CD8 T Cell Memory Transition and Maintenance of Functional Foxo1
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Role of Mitochondrial Ribosomal Protein S18-2 in Cancerogenesis and in Regulation of Stemness and Differentiation
From THE DEPARTMENT OF MICROBIOLOGY TUMOR AND CELL BIOLOGY (MTC) Karolinska Institutet, Stockholm, Sweden ROLE OF MITOCHONDRIAL RIBOSOMAL PROTEIN S18-2 IN CANCEROGENESIS AND IN REGULATION OF STEMNESS AND DIFFERENTIATION Muhammad Mushtaq Stockholm 2017 All previously published papers were reproduced with permission from the publisher. Published by Karolinska Institutet. Printed by E-Print AB 2017 © Muhammad Mushtaq, 2017 ISBN 978-91-7676-697-2 Role of Mitochondrial Ribosomal Protein S18-2 in Cancerogenesis and in Regulation of Stemness and Differentiation THESIS FOR DOCTORAL DEGREE (Ph.D.) By Muhammad Mushtaq Principal Supervisor: Faculty Opponent: Associate Professor Elena Kashuba Professor Pramod Kumar Srivastava Karolinska Institutet University of Connecticut Department of Microbiology Tumor and Cell Center for Immunotherapy of Cancer and Biology (MTC) Infectious Diseases Co-supervisor(s): Examination Board: Professor Sonia Lain Professor Ola Söderberg Karolinska Institutet Uppsala University Department of Microbiology Tumor and Cell Department of Immunology, Genetics and Biology (MTC) Pathology (IGP) Professor George Klein Professor Boris Zhivotovsky Karolinska Institutet Karolinska Institutet Department of Microbiology Tumor and Cell Institute of Environmental Medicine (IMM) Biology (MTC) Professor Lars-Gunnar Larsson Karolinska Institutet Department of Microbiology Tumor and Cell Biology (MTC) Dedicated to my parents ABSTRACT Mitochondria carry their own ribosomes (mitoribosomes) for the translation of mRNA encoded by mitochondrial DNA. The architecture of mitoribosomes is mainly composed of mitochondrial ribosomal proteins (MRPs), which are encoded by nuclear genomic DNA. Emerging experimental evidences reveal that several MRPs are multifunctional and they exhibit important extra-mitochondrial functions, such as involvement in apoptosis, protein biosynthesis and signal transduction. Dysregulations of the MRPs are associated with severe pathological conditions, including cancer. -

1 AGING Supplementary Table 2
SUPPLEMENTARY TABLES Supplementary Table 1. Details of the eight domain chains of KIAA0101. Serial IDENTITY MAX IN COMP- INTERFACE ID POSITION RESOLUTION EXPERIMENT TYPE number START STOP SCORE IDENTITY LEX WITH CAVITY A 4D2G_D 52 - 69 52 69 100 100 2.65 Å PCNA X-RAY DIFFRACTION √ B 4D2G_E 52 - 69 52 69 100 100 2.65 Å PCNA X-RAY DIFFRACTION √ C 6EHT_D 52 - 71 52 71 100 100 3.2Å PCNA X-RAY DIFFRACTION √ D 6EHT_E 52 - 71 52 71 100 100 3.2Å PCNA X-RAY DIFFRACTION √ E 6GWS_D 41-72 41 72 100 100 3.2Å PCNA X-RAY DIFFRACTION √ F 6GWS_E 41-72 41 72 100 100 2.9Å PCNA X-RAY DIFFRACTION √ G 6GWS_F 41-72 41 72 100 100 2.9Å PCNA X-RAY DIFFRACTION √ H 6IIW_B 2-11 2 11 100 100 1.699Å UHRF1 X-RAY DIFFRACTION √ www.aging-us.com 1 AGING Supplementary Table 2. Significantly enriched gene ontology (GO) annotations (cellular components) of KIAA0101 in lung adenocarcinoma (LinkedOmics). Leading Description FDR Leading Edge Gene EdgeNum RAD51, SPC25, CCNB1, BIRC5, NCAPG, ZWINT, MAD2L1, SKA3, NUF2, BUB1B, CENPA, SKA1, AURKB, NEK2, CENPW, HJURP, NDC80, CDCA5, NCAPH, BUB1, ZWILCH, CENPK, KIF2C, AURKA, CENPN, TOP2A, CENPM, PLK1, ERCC6L, CDT1, CHEK1, SPAG5, CENPH, condensed 66 0 SPC24, NUP37, BLM, CENPE, BUB3, CDK2, FANCD2, CENPO, CENPF, BRCA1, DSN1, chromosome MKI67, NCAPG2, H2AFX, HMGB2, SUV39H1, CBX3, TUBG1, KNTC1, PPP1CC, SMC2, BANF1, NCAPD2, SKA2, NUP107, BRCA2, NUP85, ITGB3BP, SYCE2, TOPBP1, DMC1, SMC4, INCENP. RAD51, OIP5, CDK1, SPC25, CCNB1, BIRC5, NCAPG, ZWINT, MAD2L1, SKA3, NUF2, BUB1B, CENPA, SKA1, AURKB, NEK2, ESCO2, CENPW, HJURP, TTK, NDC80, CDCA5, BUB1, ZWILCH, CENPK, KIF2C, AURKA, DSCC1, CENPN, CDCA8, CENPM, PLK1, MCM6, ERCC6L, CDT1, HELLS, CHEK1, SPAG5, CENPH, PCNA, SPC24, CENPI, NUP37, FEN1, chromosomal 94 0 CENPL, BLM, KIF18A, CENPE, MCM4, BUB3, SUV39H2, MCM2, CDK2, PIF1, DNA2, region CENPO, CENPF, CHEK2, DSN1, H2AFX, MCM7, SUV39H1, MTBP, CBX3, RECQL4, KNTC1, PPP1CC, CENPP, CENPQ, PTGES3, NCAPD2, DYNLL1, SKA2, HAT1, NUP107, MCM5, MCM3, MSH2, BRCA2, NUP85, SSB, ITGB3BP, DMC1, INCENP, THOC3, XPO1, APEX1, XRCC5, KIF22, DCLRE1A, SEH1L, XRCC3, NSMCE2, RAD21. -

NIH Public Access Author Manuscript Mitochondrion
NIH Public Access Author Manuscript Mitochondrion. Author manuscript; available in PMC 2009 June 1. NIH-PA Author ManuscriptPublished NIH-PA Author Manuscript in final edited NIH-PA Author Manuscript form as: Mitochondrion. 2008 June ; 8(3): 254±261. The effect of mutated mitochondrial ribosomal proteins S16 and S22 on the assembly of the small and large ribosomal subunits in human mitochondria Md. Emdadul Haque1, Domenick Grasso1,2, Chaya Miller3, Linda L Spremulli1, and Ann Saada3,# 1 Department of Chemistry, University of North Carolina at Chapel Hill, Chapel Hill, NC-27599-3290 3 Metabolic Disease Unit, Hadassah Medical Center, P.O.B. 12000, 91120 Jerusalem, Israel Abstract Mutations in mitochondrial small subunit ribosomal proteins MRPS16 or MRPS22 cause severe, fatal respiratory chain dysfunction due to impaired translation of mitochondrial mRNAs. The loss of either MRPS16 or MRPS22 was accompanied by the loss of most of another small subunit protein MRPS11. However, MRPS2 was reduced only about 2-fold in patient fibroblasts. This observation suggests that the small ribosomal subunit is only partially able to assemble in these patients. Two large subunit ribosomal proteins, MRPL13 and MRPL15, were present in substantial amounts suggesting that the large ribosomal subunit is still present despite a nonfunctional small subunit. Keywords Mitochondria; ribosome; ribosomal subunit; respiratory chain complexes; ribosomal proteins 1. Introduction The 16.5 kb human mitochondrial genome encodes 22 tRNAs, 2 rRNAs and thirteen polypeptides. These proteins, which are inserted into the inner membrane and assembled with nuclearly encoded polypeptides, are essential components of the mitochondrial respiratory chain complexes (MRC) I, III, IV and V. -

MRPS12 Rabbit Pab
Leader in Biomolecular Solutions for Life Science MRPS12 Rabbit pAb Catalog No.: A10573 Basic Information Background Catalog No. Mammalian mitochondrial ribosomal proteins are encoded by nuclear genes and help in A10573 protein synthesis within the mitochondrion. Mitochondrial ribosomes (mitoribosomes) consist of a small 28S subunit and a large 39S subunit. They have an estimated 75% Observed MW protein to rRNA composition compared to prokaryotic ribosomes, where this ratio is 14kDa reversed. Another difference between mammalian mitoribosomes and prokaryotic ribosomes is that the latter contain a 5S rRNA. Among different species, the proteins Calculated MW comprising the mitoribosome differ greatly in sequence, and sometimes in biochemical 15kDa properties, which prevents easy recognition by sequence homology. This gene encodes a 28S subunit protein that belongs to the ribosomal protein S12P family. The encoded Category protein is a key component of the ribosomal small subunit and controls the decoding fidelity and susceptibility to aminoglycoside antibiotics. The gene for mitochondrial Primary antibody seryl-tRNA synthetase is located upstream and adjacent to this gene, and both genes are possible candidates for the autosomal dominant deafness gene (DFNA4). Splice Applications variants that differ in the 5' UTR have been found for this gene; all three variants encode WB,IHC the same protein. Cross-Reactivity Human Recommended Dilutions Immunogen Information WB 1:1000 - 1:2000 Gene ID Swiss Prot 6183 O15235 IHC 1:50 - 1:200 Immunogen Recombinant fusion protein containing a sequence corresponding to amino acids 30-138 of human MRPS12 (NP_203527.1). Synonyms MRPS12;MPR-S12;MT-RPS12;RPMS12;RPS12;RPSM12 Contact Product Information www.abclonal.com Source Isotype Purification Rabbit IgG Affinity purification Storage Store at -20℃. -

MPV17L2 Is Required for Ribosome Assembly in Mitochondria Ilaria Dalla Rosa1,†, Romina Durigon1,†, Sarah F
8500–8515 Nucleic Acids Research, 2014, Vol. 42, No. 13 Published online 19 June 2014 doi: 10.1093/nar/gku513 MPV17L2 is required for ribosome assembly in mitochondria Ilaria Dalla Rosa1,†, Romina Durigon1,†, Sarah F. Pearce2,†, Joanna Rorbach2, Elizabeth M.A. Hirst1, Sara Vidoni2, Aurelio Reyes2, Gloria Brea-Calvo2, Michal Minczuk2, Michael W. Woellhaf3, Johannes M. Herrmann3, Martijn A. Huynen4,IanJ.Holt1 and Antonella Spinazzola1,* 1MRC National Institute for Medical Research, Mill Hill, London NW7 1AA, UK, 2MRC Mitochondrial Biology Unit, Wellcome Trust-MRC Building, Hills Road, Cambridge CB2 0XY, UK, 3Cell Biology, University of Kaiserslautern, 67663 Kaiserslautern, Germany and 4Centre for Molecular and Biomolecular Informatics, Radboud University Medical Centre, Geert Grooteplein Zuid 26–28, 6525 GA Nijmegen, Netherlands Received January 15, 2014; Revised May 7, 2014; Accepted May 23, 2014 ABSTRACT INTRODUCTION MPV17 is a mitochondrial protein of unknown func- The mammalian mitochondrial proteome comprises 1500 tion, and mutations in MPV17 are associated with or more gene products. The deoxyribonucleic acid (DNA) mitochondrial deoxyribonucleic acid (DNA) mainte- inside mitochondria DNA (mtDNA) contributes only 13 ∼ nance disorders. Here we investigated its most sim- of these proteins, and they make up 20% of the subunits ilar relative, MPV17L2, which is also annotated as of the oxidative phosphorylation (OXPHOS) system, which produces much of the cells energy. All the other proteins a mitochondrial protein. Mitochondrial fractionation -

Integrative Genomic Analysis Predicts Causative Cis-Regulatory Mechanisms of the Breast Cancer-Associated Genetic Variant Rs4415084
Author Manuscript Published OnlineFirst on January 19, 2018; DOI: 10.1158/0008-5472.CAN-17-3486 Author manuscripts have been peer reviewed and accepted for publication but have not yet been edited. Cancer Research Mathematical Oncology Title: Integrative genomic analysis predicts causative cis-regulatory mechanisms of the breast cancer-associated genetic variant rs4415084 Authors and affiliations: Yi Zhang1,2*, Mohith Manjunath2*, Shilu Zhang3, Deborah Chasman3, Sushmita Roy3,4, and Jun S. Song2,5 *Equal contribution 1Department of Bioengineering, 2Carl R. Woese Institute for Genomic Biology, University of Illinois at Urbana-Champaign, Urbana, IL 61801, USA. 3Wisconsin Institute for Discovery, 4Department of Biostatistics and Medical Informatics, University of Wisconsin–Madison, Madison, WI 53792, USA. 5Department of Physics, University of Illinois at Urbana-Champaign, Urbana, IL 61801, USA. Running title: Regulatory genomics of GWAS SNPs Abbreviations: GWAS - genome-wide association study; eQTL - expression quantitative trait loci; ASE - allele-specific expression; TF - transcription factor; LD - linkage disequilibrium; FPKM - fragments per kilobase of transcript per million mapped reads; LCASE - local chromosome allele-specific expression; DHS - DNase I hypersensitive sites; PWM - position weight matrix; TCGA - The Cancer Genome Atlas; ER+ - estrogen receptor positive; TAD - topologically associated domain; MAF - minor allele frequency; RPKM - reads per kilobase of transcript per million mapped reads; SNP - single nucleotide polymorphism; MAPQ - mapping quality; ChIP-seq - chromatin immunoprecipitation sequencing; ASB - allele-specific binding; ncRNA - non-coding RNA; TSS - transcription start site; DNase-seq - DNase I 1 Downloaded from cancerres.aacrjournals.org on September 29, 2021. © 2018 American Association for Cancer Research. Author Manuscript Published OnlineFirst on January 19, 2018; DOI: 10.1158/0008-5472.CAN-17-3486 Author manuscripts have been peer reviewed and accepted for publication but have not yet been edited. -

Using Edsurvey to Analyze NCES Data: an Illustration of Analyzing NAEP Primer
Using EdSurvey to Analyze NCES Data: An Illustration of Analyzing NAEP Primer Developed by Michael Lee, Paul Bailey, Ahmad Emad, Ting Zhang, Trang Nguyen, and Jiao Yu*† February 21, 2020 Overview of the EdSurvey Package National Assessment of Educational Progress (NAEP) datasets from the National Center for Education Statistics (NCES) require special statistical methods to analyze. Because of their scope and complexity, the EdSurvey package gives users functions to perform analyses that account for both complex sample survey designs and the use of plausible values. The EdSurvey package also seamlessly takes advantage of the LaF package to read in data only when required for an analysis. Users with computers that have insuÿcient memory to read in entire NAEP datasets can still do analyses without having to write special code to read in just the appropriate variables. This situation is addressed directly in the EdSurvey package—behind the scenes and without any special tuning by the user. Vignette Outline This vignette will describe the basics of using the EdSurvey package for analyzing NAEP data as follows. • Notes – Additional resources – Vignette notation – Software requirements • Setting up the environment for analyzing NCES data – Installing and loading EdSurvey – Philosophy of Conducting Analyses Using the EdSurvey Package – Downloading data – Reading in data – Getting to know the data format – Removing special values • Explore Variable Distributions with summary2 • Subsetting the data • Retrieving data for further manipulation with getData *This publication was prepared for NCES under Contract No. ED-IES-12-D-0002 with the American Institutes for Research. Mention of trade names, commercial products, or organizations does not imply endorsement by the U.S. -
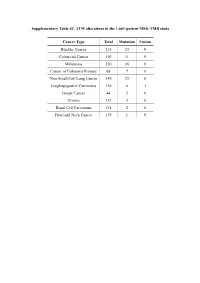
Supplementary Table S1. ATM Alterations in the 1,661-Patient MSK-TMB Study
Supplementary Table S1. ATM alterations in the 1,661-patient MSK-TMB study Cancer Type Total Mutation Fusion Bladder Cancer 215 23 0 Colorectal Cancer 110 11 0 Melanoma 320 26 0 Cancer of Unknown Primary 88 7 0 Non-Small Cell Lung Cancer 350 23 0 Esophagogastric Carcinoma 126 6 1 Breast Cancer 44 2 0 Glioma 117 3 0 Renal Cell Carcinoma 151 2 0 Head and Neck Cancer 139 1 0 Supplementary Table S2. ATM alternations in the 10,945-patient MSK-IMPACT study Cancer Type Total Mutation Deep deletion Amplification Multi-alterations Fusion Small Bowl Cancer 35 7 0 0 0 0 Skin Cancer, Non-Melaoma 148 19 1 0 0 0 Bladder Cancer 423 45 2 0 0 0 Endometrial Cancer 218 24 0 0 0 0 Hepatobiliary Cancer 355 27 0 0 0 0 Colorectal Cancer 1007 75 0 0 0 1 Mature B-Cell Neoplasms 134 8 0 0 2 0 Non-Small Cell Lung Cancer 1668 120 0 2 1 0 Melanoma 365 23 0 0 0 0 Appendiceal Cancer 79 4 0 0 0 0 Small Cell Lung Cancer 82 4 0 0 0 0 Prostate Cancer 717 25 7 1 0 0 Histiocytosis 22 1 0 0 0 0 Salivary Gland Cancer 114 5 0 0 0 0 Thyroid Cancer 231 10 0 0 0 0 Cancer of Unknown Primary 186 8 0 0 0 0 Breast Cancer 1324 48 3 2 0 0 Adrenocortical Carcinoma 25 1 0 0 0 0 Mature T and NK Neoplasms 29 1 0 0 0 0 Renal Cell Carcinoma 361 12 0 0 0 0 Head and Neck Cancer 186 14 0 0 0 1 Pancreatic Cancer 502 69 7 2 0 0 Glioma 553 14 1 0 0 0 Soft Tissue Sarcoma 443 13 1 0 0 0 Esophagogastric Cancer 341 8 1 0 0 0 Peripheral Nervous System 80 0 2 0 0 0 Germ Cell Tumor 288 2 0 1 0 1 Ovarian Can 244 2 1 0 0 0 Uterine Sarcoma 93 1 0 0 0 0 Mesothelioma 107 1 0 0 0 0 Bone Cancer 134 1 0 0 0 0 Gastrointestinal Stromal Ca 137 0 0 0 0 1 Supplementary Table S3. -

MRPS22 Rabbit Pab
Leader in Biomolecular Solutions for Life Science MRPS22 Rabbit pAb Catalog No.: A12597 Basic Information Background Catalog No. Mammalian mitochondrial ribosomal proteins are encoded by nuclear genes and help in A12597 protein synthesis within the mitochondrion. Mitochondrial ribosomes (mitoribosomes) consist of a small 28S subunit and a large 39S subunit. They have an estimated 75% Observed MW protein to rRNA composition compared to prokaryotic ribosomes, where this ratio is 41kDa reversed. Another difference between mammalian mitoribosomes and prokaryotic ribosomes is that the latter contain a 5S rRNA. Among different species, the proteins Calculated MW comprising the mitoribosome differ greatly in sequence, and sometimes in biochemical 18kDa/41kDa properties, which prevents easy recognition by sequence homology. This gene encodes a 28S subunit protein that does not seem to have a counterpart in prokaryotic and Category fungal-mitochondrial ribosomes. This gene lies telomeric of and is transcribed in the opposite direction from the forkhead box L2 gene. A pseudogene corresponding to this Primary antibody gene is found on chromosome Xq. Applications WB, IHC Cross-Reactivity Human, Mouse, Rat Recommended Dilutions Immunogen Information WB 1:500 - 1:2000 Gene ID Swiss Prot 56945 P82650 IHC 1:100 - 1:200 Immunogen Recombinant fusion protein containing a sequence corresponding to amino acids 1-360 of human MRPS22 (NP_064576.1). Synonyms MRPS22;C3orf5;COXPD5;GIBT;GK002;MRP-S22;RPMS22 Contact Product Information www.abclonal.com Source Isotype Purification Rabbit IgG Affinity purification Storage Store at -20℃. Avoid freeze / thaw cycles. Buffer: PBS with 0.02% sodium azide,50% glycerol,pH7.3. Validation Data Western blot analysis of extracts of various cell lines, using MRPS22 Antibody (A12597) at 1:1000 dilution. -

The RNA Helicase DHX30 Coordinates Cytoplasmic Translation and Mitochondrial Function Contributing to Cancer Cell Survival
bioRxiv preprint doi: https://doi.org/10.1101/2020.07.13.196709; this version posted July 13, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. The RNA helicase DHX30 coordinates cytoplasmic translation and mitochondrial function contributing to cancer cell survival Bartolomeo Bosco1, Annalisa Rossi1, Dario Rizzotto1, Sebastiano Giorgetta1, Alicia Perzolli1, Francesco Bonollo1, Angeline Gaucherot2, Frédéric Catez2, Jean-Jacques Diaz2, Erik Dassi1#, Alberto Inga1# 1Department of Cellular, Computational and Integrative Biology, CIBIO, University of Trento, via Sommarive 9, 38123, Trento, Italy 2Inserm U1052, CNRS UMR5286 Centre de Recherche en Cancérologie de Lyon, F-69000 Lyon, France. Centre Léon Bérard, F-69008 Lyon, France, Université de Lyon 1, F-69000 Lyon, France #corresponding authors: [email protected] [email protected] Running title: DHX30 controls global translation Keywords: DHX30, translation efficiency, polysomal profiling, mitoribosome, ribosome biogenesis bioRxiv preprint doi: https://doi.org/10.1101/2020.07.13.196709; this version posted July 13, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Abstract DHX30 was recently identified as a critical element of an RBP complex participating in translational control of mRNAs containing a target 3’UTR cis element, in the context of p53- dependent apoptosis. Here we show that DHX30 exhibits a more general, housekeeping function that comprises a combination of activities exerted by an isoform expressed in the cytoplasm and one, more abundant, localized in the mitochondria. -

Microarray Comparative Genomic Hybridization Analysis of 59 Patients with Schizophrenia
J Hum Genet (2008) 53:914–919 DOI 10.1007/s10038-008-0327-6 ORIGINAL ARTICLE Microarray comparative genomic hybridization analysis of 59 patients with schizophrenia Takeshi Mizuguchi Æ Ryota Hashimoto Æ Masanari Itokawa Æ Akira Sano Æ Osamu Shimokawa Æ Yukiko Yoshimura Æ Naoki Harada Æ Noriko Miyake Æ Akira Nishimura Æ Hirotomo Saitsu Æ Nadiya Sosonkina Æ Norio Niikawa Æ Hiroshi Kunugi Æ Naomichi Matsumoto Received: 18 April 2008 / Accepted: 8 July 2008 / Published online: 7 August 2008 Ó The Japan Society of Human Genetics and Springer 2008 Abstract Schizophrenia is a common psychiatric disor- that genome-wide copy number survey should be consid- der with a strong genetic contribution. Disease-associated ered in genetic studies of schizophrenia. chromosomal abnormalities in this condition may provide important clues, such as DISC1. In this study, 59 schizo- Keywords Schizophrenia Á Chromosomal abnormality Á phrenia patients were analyzed by microarray comparative Array comparative genomic hybridization Á genomic hybridization (CGH) using custom bacterial arti- Copy number variation ficial chromosome (BAC) microarray (4,219 BACs with 0.7-Mb resolution). Chromosomal abnormalities were found in six patients (10%): 46,XY,der(13)t(12;13)(p12.1; Introduction p11).ish del(5)(p11p12); 46,XY, ish del(17)(p12p12); 46,XX.ish dup(11)(p13p13); and 46,X,idic(Y)(q11.2); and Schizophrenia is a common psychiatric disorder involving in two cases, mos 45,X/46XX. Autosomal abnormalities in approximately 1% of the population worldwide. Family, three cases are likely to be pathogenic, and sex chromo- twin, and adoption studies suggest genetic factors con- some abnormalities in three follow previous findings. -

Transcriptomic and Proteomic Landscape of Mitochondrial
TOOLS AND RESOURCES Transcriptomic and proteomic landscape of mitochondrial dysfunction reveals secondary coenzyme Q deficiency in mammals Inge Ku¨ hl1,2†*, Maria Miranda1†, Ilian Atanassov3, Irina Kuznetsova4,5, Yvonne Hinze3, Arnaud Mourier6, Aleksandra Filipovska4,5, Nils-Go¨ ran Larsson1,7* 1Department of Mitochondrial Biology, Max Planck Institute for Biology of Ageing, Cologne, Germany; 2Department of Cell Biology, Institute of Integrative Biology of the Cell (I2BC) UMR9198, CEA, CNRS, Univ. Paris-Sud, Universite´ Paris-Saclay, Gif- sur-Yvette, France; 3Proteomics Core Facility, Max Planck Institute for Biology of Ageing, Cologne, Germany; 4Harry Perkins Institute of Medical Research, The University of Western Australia, Nedlands, Australia; 5School of Molecular Sciences, The University of Western Australia, Crawley, Australia; 6The Centre National de la Recherche Scientifique, Institut de Biochimie et Ge´ne´tique Cellulaires, Universite´ de Bordeaux, Bordeaux, France; 7Department of Medical Biochemistry and Biophysics, Karolinska Institutet, Stockholm, Sweden Abstract Dysfunction of the oxidative phosphorylation (OXPHOS) system is a major cause of human disease and the cellular consequences are highly complex. Here, we present comparative *For correspondence: analyses of mitochondrial proteomes, cellular transcriptomes and targeted metabolomics of five [email protected] knockout mouse strains deficient in essential factors required for mitochondrial DNA gene (IKu¨ ); expression, leading to OXPHOS dysfunction. Moreover,