali Daniyal, Samina Abidi, Ashraf AbuSharekh, Mei Kuan Wong and S. S. R. Abidi Department of Computer Science, Dalhousie University, 6050 Univeristy Ave, Halifax, Canada Keywords: e-Research, Knowledge management, Web services, Marine life, Oceanography. Abstract: In this paper we present a knowledge centric e-Research platform to support collaboration between two diverse scientific communities—i.e. Oceanography and Marine Life domains. The Platform for Ocean Knowledge Management (POKM) offers a services oriented framework to facilitate the sharing, discovery and visualization of multi-modal data and scientific models. To establish interoperability between two diverse domain, we have developed a common OWL-based domain ontology that captures and interrelates concepts from the two different domain. POKM also provide semantic descriptions of the functionalities of a range of e-research oriented web services through a OWL-S service ontology that supports dynamic discovery and invocation of services. POKM has been deployed as a web-based prototype system that is capable of fetching, sharing and visualizing marine animal detection and oceanographic data from multiple global data sources. 1 INTRODUCTION (POKM)—that offers a suite of knowledge-centric services for oceanographic researchers to (a) access, To comprehensively understand how changes to the share, integrate and operationalize the data, models ecosystem impact the ocean’s physical and and knowledge resources available at multiple sites; biological parameters (Cummings, 2005) (b) collaborate in joint scientific research oceanographers and marine biologists—termed as experiments by sharing resources, results, expertise the oceanographic research community—are seeking and models; and (c) form a virtual community of more collaboration in terms of sharing domain- researchers, marine resource managers, policy specific data and knowledge (Bos, 2007). -

Characterization of Methanosarcina Barkeri MST and 227, Methanosarcina Mazei S-6T, and Methanosarcina Vacuolata Z-76IT GLORIA M
INTERNATIONALJOURNAL OF SYSTEMATICBACTERIOLOGY, Apr. 1991, p. 267-274 Vol. 41, No. 2 0020-7713/91/020267-08$02.OO/O Copyright 0 1991, International Union of Microbiological Societies Characterization of Methanosarcina barkeri MST and 227, Methanosarcina mazei S-6T, and Methanosarcina vacuolata Z-76IT GLORIA M. MAESTROJUAN' AND DAVID R. BOONE172* Departments of Environmental Science and Engineering' and Chemical and Biological Science,2 Oregon Graduate Institute, 19600 N.W. von Neumann Drive, Beaverton, Oregon 97006-1999 Members of the genus Methanosarcina are recognized as major aceticlastic methanogens, and several species which thrive in low-salt, pH-neutral culture medium at mesophilic temperatures have been described. However, the environmental conditions which support the fastest growth of these species (Methanosarcina barkeri MST [T = type strain] and 227, Methanosarcina mazei S-6T, and Methanosarcina vacuolata Z-761T) have not been reported previously. Although the members of the genus Methanosarcina are widely assumed to grow best at pH values near neutrality, we found that some strains prefer acidic pH values. M. vacuolata and the two strains of M. barkeri which we tested were acidophilic when they were grown on H, plus methanol, growing most rapidly at pH 5 and growing at pH values as low as 4.3. M. mazei grew best at pH values near neutrality. We found that all of the strains tested grew most rapidly at 37 to 42°C on all of the growth substrates which we tested. None of the strains was strongly halophilic, although the growth of some strains was slightly stimulated by small amounts of added NaCI.