Restoration of Sp4 in Forebrain Gabaergic Neurons Rescues Hypersensitivity to Ketamine in Sp4 Hypomorphic Mice
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

Supplemental Materials ZNF281 Enhances Cardiac Reprogramming
Supplemental Materials ZNF281 enhances cardiac reprogramming by modulating cardiac and inflammatory gene expression Huanyu Zhou, Maria Gabriela Morales, Hisayuki Hashimoto, Matthew E. Dickson, Kunhua Song, Wenduo Ye, Min S. Kim, Hanspeter Niederstrasser, Zhaoning Wang, Beibei Chen, Bruce A. Posner, Rhonda Bassel-Duby and Eric N. Olson Supplemental Table 1; related to Figure 1. Supplemental Table 2; related to Figure 1. Supplemental Table 3; related to the “quantitative mRNA measurement” in Materials and Methods section. Supplemental Table 4; related to the “ChIP-seq, gene ontology and pathway analysis” and “RNA-seq” and gene ontology analysis” in Materials and Methods section. Supplemental Figure S1; related to Figure 1. Supplemental Figure S2; related to Figure 2. Supplemental Figure S3; related to Figure 3. Supplemental Figure S4; related to Figure 4. Supplemental Figure S5; related to Figure 6. Supplemental Table S1. Genes included in human retroviral ORF cDNA library. Gene Gene Gene Gene Gene Gene Gene Gene Symbol Symbol Symbol Symbol Symbol Symbol Symbol Symbol AATF BMP8A CEBPE CTNNB1 ESR2 GDF3 HOXA5 IL17D ADIPOQ BRPF1 CEBPG CUX1 ESRRA GDF6 HOXA6 IL17F ADNP BRPF3 CERS1 CX3CL1 ETS1 GIN1 HOXA7 IL18 AEBP1 BUD31 CERS2 CXCL10 ETS2 GLIS3 HOXB1 IL19 AFF4 C17ORF77 CERS4 CXCL11 ETV3 GMEB1 HOXB13 IL1A AHR C1QTNF4 CFL2 CXCL12 ETV7 GPBP1 HOXB5 IL1B AIMP1 C21ORF66 CHIA CXCL13 FAM3B GPER HOXB6 IL1F3 ALS2CR8 CBFA2T2 CIR1 CXCL14 FAM3D GPI HOXB7 IL1F5 ALX1 CBFA2T3 CITED1 CXCL16 FASLG GREM1 HOXB9 IL1F6 ARGFX CBFB CITED2 CXCL3 FBLN1 GREM2 HOXC4 IL1F7 -

SUPPLEMENTARY MATERIAL Bone Morphogenetic Protein 4 Promotes
www.intjdevbiol.com doi: 10.1387/ijdb.160040mk SUPPLEMENTARY MATERIAL corresponding to: Bone morphogenetic protein 4 promotes craniofacial neural crest induction from human pluripotent stem cells SUMIYO MIMURA, MIKA SUGA, KAORI OKADA, MASAKI KINEHARA, HIROKI NIKAWA and MIHO K. FURUE* *Address correspondence to: Miho Kusuda Furue. Laboratory of Stem Cell Cultures, National Institutes of Biomedical Innovation, Health and Nutrition, 7-6-8, Saito-Asagi, Ibaraki, Osaka 567-0085, Japan. Tel: 81-72-641-9819. Fax: 81-72-641-9812. E-mail: [email protected] Full text for this paper is available at: http://dx.doi.org/10.1387/ijdb.160040mk TABLE S1 PRIMER LIST FOR QRT-PCR Gene forward reverse AP2α AATTTCTCAACCGACAACATT ATCTGTTTTGTAGCCAGGAGC CDX2 CTGGAGCTGGAGAAGGAGTTTC ATTTTAACCTGCCTCTCAGAGAGC DLX1 AGTTTGCAGTTGCAGGCTTT CCCTGCTTCATCAGCTTCTT FOXD3 CAGCGGTTCGGCGGGAGG TGAGTGAGAGGTTGTGGCGGATG GAPDH CAAAGTTGTCATGGATGACC CCATGGAGAAGGCTGGGG MSX1 GGATCAGACTTCGGAGAGTGAACT GCCTTCCCTTTAACCCTCACA NANOG TGAACCTCAGCTACAAACAG TGGTGGTAGGAAGAGTAAAG OCT4 GACAGGGGGAGGGGAGGAGCTAGG CTTCCCTCCAACCAGTTGCCCCAAA PAX3 TTGCAATGGCCTCTCAC AGGGGAGAGCGCGTAATC PAX6 GTCCATCTTTGCTTGGGAAA TAGCCAGGTTGCGAAGAACT p75 TCATCCCTGTCTATTGCTCCA TGTTCTGCTTGCAGCTGTTC SOX9 AATGGAGCAGCGAAATCAAC CAGAGAGATTTAGCACACTGATC SOX10 GACCAGTACCCGCACCTG CGCTTGTCACTTTCGTTCAG Suppl. Fig. S1. Comparison of the gene expression profiles of the ES cells and the cells induced by NC and NC-B condition. Scatter plots compares the normalized expression of every gene on the array (refer to Table S3). The central line -

Exclusion of Dlx5/6 Expression from the Distal-Most Mandibular Arches Enables BMP-Mediated Specification of the Distal Cap
Exclusion of Dlx5/6 expression from the distal-most mandibular arches enables BMP-mediated specification of the distal cap Joshua W. Vincentza, Jose J. Casasnovasa, Ralston M. Barnesb, Jianwen Quec, David E. Clouthierd, Jun Wange, and Anthony B. Firullia,1 aRiley Heart Research Center, Herman B. Wells Center for Pediatric Research Division of Pediatric Cardiology, Departments of Anatomy, Biochemistry, and Medical and Molecular Genetics, Indiana University Medical School, Indianapolis, IN 46202; bBiologics Discovery California, Bristol-Myers Squibb, Redwood City, CA 94063; cDepartment of Medicine, Columbia University Medical Center, New York, NY 10032; dDepartment of Craniofacial Biology, University of Colorado Anschutz Medical Campus, Aurora, CO 80108; and eDepartment of Molecular Physiology and Biophysics, Texas Heart Institute, Baylor College of Medicine, Houston, TX 77030 Edited by Clifford J. Tabin, Harvard Medical School, Boston, MA, and approved May 26, 2016 (received for review March 8, 2016) Cranial neural crest cells (crNCCs) migrate from the neural tube to Smads, H2, and GATA2/3 provide positive transcriptional inputs the pharyngeal arches (PAs) of the developing embryo and, sub- that serve to counteract the activity of Dlx proteins, thereby re- sequently, differentiate into bone and connective tissue to form the lieving EDN1-meditated repression. Together, these findings in- mandible. Within the PAs, crNCCs respond to local signaling cues to tegrate the communication between BMP and EDN1 signaling partition into the proximo-distally oriented subdomains that convey that establishes the distal cap of the mandible. positional information to these developing tissues. Here, we show that the distal-most of these subdomains, the distal cap, is marked Results by expression of the transcription factor Hand1 (H1) and gives rise to The H1Cre Lineage Gives Rise to the Lower Incisors, in a HAND Factor- the ectomesenchymal derivatives of the lower incisors. -

Dlx5 and Dlx6 Expression in Gabaergic Neurons Controls Behavior, Metabolism, Healthy Aging and Lifespan
Dlx5 and Dlx6 expression in GABAergic neurons controls behavior, metabolism, healthy aging and lifespan. Camille de Lombares, Eglantine Heude, Gladys Alfama, Anastasia Fontaine, Rim Hassouna, Cecile Vernochet, Fabrice de Chaumont, Christophe Olivo-Marin, Elodie Ey, Sebastien Parnaudeau, et al. To cite this version: Camille de Lombares, Eglantine Heude, Gladys Alfama, Anastasia Fontaine, Rim Hassouna, et al.. Dlx5 and Dlx6 expression in GABAergic neurons controls behavior, metabolism, healthy aging and lifespan.. Aging, Impact Journals, 2019, 11 (17), pp.6638-6656. 10.18632/aging.102141. mnhn- 02295464 HAL Id: mnhn-02295464 https://hal-mnhn.archives-ouvertes.fr/mnhn-02295464 Submitted on 4 Feb 2020 HAL is a multi-disciplinary open access L’archive ouverte pluridisciplinaire HAL, est archive for the deposit and dissemination of sci- destinée au dépôt et à la diffusion de documents entific research documents, whether they are pub- scientifiques de niveau recherche, publiés ou non, lished or not. The documents may come from émanant des établissements d’enseignement et de teaching and research institutions in France or recherche français ou étrangers, des laboratoires abroad, or from public or private research centers. publics ou privés. Distributed under a Creative Commons Attribution| 4.0 International License www.aging-us.com AGING 2019, Vol. 11, No. 17 Research Paper Dlx5 and Dlx6 expression in GABAergic neurons controls behavior, metabolism, healthy aging and lifespan Camille de Lombares1, Eglantine Heude1, Gladys Alfama1, Anastasia Fontaine1, -

DLX Genes: Roles in Development and Cancer
cancers Review DLX Genes: Roles in Development and Cancer Yinfei Tan 1,* and Joseph R. Testa 1,2,* 1 Genomics Facility, Fox Chase Cancer Center, Philadelphia, PA 19111, USA 2 Cancer Signaling and Epigenetics Program, Fox Chase Cancer Center, Philadelphia, PA 19111, USA * Correspondence: [email protected] (Y.T.); [email protected] (J.R.T.) Simple Summary: DLX homeobox family genes encode transcription factors that have indispensable roles in embryonic and postnatal development. These genes are critically linked to the morphogene- sis of craniofacial structures, branchial arches, forebrain, and sensory organs. DLX genes are also involved in postnatal homeostasis, particularly hematopoiesis and, when dysregulated, oncogen- esis. DLX1/2, DLX3/4, and DLX5/6 exist as bigenes on different chromosomes, sharing intergenic enhancers between gene pairs, which allows orchestrated spatiotemporal expression. Genomic alterations of human DLX gene enhancers or coding sequences result in congenital disorders such as split-hand/foot malformation. Aberrant postnatal expression of DLX genes is associated with hematological malignancies, including leukemias and lymphomas. In several mouse models of T-cell lymphoma, Dlx5 has been shown to act as an oncogene by cooperating with activated Akt, Notch1/3, and/or Wnt to drive tumor formation. In humans, DLX5 is aberrantly expressed in lung and ovarian carcinomas and holds promise as a therapeutic target. Abstract: Homeobox genes control body patterning and cell-fate decisions during development. The homeobox genes consist of many families, only some of which have been investigated regarding a possible role in tumorigenesis. Dysregulation of HOX family genes have been widely implicated in cancer etiology. -

Transcriptional Profiling of Mesc-Derived Tendon And
ARTICLE https://doi.org/10.1038/s41467-021-24535-5 OPEN Transcriptional profiling of mESC-derived tendon and fibrocartilage cell fate switch ✉ Deepak A. Kaji1, Angela M. Montero1, Roosheel Patel2 & Alice H. Huang 1 The transcriptional regulators underlying induction and differentiation of dense connective tissues such as tendon and related fibrocartilaginous tissues (meniscus and annulus fibrosus) remain largely unknown. Using an iterative approach informed by developmental cues and 1234567890():,; single cell RNA sequencing (scRNA-seq), we establish directed differentiation models to generate tendon and fibrocartilage cells from mouse embryonic stem cells (mESCs) by activation of TGFβ and hedgehog pathways, achieving 90% induction efficiency. Transcrip- tional signatures of the mESC-derived cells recapitulate embryonic tendon and fibrocartilage signatures from the mouse tail. scRNA-seq further identify retinoic acid signaling as a critical regulator of cell fate switch between TGFβ-induced tendon and fibrocartilage lineages. Tra- jectory analysis by RNA sequencing define transcriptional modules underlying tendon and fibrocartilage fate induction and identify molecules associated with lineage-specific differ- entiation. Finally, we successfully generate 3-dimensional engineered tissues using these differentiation protocols and show activation of mechanotransduction markers with dynamic tensile loading. These findings provide a serum-free approach to generate tendon and fibrocartilage cells and tissues at high efficiency for modeling development and -
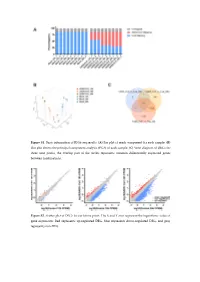
Figure S1. Basic Information of RNA-Seq Results. (A) Bar Plot of Reads Component for Each Sample
Figure S1. Basic information of RNA-seq results. (A) Bar plot of reads component for each sample. (B) Dot plot shows the principal component analysis (PCA) of each sample. (C) Venn diagram of DEGs for three time points, the overlap part of the circles represents common differentially expressed genes between combinations. Figure S2. Scatter plot of DEGs for each time point. The X and Y axes represent the logarithmic value of gene expression. Red represents up-regulated DEG, blue represents down-regulated DEG, and gray represents non-DEG. Table S1. Primers used for quantitative real-time PCR analysis of DEGs. Gene Primer Sequence Forward 5’-CTACGAGTGGATGGTCAAGAGC-3’ FOXO1 Reverse 5’-CCAGTTCCTTCATTCTGCACACG-3’ Forward 5’-GACGTCCGGCATCAGAGAAA-3’ IRS2 Reverse 5’-TCCACGGCTAATCGTCACAG-3’ Forward 5’-CACAACCAGGACCTCACACC-3’ IRS1 Reverse 5’-CTTGGCACGATAGAGAGCGT-3’ Forward 5’-AGGATACCACTCCCAACAGACCT-3’ IL6 Reverse 5’-CAAGTGCATCATCGTTGTTCATAC-3’ Forward 5’-TCACGTTGTACGCAGCTACC-3’ CCL5 Reverse 5’-CAGTCCTCTTACAGCCTTTGG-3’ Forward 5’-CTGTGCAGCCGCAGTGCCTACC-3’ BMP7 Reverse 5’-ATCCCTCCCCACCCCACCATCT-3’ Forward 5’-CTCTCCCCCTCGACTTCTGA-3’ BCL2 Reverse 5’-AGTCACGCGGAACACTTGAT-3’ Forward 5’-CTGTCGAACACAGTGGTACCTG-3’ FGF7 Reverse 5’-CCAACTGCCACTGTCCTGATTTC-3’ Forward 5’-GGGAGCCAAAAGGGTCATCA-3’ GAPDH Reverse 5’-CGTGGACTGTGGTCATGAGT-3’ Supplementary material: Differentially expressed genes log2(SADS-CoV_12h/ Qvalue (SADS-CoV _12h/ Gene Symbol Control_12h) Control_12h) PTGER4 -1.03693 6.79E-04 TMEM72 -3.08132 3.66E-04 IFIT2 -1.02918 2.11E-07 FRAT2 -1.09282 4.66E-05 -

Restoration of Sp4 in Forebrain Gabaergic Neurons Rescues Hypersensitivity to Ketamine in Sp4 Hypomorphic Mice Kerin K
International Journal of Neuropsychopharmacology, 2015, 1–9 doi:10.1093/ijnp/pyv063 Research Article research article Restoration of Sp4 in Forebrain GABAergic Neurons Rescues Hypersensitivity to Ketamine in Sp4 Hypomorphic Mice Kerin K. Higa, MS; Baohu Ji, PhD; Mahalah R. Buell, BA; Victoria B. Risbrough, PhD; Susan B. Powell, PhD; Jared W. Young, PhD; Mark A. Geyer, PhD; and Xianjin Zhou, PhD Department of Psychiatry, University of California San Diego, La Jolla, CA (Ms Higa, Drs Ji, Risbrough, Powell, Young, Geyer, and Zhou, and Ms Buell); Research Service, VA San Diego Healthcare System, La Jolla, CA (Drs Risbrough, Powell, Young, Geyer, and Zhou, and Ms Buell); Neurosciences Graduate Program, University of California San Diego, La Jolla, CA (Ms Higa). Correspondence: Xianjin Zhou, PhD, Department of Psychiatry, University of California San Diego, La Jolla, CA ([email protected]). Abstract Background: Ketamine produces schizophrenia-like behavioral phenotypes in healthy people. Prolonged ketamine effects and exacerbation of symptoms after the administration of ketamine have been observed in patients with schizophrenia. More recently, ketamine has been used as a potent antidepressant to treat patients with major depression. The genes and neurons that regulate behavioral responses to ketamine, however, remain poorly understood. Sp4 is a transcription factor for which gene expression is restricted to neuronal cells in the brain. Our previous studies demonstrated that Sp4 hypomorphic mice display several behavioral phenotypes relevant to psychiatric disorders, consistent with human SP4 gene associations with schizophrenia, bipolar disorder, and major depression. Among those behavioral phenotypes, hypersensitivity to ketamine- induced hyperlocomotion has been observed in Sp4 hypomorphic mice. Methods: In the present study, we used the Cre-LoxP system to restore Sp4 gene expression, specifically in either forebrain excitatory or GABAergic inhibitory neurons in Sp4 hypomorphic mice. -

Genomic Architecture of Histone 3 Lysine 27 Trimethylation During Late Ovine Skeletal Muscle Development
doi: 10.1111/age.12145 Genomic architecture of histone 3 lysine 27 trimethylation during late ovine skeletal muscle development † ‡ K. Byrne*1, S. McWilliam*1, T. Vuocolo*, C. Gondro , N. E. Cockett and R. L. Tellam* † *CSIRO Animal, Food and Health Sciences, Queensland Bioscience Precinct, 306 Carmody Rd, St Lucia, QLD, 4067, Australia. School of ‡ Environmental and Rural Science, University of New England, Armidale, NSW, Australia. Department of Animal, Dairy and Veterinary Sciences, Utah State University, Logan, UT, USA. Summary The ruminant developmental transition from late foetus to lamb is associated with marked changes in skeletal muscle structure and function that reflect programming for new physiological demands following birth. To determine whether epigenetic changes are involved in this transition, we investigated the genomic architecture of the chromatin modification, histone 3 lysine 27 trimethylation (H3K27me3), which typically regulates early life developmental processes; however, its role in later life processes is unclear. Chromatin immunoprecipitation coupled with next-generation sequencing was used to map H3K27me3 nucleosomes in ovine longissimus lumborum skeletal muscle at 100 days of gestation and 12 weeks post-partum. In both states, H3K27me3 modification was associated with genes, transcription start sites and CpG islands and with transcriptional silencing. The H3K27me3 peaks consisted of two major categories, promoter specific and regional, with the latter the dominant feature. Genes encoding homeobox transcription factors regulating early life development and genes involved in neural functions, particularly gated ion channels, were strongly modified by H3K27me3. Gene promoters differentially modified by H3K27me3 in the foetus and lamb were enriched for gated ion channels, which may reflect changes in neuromuscular function. -

Targeted Single-Cell RNA Sequencing of Transcription Factors Enhances the Identification of Cell Types and Trajectories
Downloaded from genome.cshlp.org on October 7, 2021 - Published by Cold Spring Harbor Laboratory Press Method Targeted single-cell RNA sequencing of transcription factors enhances the identification of cell types and trajectories Alexandra Pokhilko,1,9 Adam E. Handel,1,9 Fabiola Curion,2 Viola Volpato,3 Emma S. Whiteley,4 Sunniva Bøstrand,4 Sarah E. Newey,4 Colin J. Akerman,4 Caleb Webber,3 Michael B. Clark,5,6 Rory Bowden,2,7,8 and M. Zameel Cader1 1Translational Molecular Neuroscience Group, Weatherall Institute of Molecular Medicine, Nuffield Department of Clinical Neurosciences, University of Oxford, Oxford, OX3 9DS, United Kingdom; 2Wellcome Centre for Human Genetics, University of Oxford, Oxford, OX3 7BN, United Kingdom; 3UK Dementia Research Institute, Cardiff University, Cardiff, CF24 4HQ, United Kingdom; 4Department of Pharmacology, University of Oxford, Oxford, OX1 3QT, United Kingdom; 5Department of Psychiatry, Warneford Hospital, University of Oxford, Oxford, OX3 7JX, United Kingdom; 6Centre for Stem Cell Systems, Department of Anatomy and Neuroscience, The University of Melbourne, Parkville, Victoria 3010, Australia; 7The Walter and Eliza Hall Institute of Medical Research, Parkville, Victoria 3052, Australia; 8University of Melbourne, Department of Medical Biology, Parkville, Victoria 3052, Australia Single-cell RNA sequencing (scRNA-seq) is a widely used method for identifying cell types and trajectories in biologically heterogeneous samples, but it is limited in its detection and quantification of lowly expressed genes. This results in missing important biological signals, such as the expression of key transcription factors (TFs) driving cellular differentiation. We show that targeted sequencing of ∼1000 TFs (scCapture-seq) in iPSC-derived neuronal cultures greatly improves the bio- logical information garnered from scRNA-seq. -

The Transcriptome of Utricle Hair Cell Regeneration in the Avian Inner Ear
The Journal of Neuroscience, March 5, 2014 • 34(10):3523–3535 • 3523 Development/Plasticity/Repair The Transcriptome of Utricle Hair Cell Regeneration in the Avian Inner Ear Yuan-Chieh Ku,1 Nicole A. Renaud,1 Rose A. Veile,1 Cynthia Helms,1 Courtney C.J. Voelker,2 Mark E. Warchol,2 and Michael Lovett1 1Department of Genetics and 2Department of Otolaryngology, Washington University School of Medicine, St Louis, Missouri 63110 Sensory hair cell loss is the major cause of hearing and balance disorders. Mammals are incapable of sustained hair cell regeneration, but lower vertebrates can regenerate these mechano-electrical transducers. We present the first comprehensive transcriptome (by mRNA- Seq) of hair cell regeneration in the chick utricle. We provide pathway and pattern annotations and correlate these with the phenotypic events that occur during regeneration. These patterns are surprisingly synchronous and highly punctuated. We show how these patterns are a new resource for identifying components of the hair cell transcriptome and identify 494 new putative hair-cell-specific genes and validate three of these (of three tested) by immunohistochemical staining. We describe many surprising new components and dynamic expression patterns, particularly within NOTCH signaling. For example, we show that HES7 is specifically expressed during utricle hair cell regeneration and closely parallels the expression of HES5. Likewise, the expression of ATOH1 is closely correlated with HEYL and the HLH inhibitory transcription factors ID1, ID2, and ID4.