BAZ1B in Nucleus Accumbens Regulates Reward-Related Behaviors in Response to Distinct Emotional Stimuli
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Functional Roles of Bromodomain Proteins in Cancer
cancers Review Functional Roles of Bromodomain Proteins in Cancer Samuel P. Boyson 1,2, Cong Gao 3, Kathleen Quinn 2,3, Joseph Boyd 3, Hana Paculova 3 , Seth Frietze 3,4,* and Karen C. Glass 1,2,4,* 1 Department of Pharmaceutical Sciences, Albany College of Pharmacy and Health Sciences, Colchester, VT 05446, USA; [email protected] 2 Department of Pharmacology, Larner College of Medicine, University of Vermont, Burlington, VT 05405, USA; [email protected] 3 Department of Biomedical and Health Sciences, University of Vermont, Burlington, VT 05405, USA; [email protected] (C.G.); [email protected] (J.B.); [email protected] (H.P.) 4 University of Vermont Cancer Center, Burlington, VT 05405, USA * Correspondence: [email protected] (S.F.); [email protected] (K.C.G.) Simple Summary: This review provides an in depth analysis of the role of bromodomain-containing proteins in cancer development. As readers of acetylated lysine on nucleosomal histones, bromod- omain proteins are poised to activate gene expression, and often promote cancer progression. We examined changes in gene expression patterns that are observed in bromodomain-containing proteins and associated with specific cancer types. We also mapped the protein–protein interaction network for the human bromodomain-containing proteins, discuss the cellular roles of these epigenetic regu- lators as part of nine different functional groups, and identify bromodomain-specific mechanisms in cancer development. Lastly, we summarize emerging strategies to target bromodomain proteins in cancer therapy, including those that may be essential for overcoming resistance. Overall, this review provides a timely discussion of the different mechanisms of bromodomain-containing pro- Citation: Boyson, S.P.; Gao, C.; teins in cancer, and an updated assessment of their utility as a therapeutic target for a variety of Quinn, K.; Boyd, J.; Paculova, H.; cancer subtypes. -

Cyclin D1 Is a Direct Transcriptional Target of GATA3 in Neuroblastoma Tumor Cells
Oncogene (2010) 29, 2739–2745 & 2010 Macmillan Publishers Limited All rights reserved 0950-9232/10 $32.00 www.nature.com/onc SHORT COMMUNICATION Cyclin D1 is a direct transcriptional target of GATA3 in neuroblastoma tumor cells JJ Molenaar1,2, ME Ebus1, J Koster1, E Santo1, D Geerts1, R Versteeg1 and HN Caron2 1Department of Human Genetics, Academic Medical Center, University of Amsterdam, Amsterdam, The Netherlands and 2Department of Pediatric Oncology, Emma Kinderziekenhuis, Academic Medical Center, University of Amsterdam, Amsterdam, The Netherlands Almost all neuroblastoma tumors express excess levels of 2000). Several checkpoints normally prevent premature Cyclin D1 (CCND1) compared to normal tissues and cell-cycle progression and cell division. The crucial G1 other tumor types. Only a small percentage of these entry point is controlled by the D-type Cyclins that can neuroblastoma tumors have high-level amplification of the activate CDK4/6 that in turn phosphorylate the pRb Cyclin D1 gene. The other neuroblastoma tumors have protein. This results in a release of the E2F transcription equally high Cyclin D1 expression without amplification. factor that causes transcriptional upregulation of Silencing of Cyclin D1 expression was previously found to numerous genes involved in further progression of the trigger differentiation of neuroblastoma cells. Over- cell cycle (Sherr, 1996). expression of Cyclin D1 is therefore one of the most Neuroblastomas are embryonal tumors that originate frequent mechanisms with a postulated function in neuro- from precursor cells of the sympathetic nervous system. blastoma pathogenesis. The cause for the Cyclin D1 This tumor has a very poor prognosis and despite the overexpression is unknown. -

Loss of ISWI Atpase SMARCA5 (SNF2H) in Acute Myeloid Leukemia Cells Inhibits Proliferation and Chromatid Cohesion
International Journal of Molecular Sciences Article Loss of ISWI ATPase SMARCA5 (SNF2H) in Acute Myeloid Leukemia Cells Inhibits Proliferation and Chromatid Cohesion 1, 1, 1 2,3,4 1 Tomas Zikmund y , Helena Paszekova y , Juraj Kokavec , Paul Kerbs , Shefali Thakur , Tereza Turkova 1, Petra Tauchmanova 1, Philipp A. Greif 2,3,4 and Tomas Stopka 1,* 1 Biocev, 1st Medical Faculty, Charles University, 25250 Vestec, Czech Republic; [email protected] (T.Z.); [email protected] (H.P.); [email protected] (J.K.); [email protected] (S.T.); [email protected] (T.T.); [email protected] (P.T.) 2 Department of Medicine III, University Hospital, LMU Munich, D-80539 Munich, Germany; [email protected] (P.K.); [email protected] (P.A.G.) 3 German Cancer Consortium (DKTK), partner site Munich, D-80336 Munich, Germany 4 German Cancer Research Center (DKFZ), D-69120 Heidelberg, Germany * Correspondence: [email protected]; Tel.: +420-32587-3001 These authors contributed equally. y Received: 26 February 2020; Accepted: 16 March 2020; Published: 18 March 2020 Abstract: ISWI chromatin remodeling ATPase SMARCA5 (SNF2H) is a well-known factor for its role in regulation of DNA access via nucleosome sliding and assembly. SMARCA5 transcriptionally inhibits the myeloid master regulator PU.1. Upregulation of SMARCA5 was previously observed in CD34+ hematopoietic progenitors of acute myeloid leukemia (AML) patients. Since high levels of SMARCA5 are necessary for intensive cell proliferation and cell cycle progression of developing hematopoietic stem and progenitor cells in mice, we reasoned that removal of SMARCA5 enzymatic activity could affect the cycling or undifferentiated state of leukemic progenitor-like clones. -

The Function and Evolution of C2H2 Zinc Finger Proteins and Transposons
The function and evolution of C2H2 zinc finger proteins and transposons by Laura Francesca Campitelli A thesis submitted in conformity with the requirements for the degree of Doctor of Philosophy Department of Molecular Genetics University of Toronto © Copyright by Laura Francesca Campitelli 2020 The function and evolution of C2H2 zinc finger proteins and transposons Laura Francesca Campitelli Doctor of Philosophy Department of Molecular Genetics University of Toronto 2020 Abstract Transcription factors (TFs) confer specificity to transcriptional regulation by binding specific DNA sequences and ultimately affecting the ability of RNA polymerase to transcribe a locus. The C2H2 zinc finger proteins (C2H2 ZFPs) are a TF class with the unique ability to diversify their DNA-binding specificities in a short evolutionary time. C2H2 ZFPs comprise the largest class of TFs in Mammalian genomes, including nearly half of all Human TFs (747/1,639). Positive selection on the DNA-binding specificities of C2H2 ZFPs is explained by an evolutionary arms race with endogenous retroelements (EREs; copy-and-paste transposable elements), where the C2H2 ZFPs containing a KRAB repressor domain (KZFPs; 344/747 Human C2H2 ZFPs) are thought to diversify to bind new EREs and repress deleterious transposition events. However, evidence of the gain and loss of KZFP binding sites on the ERE sequence is sparse due to poor resolution of ERE sequence evolution, despite the recent publication of binding preferences for 242/344 Human KZFPs. The goal of my doctoral work has been to characterize the Human C2H2 ZFPs, with specific interest in their evolutionary history, functional diversity, and coevolution with LINE EREs. -

Pathway and Network Analysis of Somatic Mutations Across Cancer
Network Analysis of Mutaons Across Cancer Types Ben Raphael Fabio Vandin, Max Leiserson, Hsin-Ta Wu Department of Computer Science Center for Computaonal Molecular Biology Significantly Mutated Genes Muta#on Matrix Stascal test Genes Paents Frequency Number Paents Study Num. Samples Num. SMG TCGA Ovarian (2011) 316 10 TCGA Breast (2012) 510 35 TCGA Colorectal (2012) 276 32 background mutaon rate (BMR), gene specific effects, etc. Significantly Mutated Genes à Pathways Stascal test Frequency Number Paents TCGA Colorectal (Nature 2012) TCGA Ovarian (Nature 2011) background mutaon rate (BMR), gene specific effects, etc. Advantages of Large Datasets Prior knowledge of groups of genes Genes Paents Known pathways Interac3on Network None Prior knowledge • Novel pathways or interac3ons between pathways (crosstalk) • Topology of interac3ons Two Algorithms Prior knowledge of groups of genes Genes Paents Known pathways Interac3on Network None Prior knowledge Number of Hypotheses HotNet subnetworks of Dendrix interac3on network Exclusive gene sets HotNet: Problem Defini3on Given: 1. Network G = (V, E) V = genes. E = interac3ons b/w genes 2. Binary mutaon matrix Genes = mutated = not mutated Paents Find: Connected subnetworks mutated in a significant number of paents. Subnetwork Properes Mutaon frequency/score AND network topology Frequency Number Paents • Moderate frequency/score • High frequency/score • Highly connected • Connected through high-degree node. Example: TP53 has 238 neighbors in HPRD network Mutated subnetworks: HotNet* Muta#on Matrix Human Interac#on Network Genes = mutated genes Paents (1) Muta#on à heat diffusion Extract “significantly hot” subnetworks Hot (2) Cold *F. Vandin, E. Upfal, and B. J. Raphael. J. Comp.Biol. (2011). Also RECOMB (2010). Stas3cal Test Muta#on Matrix Random Binary Matrix Genes Genes Paents Paents Xs = number of subnetworks ≥ s genes Two-stage mul-hypothesis test: Rigorously bound FDR. -

Supplementary Table 1
Supplementary Table 1. 492 genes are unique to 0 h post-heat timepoint. The name, p-value, fold change, location and family of each gene are indicated. Genes were filtered for an absolute value log2 ration 1.5 and a significance value of p ≤ 0.05. Symbol p-value Log Gene Name Location Family Ratio ABCA13 1.87E-02 3.292 ATP-binding cassette, sub-family unknown transporter A (ABC1), member 13 ABCB1 1.93E-02 −1.819 ATP-binding cassette, sub-family Plasma transporter B (MDR/TAP), member 1 Membrane ABCC3 2.83E-02 2.016 ATP-binding cassette, sub-family Plasma transporter C (CFTR/MRP), member 3 Membrane ABHD6 7.79E-03 −2.717 abhydrolase domain containing 6 Cytoplasm enzyme ACAT1 4.10E-02 3.009 acetyl-CoA acetyltransferase 1 Cytoplasm enzyme ACBD4 2.66E-03 1.722 acyl-CoA binding domain unknown other containing 4 ACSL5 1.86E-02 −2.876 acyl-CoA synthetase long-chain Cytoplasm enzyme family member 5 ADAM23 3.33E-02 −3.008 ADAM metallopeptidase domain Plasma peptidase 23 Membrane ADAM29 5.58E-03 3.463 ADAM metallopeptidase domain Plasma peptidase 29 Membrane ADAMTS17 2.67E-04 3.051 ADAM metallopeptidase with Extracellular other thrombospondin type 1 motif, 17 Space ADCYAP1R1 1.20E-02 1.848 adenylate cyclase activating Plasma G-protein polypeptide 1 (pituitary) receptor Membrane coupled type I receptor ADH6 (includes 4.02E-02 −1.845 alcohol dehydrogenase 6 (class Cytoplasm enzyme EG:130) V) AHSA2 1.54E-04 −1.6 AHA1, activator of heat shock unknown other 90kDa protein ATPase homolog 2 (yeast) AK5 3.32E-02 1.658 adenylate kinase 5 Cytoplasm kinase AK7 -

Effects of Targeting the Transcription Factors Ikaros and Aiolos on B Cell Activation and Differentiation in Systemic Lupus Erythematosus
Immunology and inflammation Lupus Sci Med: first published as 10.1136/lupus-2020-000445 on 16 March 2021. Downloaded from Effects of targeting the transcription factors Ikaros and Aiolos on B cell activation and differentiation in systemic lupus erythematosus Felice Rivellese ,1 Sotiria Manou- Stathopoulou,1 Daniele Mauro,1 Katriona Goldmann,1 Debasish Pyne,2 Ravindra Rajakariar,3 Patrick Gordon,4 Peter Schafer,5 Michele Bombardieri,1 Costantino Pitzalis,1 Myles J Lewis 1 To cite: Rivellese F, ABSTRACT Manou- Stathopoulou S, Objective To evaluate the effects of targeting Ikaros and Key messages Mauro D, et al. Effects of Aiolos by cereblon modulator iberdomide on the activation What is already known about this subject? targeting the transcription and differentiation of B- cells from patients with systemic factors Ikaros and Aiolos The transcription factors Ikaros and Aiolos, which lupus erythematosus (SLE). ► on B cell activation and are critical for B cell differentiation, are implicated in Methods CD19+ B- cells isolated from the peripheral differentiation in systemic systemic lupus erythematosus (SLE) pathogenesis. blood of patients with SLE (n=41) were cultured with lupus erythematosus. Targeting Ikaros and Aiolos using the cereblon mod- TLR7 ligand resiquimod ±IFNα together with iberdomide ► Lupus Science & Medicine ulator iberdomide has been proposed as a promising 2021;8:e000445. doi:10.1136/ or control from day 0 (n=16). Additionally, in vitro B- cell therapeutic agent. lupus-2020-000445 differentiation was induced by stimulation with IL-2/IL-10/ IL-15/CD40L/resiquimod with iberdomide or control, given What does this study add? at day 0 or at day 4. -

Chromatin Remodeling: a Complex Affair
News & Views Chromatin remodeling: a complex affair Nathan Gioacchini & Craig L Peterson ATP-dependent chromatin remodelers are spacing of nucleosomes during ISWI-cata- followed by mass spectrometry. They find multi-subunit enzymes that catalyze lyzed chromatin remodeling events (for a that each ISWI isoform co-purifies with all nucleosome dynamics essential for chro- recent review, see [2]). Mammals contain two of the known accessory subunits (BAZ1A/ mosomal functions, and their inactivation isoforms of the ISWI ATPase, encoded by two Acf1, BAZ1B/Wstf, BAZ2A/Tip5, CECR2, or dysregulation can lead to numerous related genes, Snf2L/SMARCA1 and Snf2H/ BPTF/NURF301, Rsf-1), and they identify diseases, including neuro-degenerative SMARCA5. In the mouse, Snf2H is essential the BAZ2B protein as a seventh accessory disorders and cancers. Each remodeler for early development and is expressed fairly factor for both ATPases. The co-purification contains a conserved ATPase “motor” ubiquitously, whereas Snf2L is expressed with all accessory subunits is not due to whose activity or targeting can be regu- highly in the brain and testes [3]. In contrast, contamination with the alternate ISWI lated by enzyme-specific, accessory sub- mice lacking Snf2L are viable and fertile, but isoform, as purifications were also carried units. The human ISWI subfamily of show delayed neurogenesis and an enhanced out with cell lines where one of the two remodelers has been defined as a group of forebrain hyper-cellularity phenotype [4]. isoforms was deleted by CRISPR/Cas9 edit- more than six different enzyme complexes Interestingly, these neural defects are not ing. Furthermore, interactions with each where one of two related ATPase subunits rescued by Snf2H overexpression, suggesting accessory subunit were observed in several (Snf2L/SMARCA1 and Snf2H/SMARCA5)is that each isoform has unique functions. -

Chemotherapeutic Drugs Inhibiting Topoisomerase 1 Activity Impede Cytokine-Induced and NF-Κb P65-Regulated Gene Expression
cancers Article Chemotherapeutic Drugs Inhibiting Topoisomerase 1 Activity Impede Cytokine-Induced and NF-κB p65-Regulated Gene Expression 1, 2 3 2 Tabea Riedlinger y, Marek Bartkuhn , Tobias Zimmermann , Sandra B. Hake , 4, 4, 5, 1, , Andrea Nist y, Thorsten Stiewe y, Michael Kracht y and M. Lienhard Schmitz * y 1 Institute of Biochemistry, Justus Liebig University, D-35392 Giessen, Germany; [email protected] 2 Institute for Genetics, Justus-Liebig University Giessen, 35392 Giessen, Germany; [email protected] (M.B.); [email protected] (S.B.H.) 3 Bioinformatics and Systems Biology, University of Giessen, Heinrich-Buff-Ring 58–62, 35392 Giessen, Germany; [email protected] 4 Genomics Core Facility and Institute of Molecular Oncology, Philipps University Marburg, D-35043 Marburg, Germany; [email protected] (A.N.); [email protected] (T.S.) 5 Rudolf-Buchheim-Institute of Pharmacology, Justus Liebig University, D-35392 Giessen, Germany; [email protected] * Correspondence: [email protected]; Tel.: +49-641-9947570; Fax: +49-641-9947589 Member of the German Center for Lung Research. y Received: 1 May 2019; Accepted: 20 June 2019; Published: 25 June 2019 Abstract: Inhibitors of DNA topoisomerase I (TOP1), an enzyme relieving torsional stress of DNA by generating transient single-strand breaks, are clinically used to treat ovarian, small cell lung and cervical cancer. As torsional stress is generated during transcription by progression of RNA polymerase II through the transcribed gene, we tested the effects of camptothecin and of the approved TOP1 inhibitors Topotecan and SN-38 on TNFα-induced gene expression. -

Xo PANEL DNA GENE LIST
xO PANEL DNA GENE LIST ~1700 gene comprehensive cancer panel enriched for clinically actionable genes with additional biologically relevant genes (at 400 -500x average coverage on tumor) Genes A-C Genes D-F Genes G-I Genes J-L AATK ATAD2B BTG1 CDH7 CREM DACH1 EPHA1 FES G6PC3 HGF IL18RAP JADE1 LMO1 ABCA1 ATF1 BTG2 CDK1 CRHR1 DACH2 EPHA2 FEV G6PD HIF1A IL1R1 JAK1 LMO2 ABCB1 ATM BTG3 CDK10 CRK DAXX EPHA3 FGF1 GAB1 HIF1AN IL1R2 JAK2 LMO7 ABCB11 ATR BTK CDK11A CRKL DBH EPHA4 FGF10 GAB2 HIST1H1E IL1RAP JAK3 LMTK2 ABCB4 ATRX BTRC CDK11B CRLF2 DCC EPHA5 FGF11 GABPA HIST1H3B IL20RA JARID2 LMTK3 ABCC1 AURKA BUB1 CDK12 CRTC1 DCUN1D1 EPHA6 FGF12 GALNT12 HIST1H4E IL20RB JAZF1 LPHN2 ABCC2 AURKB BUB1B CDK13 CRTC2 DCUN1D2 EPHA7 FGF13 GATA1 HLA-A IL21R JMJD1C LPHN3 ABCG1 AURKC BUB3 CDK14 CRTC3 DDB2 EPHA8 FGF14 GATA2 HLA-B IL22RA1 JMJD4 LPP ABCG2 AXIN1 C11orf30 CDK15 CSF1 DDIT3 EPHB1 FGF16 GATA3 HLF IL22RA2 JMJD6 LRP1B ABI1 AXIN2 CACNA1C CDK16 CSF1R DDR1 EPHB2 FGF17 GATA5 HLTF IL23R JMJD7 LRP5 ABL1 AXL CACNA1S CDK17 CSF2RA DDR2 EPHB3 FGF18 GATA6 HMGA1 IL2RA JMJD8 LRP6 ABL2 B2M CACNB2 CDK18 CSF2RB DDX3X EPHB4 FGF19 GDNF HMGA2 IL2RB JUN LRRK2 ACE BABAM1 CADM2 CDK19 CSF3R DDX5 EPHB6 FGF2 GFI1 HMGCR IL2RG JUNB LSM1 ACSL6 BACH1 CALR CDK2 CSK DDX6 EPOR FGF20 GFI1B HNF1A IL3 JUND LTK ACTA2 BACH2 CAMTA1 CDK20 CSNK1D DEK ERBB2 FGF21 GFRA4 HNF1B IL3RA JUP LYL1 ACTC1 BAG4 CAPRIN2 CDK3 CSNK1E DHFR ERBB3 FGF22 GGCX HNRNPA3 IL4R KAT2A LYN ACVR1 BAI3 CARD10 CDK4 CTCF DHH ERBB4 FGF23 GHR HOXA10 IL5RA KAT2B LZTR1 ACVR1B BAP1 CARD11 CDK5 CTCFL DIAPH1 ERCC1 FGF3 GID4 HOXA11 -
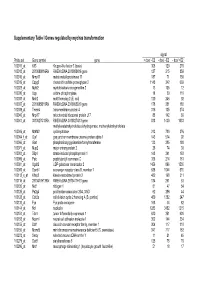
Supplementary Table I Genes Regulated by Myc/Ras Transformation
Supplementary Table I Genes regulated by myc/ras transformation signal Probe set Gene symbol gene + dox/ - E2 - dox/ - E2 - dox/ +E2 100011_at Klf3 Kruppel-like factor 3 (basic) 308 120 270 100013_at 2010008K16Rik RIKEN cDNA 2010008K16 gene 127 215 859 100016_at Mmp11 matrix metalloproteinase 11 187 71 150 100019_at Cspg2 chondroitin sulfate proteoglycan 2 1148 342 669 100023_at Mybl2 myeloblastosis oncogene-like 2 13 105 12 100030_at Upp uridine phosphorylase 18 39 110 100033_at Msh2 mutS homolog 2 (E. coli) 120 246 92 100037_at 2310005B10Rik RIKEN cDNA 2310005B10 gene 178 351 188 100039_at Tmem4 transmembrane protein 4 316 135 274 100040_at Mrpl17 mitochondrial ribosomal protein L17 88 142 89 100041_at 3010027G13Rik RIKEN cDNA 3010027G13 gene 818 1430 1033 methylenetetrahydrofolate dehydrogenase, methenyltetrahydrofolate 100046_at Mthfd2 cyclohydrolase 213 720 376 100064_f_at Gja1 gap junction membrane channel protein alpha 1 143 574 92 100066_at Gart phosphoribosylglycinamide formyltransferase 133 385 180 100071_at Mup2 major urinary protein 2 26 74 30 100081_at Stip1 stress-induced phosphoprotein 1 148 341 163 100089_at Ppic peptidylprolyl isomerase C 358 214 181 100091_at Ugalt2 UDP-galactose translocator 2 1456 896 1530 100095_at Scarb1 scavenger receptor class B, member 1 638 1044 570 100113_s_at Kifap3 kinesin-associated protein 3 482 168 311 100116_at 2810417H13Rik RIKEN cDNA 2810417H13 gene 134 261 53 100120_at Nid1 nidogen 1 81 47 54 100125_at Pa2g4 proliferation-associated 2G4, 38kD 62 286 44 100128_at Cdc2a cell division cycle 2 homolog A (S. pombe) 459 1382 247 100133_at Fyn Fyn proto-oncogene 168 44 82 100144_at Ncl nucleolin 1283 3452 1215 100151_at Tde1 tumor differentially expressed 1 620 351 620 100153_at Ncam1 neural cell adhesion molecule 1 302 144 234 100155_at Ddr1 discoidin domain receptor family, member 1 304 117 310 100156_at Mcmd5 mini chromosome maintenance deficient 5 (S. -

Identifying Proteins Bound to Native Mitotic ESC Chromosomes Reveals Chromatin Repressors Are Important for Compaction
ARTICLE https://doi.org/10.1038/s41467-020-17823-z OPEN Identifying proteins bound to native mitotic ESC chromosomes reveals chromatin repressors are important for compaction Dounia Djeghloul 1, Bhavik Patel2, Holger Kramer 3, Andrew Dimond 1, Chad Whilding4, Karen Brown1, Anne-Céline Kohler1, Amelie Feytout1, Nicolas Veland 1, James Elliott2, Tanmay A. M. Bharat 5, Abul K. Tarafder5, Jan Löwe 6, Bee L. Ng 7, Ya Guo1, Jacky Guy 8, Miles K. Huseyin 9, Robert J. Klose9, ✉ Matthias Merkenschlager 1 & Amanda G. Fisher 1 1234567890():,; Epigenetic information is transmitted from mother to daughter cells through mitosis. Here, to identify factors that might play a role in conveying epigenetic memory through cell division, we report on the isolation of unfixed, native chromosomes from metaphase-arrested cells using flow cytometry and perform LC-MS/MS to identify chromosome-bound proteins. A quantitative proteomic comparison between metaphase-arrested cell lysates and chromosome-sorted samples reveals a cohort of proteins that were significantly enriched on mitotic ESC chromosomes. These include pluripotency-associated transcription factors, repressive chromatin-modifiers such as PRC2 and DNA methyl-transferases, and proteins governing chromosome architecture. Deletion of PRC2, Dnmt1/3a/3b or Mecp2 in ESCs leads to an increase in the size of individual mitotic chromosomes, consistent with de- condensation. Similar results were obtained by the experimental cleavage of cohesin. Thus, we identify chromosome-bound factors in pluripotent stem cells during mitosis and reveal that PRC2, DNA methylation and Mecp2 are required to maintain chromosome compaction. 1 Lymphocyte Development Group, MRC London Institute of Medical Sciences, Imperial College London, Hammersmith Hospital Campus, Du Cane Road, London W12 0NN, UK.