What Are DNA Sequence Motifs?
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

DNA Sequencing and Sorting: Identifying Genetic Variations
BioMath DNA Sequencing and Sorting: Identifying Genetic Variations Student Edition Funded by the National Science Foundation, Proposal No. ESI-06-28091 This material was prepared with the support of the National Science Foundation. However, any opinions, findings, conclusions, and/or recommendations herein are those of the authors and do not necessarily reflect the views of the NSF. At the time of publishing, all included URLs were checked and active. We make every effort to make sure all links stay active, but we cannot make any guaranties that they will remain so. If you find a URL that is inactive, please inform us at [email protected]. DIMACS Published by COMAP, Inc. in conjunction with DIMACS, Rutgers University. ©2015 COMAP, Inc. Printed in the U.S.A. COMAP, Inc. 175 Middlesex Turnpike, Suite 3B Bedford, MA 01730 www.comap.com ISBN: 1 933223 71 5 Front Cover Photograph: EPA GULF BREEZE LABORATORY, PATHO-BIOLOGY LAB. LINDA SHARP ASSISTANT This work is in the public domain in the United States because it is a work prepared by an officer or employee of the United States Government as part of that person’s official duties. DNA Sequencing and Sorting: Identifying Genetic Variations Overview Each of the cells in your body contains a copy of your genetic inheritance, your DNA which has been passed down to you, one half from your biological mother and one half from your biological father. This DNA determines physical features, like eye color and hair color, and can determine susceptibility to medical conditions like hypertension, heart disease, diabetes, and cancer. -

Representation for Discovery of Protein Motifs
From: ISMB-93 Proceedings. Copyright © 1993, AAAI (www.aaai.org). All rights reserved. Representation for Discovery of Protein Motifs Darrell ConklinT, Suzanne Fortier~, Janice Glasgowt Departments~t of Computing and Information Sciencet and Chemistry Queen’s University Kingston, Ontario, Canada K7L 3N6 [email protected] Abstract pattern of amino acid residues. Protein motifs can be roughly classified into four categories. Sequence There are several dimensions and levels of com- motifs are linear strings of residues with an implicit plexity in which information on protein mo- topological ordering. Sequence-structure motifs are tifs may be available. For example, one- sequence motifs with secondary structure identifiers at- dimensional sequence motifs may be associated tached to one or more residues in the motif. Structure with secondary structure identifiers. Alterna- motifs are 3D structural objects, described by posi- tively, three-dimensional information on polypep- tions of residue objects in 3D Euclidian space. Apart tide segments may be used to induce prototypical from a topological linear ordering on the residues, three-dimensional structure templates. This pa- structure motifs are free of sequence information. Fi- per surveys various representations encountered nally, structure-sequence motifs are combined 1D- in the protein motif discovery literature. Many 3D structures that associate sequence information with of the representations are based on incompatible a structure motif. Figure 1 illustrates these four types semantics, making difficult the comparison and of protein motifs, along with somefurther subclassifi- combination of previous results. To make better cations which are elaborated upon later in the paper. use of machine learning techniques and to pro- The first three motif types are discussed by Thornton vide for an integrated knowledge representation and Gardner (1989); the structure-sequence motif will framework, a general representation language -- be presented in this paper. -

Introduction to Sequence Motif Discovery and Search the Complete
Introduction to sequence motif discovery and search EMBNET course Bioinformatics of transcriptional regulation Jan 28 2008 Christoph Schmid The complete genome era Components of transcriptional regulation Distal transcription-factor binding sites (enhancer) cis-regulatory modules Wasserman 5, 276-287 (2004) DNA-protein interaction Allen et al. © 1998 EMBL Jordan et al. © 1991 Prentice-Hall Sequence motifs sequence variants of site: ..A T C G C A.. ..T T G G A C.. ..T T G G T G.. ..A T C G G T.. matrix: A 2 0 0 0 1 1 (simplest version) C 0 0 2 0 1 1 G 0 0 2 4 1 1 T 2 4 0 0 1 1 + cutoff ! sequence logo: Title Representation of the binding specificity by a scoring matrix (also referred to as weight matrix) 1 2 3 4 5 6 7 8 9 A -10 -10 -14 -12 -10 5 -2 -10 -6 C 5 -10 -13 -13 -7 -15 -13 3 -4 G -3 -14 -13 -11 5 -12 -13 2 -7 T -5 5 5 5 -10 -9 5 -11 5 Strong C T T T G A T C T Binding site 5 + 5 + 5 + 5 + 5 + 5 + 5 + 3 + 5 = 43 Random A C G T A C G T A Sequence -10 -10 -13 + 5 -10 -15 -13 -11 - 6 = -83 Biophysical interpretation of protein binding sites Columns of a weight matrix characterize the specificity of base-pair acceptor sites on the protein surface. Weight matrix elements represent negated energy contributions to the total binding energy → weight matrix score inversely proportional to binding energy Motif search: Statistical over-representation of genomic sequence motifs Conserved motifs represent protein binding sites Putative conservation in nucleotide sequence (motif) position relative to: TSS other binding sites (protein complexes) -

Tools for Motif and Pattern Searching
TOOLS FOR MOTIF AND PATTERN SEARCHING Prat Thiru OUTLINE • What are motifs? • Algorithms Used and Programs Available • Workflow and Strategies • MEME/MAST Demo (online and command line) Protein Motifs DNA Motifs MEME Output Definitions • Motif: Conserved regions of protein or DNA sequences • Pattern: Qualitative description of a motif eg. regular expression C[AT]AAT[CG]X •Profile: Quantitative description of a motif eg. position weight matrix Patterns • Regular Expression Symbols ¾[ ] – OR eg. [GA] means G or A ¾{ } – NOT eg. {P,V} means not P or V ¾( ) – repeats eg. A(3) means AAA ¾X or N or “.” – any • Complex patterns representation difficult • Loose frequency information eg. [AT] vs 20%A 80%T Profiles Sequence Logos Algorithms • Enumeration • Probabilistic Optimization • Deterministic Optimization 1. Identify motifs 2. Build a consensus Enumeration • Exhaustive search: word counting method, count all n-mers and look for overrepresentation • Less likely to get stuck in a local optimum • Computationally expensive ¾YMF http://wingless.cs.washington.edu/YMF/YMFWeb/YMFInput.pl ¾Weeder http://159.149.109.9/weederaddons/locator.html Probabilistic Optimization • Uses a Gibbs sampling approach • One n-mer from each sequence is randomly picked to determine initial model. In subsequent iterations, one sequence, i, is removed and the model is recalculated. Pick a new location of motif in sequence i iterate until convergence • Assumes most sequences will have the motif ¾ AlignAce http://atlas.med.harvard.edu/cgi-bin/alignace.pl ¾ Gibbs Motif Sampler http://bayesweb.wadsworth.org/gibbs/gibbs.html Deterministic Optimization • Based on expectation maximization (EM) • EM: iteratively estimates the likelihood given the data that is present I. -

Degsampler: Gibbs Sampling Strategy for Predicting E3-Binding Sites with Position-Specific Prior Information
DegSampler: Gibbs sampling strategy for predicting E3-binding sites with position-specic prior information Osamu Maruyama ( [email protected] ) Kyushu University https://orcid.org/0000-0002-5760-1507 Fumiko Matsuzaki Kyushu University Research article Keywords: motif, substrate protein, E3 ubiquitin ligase, prior, posterior, disorder, collapsed, Gibbs sampling, DegSampler Posted Date: November 6th, 2020 DOI: https://doi.org/10.21203/rs.3.rs-101967/v1 License: This work is licensed under a Creative Commons Attribution 4.0 International License. Read Full License Maruyama and Matsuzaki RESEARCH DegSampler: Gibbs sampling strategy for predicting E3-binding sites with position-specific prior information Osamu Maruyama1* and Fumiko Matsuzaki2 Background Eukaryotic cells have two major pathways for degrad- ing proteins in order to control cellular processes and maintain intracellular homeostasis. One of the path- ways is autophagy, a catabolic process that deliv- ers intracellular components to lysosomes or vacuoles Abstract [1]. The other pathway is the ubiquitin-proteasome Background: The ubiquitin-proteasome system is system, which degrades polyubiquitin-tagged pro- a pathway in eukaryotic cells for degrading teins through the proteasomal machinery [2]. In the polyubiquitin-tagged proteins through the ubiquitin-proteasome system, an E3 ubiquitin ligase proteasomal machinery to control various cellular (hereinafter E3) selectively recognizes and binds to processes and maintain intracellular homeostasis. specific regions of the target substrate proteins. These In this system, the E3 ubiquitin ligase (hereinafter binding sites are called degrons [3]. E3) plays an important role in selectively It is important to identify the E3s and substrate recognizing and binding to specific regions of its proteins that interact with one another to character- substrate proteins. -

Bioinformatics: a Practical Guide to the Analysis of Genes and Proteins, Second Edition Andreas D
BIOINFORMATICS A Practical Guide to the Analysis of Genes and Proteins SECOND EDITION Andreas D. Baxevanis Genome Technology Branch National Human Genome Research Institute National Institutes of Health Bethesda, Maryland USA B. F. Francis Ouellette Centre for Molecular Medicine and Therapeutics Children’s and Women’s Health Centre of British Columbia University of British Columbia Vancouver, British Columbia Canada A JOHN WILEY & SONS, INC., PUBLICATION New York • Chichester • Weinheim • Brisbane • Singapore • Toronto BIOINFORMATICS SECOND EDITION METHODS OF BIOCHEMICAL ANALYSIS Volume 43 BIOINFORMATICS A Practical Guide to the Analysis of Genes and Proteins SECOND EDITION Andreas D. Baxevanis Genome Technology Branch National Human Genome Research Institute National Institutes of Health Bethesda, Maryland USA B. F. Francis Ouellette Centre for Molecular Medicine and Therapeutics Children’s and Women’s Health Centre of British Columbia University of British Columbia Vancouver, British Columbia Canada A JOHN WILEY & SONS, INC., PUBLICATION New York • Chichester • Weinheim • Brisbane • Singapore • Toronto Designations used by companies to distinguish their products are often claimed as trademarks. In all instances where John Wiley & Sons, Inc., is aware of a claim, the product names appear in initial capital or ALL CAPITAL LETTERS. Readers, however, should contact the appropriate companies for more complete information regarding trademarks and registration. Copyright ᭧ 2001 by John Wiley & Sons, Inc. All rights reserved. No part of this publication may be reproduced, stored in a retrieval system or transmitted in any form or by any means, electronic or mechanical, including uploading, downloading, printing, decompiling, recording or otherwise, except as permitted under Sections 107 or 108 of the 1976 United States Copyright Act, without the prior written permission of the Publisher. -

A Tool for Detecting Base Mis-Calls in Multiple Sequence Alignments by Semi-Automatic Chromatogram Inspection
ChromatoGate: A Tool for Detecting Base Mis-Calls in Multiple Sequence Alignments by Semi-Automatic Chromatogram Inspection Nikolaos Alachiotis Emmanouella Vogiatzi∗ Scientific Computing Group Institute of Marine Biology and Genetics HITS gGmbH HCMR Heidelberg, Germany Heraklion Crete, Greece [email protected] [email protected] Pavlos Pavlidis Alexandros Stamatakis Scientific Computing Group Scientific Computing Group HITS gGmbH HITS gGmbH Heidelberg, Germany Heidelberg, Germany [email protected] [email protected] ∗ Affiliated also with the Department of Genetics and Molecular Biology of the Democritian University of Thrace at Alexandroupolis, Greece. Corresponding author: Nikolaos Alachiotis Keywords: chromatograms, software, mis-calls Abstract Automated DNA sequencers generate chromatograms that contain raw sequencing data. They also generate data that translates the chromatograms into molecular sequences of A, C, G, T, or N (undetermined) characters. Since chromatogram translation programs frequently introduce errors, a manual inspection of the generated sequence data is required. As sequence numbers and lengths increase, visual inspection and manual correction of chromatograms and corresponding sequences on a per-peak and per-nucleotide basis becomes an error-prone, time-consuming, and tedious process. Here, we introduce ChromatoGate (CG), an open-source software that accelerates and partially automates the inspection of chromatograms and the detection of sequencing errors for bidirectional sequencing runs. To provide users full control over the error correction process, a fully automated error correction algorithm has not been implemented. Initially, the program scans a given multiple sequence alignment (MSA) for potential sequencing errors, assuming that each polymorphic site in the alignment may be attributed to a sequencing error with a certain probability. -

A Brief History of Sequence Logos
Biostatistics and Biometrics Open Access Journal ISSN: 2573-2633 Mini-Review Biostat Biometrics Open Acc J Volume 6 Issue 3 - April 2018 Copyright © All rights are reserved by Kushal K Dey DOI: 10.19080/BBOAJ.2018.06.555690 A Brief History of Sequence Logos Kushal K Dey* Department of Statistics, University of Chicago, USA Submission: February 12, 2018; Published: April 25, 2018 *Corresponding author: Kushal K Dey, Department of Statistics, University of Chicago, 5747 S Ellis Ave, Chicago, IL 60637, USA. Tel: 312-709- 0680; Email: Abstract For nearly three decades, sequence logo plots have served as the standard tool for graphical representation of aligned DNA, RNA and protein sequences. Over the years, a large number of packages and web applications have been developed for generating these logo plots and using them handling and the overall scope of these plots in biological applications and beyond. Here I attempt to review some popular tools for generating sequenceto identify logos, conserved with a patterns focus on in how sequences these plots called have motifs. evolved Also, over over time time, since we their have origin seen anda considerable how I view theupgrade future in for the these look, plots. flexibility of data Keywords : Graphical representation; Sequence logo plots; Standard tool; Motifs; Biological applications; Flexibility of data; DNA sequence data; Python library; Interdependencies; PLogo; Depletion of symbols; Alphanumeric strings; Visualizes pairwise; Oligonucleotide RNA sequence data; Visualize succinctly; Predictive power; Initial attempts; Widespread; Stylistic configurations; Multiple sequence alignment; Introduction based comparisons and predictions. In the next section, we The seeds of the origin of sequence logos were planted in review the modeling frameworks and functionalities of some of early 1980s when researchers, equipped with large amounts of these tools [5]. -
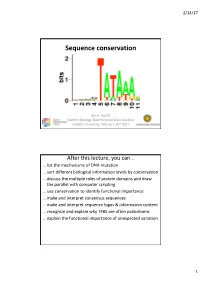
Sequence Conservation
2/14/17 Sequence conservation Bas E. Dutilh Systems Biology: Bioinformatic Data Analysis Utrecht University, FeBruary 16th 2017 After this lecture, you can… … list the mechanisms of DNA mutation … sort different biological information levels By conservation … discuss the multiple roles of protein domains and draw the parallel with computer scripting … use conservation to identify functional importance … make and interpret consensus sequences … make and interpret sequence logos & information content … recognize and explain why TFBS are often palindromic … explain the functional importance of unexpected variation 1 2/14/17 Evolution • Replication (copy) • Mutation (modify) • Selection (fitness) Time Mutations • Nucleotide suBstitutions – Replication error – Physical or chemical reaction • Insertions or deletions (indels) – Unequal crossing over during meiosis – Replication slippage • Inversions or rearrangements • Duplications of: – Partial or whole gene – Partial (polysomy) or whole chromosome (aneuploidy, polysomy) – Whole genome (polyploidy) • Horizontal gene transfer (HGT) – Conjugation (direct transfer between Bacteria) – Transformation by naturally competent Bacteria – Transduction by bacteriophages 2 2/14/17 Phenotypic/genotypic similarity • We exploit similarity to… …identify homology (shared ancestry) …determine evolutionary relationships …transfer functional information • Sequences (genotype) rarely converge • Functions (phenotype) can converge • Analogy • Homology – Similar function – Similar ancestry Low-complexity regions • -

Bioinformatics Manual Sequence Data Analysis with CLC Main Workbench
Introduction to Molecular Biology and Bioinformatics Workshop Biosciences eastern and central Africa - International Livestock Research Institute Hub (BecA-ILRI Hub), Nairobi, Kenya May 5-16, 2014 Bioinformatics Manual Sequence data analysis with CLC Main Workbench Written and compiled by Joyce Njuguna, Mark Wamalwa, Rob Skilton 1 Getting started with CLC Main Workbench A. Quality control of sequenced data B. Assembling sequences C. Nucleotide and protein sequence manipulation D. BLAST sequence search E. Aligning sequences F. Building phylogenetic trees Welcome to CLC Main Workbench -- a user-friendly sequence analysis software package for analysing Sanger sequencing data and for supporting your daily bioinformatics work. Definition Sequence: the order of nucleotide bases [or amino acids] in a DNA [or protein] molecule DNA Sequencing: Biochemical methods used to determine the order of nucleotide bases, adenine(A), guanine(G),cytosine(C)and thymine(T) in a DNA strand TIP CLC Main Workbench includes an extensive Help function, which can be found in the Help menu of the program’s Menu bar. The Help can also be shown by pressing F1. The help topics are sorted in a table of contents and the topics can be searched. Also, it is recommended that you view the Online presentations where a product specialist from CLC bio demonstrates the software. This is a very easy way to get started using the program. Read more about online presentations here: http://clcbio.com/presentation. A. Getting Started with CLC Main Workbench 1. Download the trial version of CLC Main Workbench 6.8.1 from the following URL: http://www.clcbio.com/products/clc-main-workbench-direct-download/ 2. -

A STAT Protein Domain That Determines DNA Sequence Recognition Suggests a Novel DNA-Binding Domain
Downloaded from genesdev.cshlp.org on September 25, 2021 - Published by Cold Spring Harbor Laboratory Press A STAT protein domain that determines DNA sequence recognition suggests a novel DNA-binding domain Curt M. Horvath, Zilong Wen, and James E. Darnell Jr. Laboratory of Molecular Cell Biology, The Rockefeller University, New York, New York 10021 Statl and Stat3 are two members of the ligand-activated transcription factor family that serve the dual functions of signal transducers and activators of transcription. Whereas the two proteins select very similar (not identical) optimum binding sites from random oligonucleotides, differences in their binding affinity were readily apparent with natural STAT-binding sites. To take advantage of these different affinities, chimeric Statl:Stat3 molecules were used to locate the amino acids that could discriminate a general binding site from a specific binding site. The amino acids between residues -400 and -500 of these -750-amino-acid-long proteins determine the DNA-binding site specificity. Mutations within this region result in Stat proteins that are activated normally by tyrosine phosphorylation and that dimerize but have greatly reduced DNA-binding affinities. [Key Words: STAT proteins; DNA binding; site selection] Received January 6, 1995; revised version accepted March 2, 1995. The STAT (signal transducers and activators if transcrip- Whereas oligonucleotides representing these selected se- tion) proteins have the dual purpose of, first, signal trans- quences exhibited slight binding preferences, the con- duction from ligand-activated receptor kinase com- sensus sites overlapped sufficiently to be recognized by plexes, followed by nuclear translocation and DNA bind- both factors. However, by screening different natural ing to activate transcription (Darnell et al. -

Bioinformatics I Sanger Sequence Analysis
Updated: October 2020 Lab 5: Bioinformatics I Sanger Sequence Analysis Project Guide The Wolbachia Project Page Contents 3 Activity at a Glance 4 Technical Overview 5-7 Activity: How to Analyze Sanger Sequences 8 DataBase Entry 9 Appendix I: Illustrated BLAST Alignment 10 Appendix II: Sanger Sequencing Quick Reference Content is made available under the Creative Commons Attribution-NonCommercial-No Derivatives International License. Contact ([email protected]) if you would like to make adaptations for distribution beyond the classroom. The Wolbachia Project: Discover the Microbes Within! was developed by a collaboration of scientists, educators, and outreach specialists. It is directed by the Bordenstein Lab at Vanderbilt University. https://www.vanderbilt.edu/wolbachiaproject 2 Activity at a Glance Goals To analyze and interpret the quality of Sanger sequences To generate a consensus DNA sequence for bioinformatics analyses Learning Objectives Upon completion of this activity, students will (i) understand the Sanger method of sequencing, also known as the chain-termination method; (ii) be able to interpret chromatograms; (iii) evaluate sequencing Quality Scores; and (iv) generate a consensus DNA sequence based on forward and reverse Sanger reactions. Prerequisite Skills While no computer programming skills are necessary to complete this work, prior exposure to personal computers and the Internet is assumed. Teaching Time: One class period Recommended Background Tutorials • DNA Learning Center Animation: Sanger Method of