Functional Effects of Variation in Transcription Factor Binding Highlight Long-Range Gene Regulation by Epromoters
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-
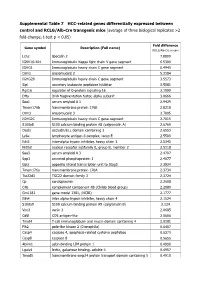
Supplemental Table 7 HCC-Related Genes Differentially Expressed Between Control and RCLG/Alb-Cre Transgenic Mice
Supplemental Table 7 HCC-related genes differentially expressed between control and RCLG/Alb-Cre transgenic mice (average of three biological replicates >2 fold-change, t-test p < 0.05) Fold difference Gene symbol Description (Full name) (RCLG/Alb-Cre vs con) Lcn2 lipocalin 2 7.8899 IGKV16-104 Immunoglobulin Kappa light chain V gene segment 6.5300 IGHG1 Immunoglobulin heavy chain C gene segment 6.4945 Orm2 orosomucoid 2 5.3184 IGHG2B Immunoglobulin heavy chain C gene segment 3.5573 Slpi secretory leukocyte peptidase inhibitor 3.5081 Rgs16 regulator of G-protein signaling 16 3.1999 Dffa DNA fragmentation factor, alpha subunit 3.0666 Saa1 serum amyloid A 1 2.9429 Tmem176b transmembrane protein 176B 2.8218 Orm3 orosomucoid 3 2.7805 IGHG2C Immunoglobulin heavy chain C gene segment 2.7015 S100a8 S100 calcium binding protein A8 (calgranulin A) 2.6769 Ocel1 occludin/ELL domain containing 1 2.6553 Ly6e lymphocyte antigen 6 complex, locus E 2.5509 Itih3 inter-alpha trypsin inhibitor, heavy chain 3 2.5345 Nr0b2 nuclear receptor subfamily 0, group B, member 2 2.5118 Saa3 serum amyloid A 3 2.4707 Spp1 secreted phosphoprotein 1 2.4077 Gats opposite strand transcription unit to Stag3 2.3934 Tmem176a transmembrane protein 176A 2.3734 Tsc22d3 TSC22 domain family 3 2.3724 Cp ceruloplasmin 2.2608 C4b complement component 4B (Childo blood group) 2.2089 Gm1381 gene model 1381, (NCBI) 2.1777 Itih4 inter alpha-trypsin inhibitor, heavy chain 4 2.1524 S100a9 S100 calcium binding protein A9 (calgranulin B) 2.124 Vnn3 vanin 3 2.0685 Cd5l CD5 antigen-like 2.0606 -

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

4-6 Weeks Old Female C57BL/6 Mice Obtained from Jackson Labs Were Used for Cell Isolation
Methods Mice: 4-6 weeks old female C57BL/6 mice obtained from Jackson labs were used for cell isolation. Female Foxp3-IRES-GFP reporter mice (1), backcrossed to B6/C57 background for 10 generations, were used for the isolation of naïve CD4 and naïve CD8 cells for the RNAseq experiments. The mice were housed in pathogen-free animal facility in the La Jolla Institute for Allergy and Immunology and were used according to protocols approved by the Institutional Animal Care and use Committee. Preparation of cells: Subsets of thymocytes were isolated by cell sorting as previously described (2), after cell surface staining using CD4 (GK1.5), CD8 (53-6.7), CD3ε (145- 2C11), CD24 (M1/69) (all from Biolegend). DP cells: CD4+CD8 int/hi; CD4 SP cells: CD4CD3 hi, CD24 int/lo; CD8 SP cells: CD8 int/hi CD4 CD3 hi, CD24 int/lo (Fig S2). Peripheral subsets were isolated after pooling spleen and lymph nodes. T cells were enriched by negative isolation using Dynabeads (Dynabeads untouched mouse T cells, 11413D, Invitrogen). After surface staining for CD4 (GK1.5), CD8 (53-6.7), CD62L (MEL-14), CD25 (PC61) and CD44 (IM7), naïve CD4+CD62L hiCD25-CD44lo and naïve CD8+CD62L hiCD25-CD44lo were obtained by sorting (BD FACS Aria). Additionally, for the RNAseq experiments, CD4 and CD8 naïve cells were isolated by sorting T cells from the Foxp3- IRES-GFP mice: CD4+CD62LhiCD25–CD44lo GFP(FOXP3)– and CD8+CD62LhiCD25– CD44lo GFP(FOXP3)– (antibodies were from Biolegend). In some cases, naïve CD4 cells were cultured in vitro under Th1 or Th2 polarizing conditions (3, 4). -

Functional Mechanisms Underlying Pleiotropic Risk Alleles at the 19P13.1 Breast&Ndash;Ovarian Cancer Susceptibility Locus
ARTICLE Received 16 Jun 2015 | Accepted 20 Jul 2016 | Published 7 Sep 2016 DOI: 10.1038/ncomms12675 OPEN Functional mechanisms underlying pleiotropic risk alleles at the 19p13.1 breast–ovarian cancer susceptibility locus Kate Lawrenson et al.# A locus at 19p13 is associated with breast cancer (BC) and ovarian cancer (OC) risk. Here we analyse 438 SNPs in this region in 46,451 BC and 15,438 OC cases, 15,252 BRCA1 mutation carriers and 73,444 controls and identify 13 candidate causal SNPs associated with serous OC (P ¼ 9.2 Â 10 À 20), ER-negative BC (P ¼ 1.1 Â 10 À 13), BRCA1-associated BC (P ¼ 7.7 Â 10 À 16) and triple negative BC (P-diff ¼ 2 Â 10 À 5). Genotype-gene expression associations are identified for candidate target genes ANKLE1 (P ¼ 2 Â 10 À 3) and ABHD8 (Po2 Â 10 À 3). Chromosome conformation capture identifies interactions between four candidate SNPs and ABHD8, and luciferase assays indicate six risk alleles increased transactivation of the ADHD8 promoter. Targeted deletion of a region containing risk SNP rs56069439 in a putative enhancer induces ANKLE1 downregulation; and mRNA stability assays indicate functional effects for an ANKLE1 30-UTR SNP. Altogether, these data suggest that multiple SNPs at 19p13 regulate ABHD8 and perhaps ANKLE1 expression, and indicate common mechanisms underlying breast and ovarian cancer risk. Correspondence and requests for materials should be addressed to A.C.A. (email: [email protected]). #A full list of authors and their affiliations appears at the end of the paper. -

Supplementary Table S4. FGA Co-Expressed Gene List in LUAD
Supplementary Table S4. FGA co-expressed gene list in LUAD tumors Symbol R Locus Description FGG 0.919 4q28 fibrinogen gamma chain FGL1 0.635 8p22 fibrinogen-like 1 SLC7A2 0.536 8p22 solute carrier family 7 (cationic amino acid transporter, y+ system), member 2 DUSP4 0.521 8p12-p11 dual specificity phosphatase 4 HAL 0.51 12q22-q24.1histidine ammonia-lyase PDE4D 0.499 5q12 phosphodiesterase 4D, cAMP-specific FURIN 0.497 15q26.1 furin (paired basic amino acid cleaving enzyme) CPS1 0.49 2q35 carbamoyl-phosphate synthase 1, mitochondrial TESC 0.478 12q24.22 tescalcin INHA 0.465 2q35 inhibin, alpha S100P 0.461 4p16 S100 calcium binding protein P VPS37A 0.447 8p22 vacuolar protein sorting 37 homolog A (S. cerevisiae) SLC16A14 0.447 2q36.3 solute carrier family 16, member 14 PPARGC1A 0.443 4p15.1 peroxisome proliferator-activated receptor gamma, coactivator 1 alpha SIK1 0.435 21q22.3 salt-inducible kinase 1 IRS2 0.434 13q34 insulin receptor substrate 2 RND1 0.433 12q12 Rho family GTPase 1 HGD 0.433 3q13.33 homogentisate 1,2-dioxygenase PTP4A1 0.432 6q12 protein tyrosine phosphatase type IVA, member 1 C8orf4 0.428 8p11.2 chromosome 8 open reading frame 4 DDC 0.427 7p12.2 dopa decarboxylase (aromatic L-amino acid decarboxylase) TACC2 0.427 10q26 transforming, acidic coiled-coil containing protein 2 MUC13 0.422 3q21.2 mucin 13, cell surface associated C5 0.412 9q33-q34 complement component 5 NR4A2 0.412 2q22-q23 nuclear receptor subfamily 4, group A, member 2 EYS 0.411 6q12 eyes shut homolog (Drosophila) GPX2 0.406 14q24.1 glutathione peroxidase -

Functional Mechanisms Underlying Pleiotropic Risk Alleles at the 19P13.1 Breast&Ndash;Ovarian Cancer Susceptibility Locus
UC Office of the President Recent Work Title Functional mechanisms underlying pleiotropic risk alleles at the 19p13.1 breast–ovarian cancer susceptibility locus Permalink https://escholarship.org/uc/item/1rx2f44n Author Lawrenson, Kate Publication Date 2016-09-07 DOI 10.1038/ncomms12675 Peer reviewed eScholarship.org Powered by the California Digital Library University of California ARTICLE Received 16 Jun 2015 | Accepted 20 Jul 2016 | Published 7 Sep 2016 DOI: 10.1038/ncomms12675 OPEN Functional mechanisms underlying pleiotropic risk alleles at the 19p13.1 breast–ovarian cancer susceptibility locus Kate Lawrenson et al.# A locus at 19p13 is associated with breast cancer (BC) and ovarian cancer (OC) risk. Here we analyse 438 SNPs in this region in 46,451 BC and 15,438 OC cases, 15,252 BRCA1 mutation carriers and 73,444 controls and identify 13 candidate causal SNPs associated with serous OC (P ¼ 9.2 Â 10 À 20), ER-negative BC (P ¼ 1.1 Â 10 À 13), BRCA1-associated BC (P ¼ 7.7 Â 10 À 16) and triple negative BC (P-diff ¼ 2 Â 10 À 5). Genotype-gene expression associations are identified for candidate target genes ANKLE1 (P ¼ 2 Â 10 À 3) and ABHD8 (Po2 Â 10 À 3). Chromosome conformation capture identifies interactions between four candidate SNPs and ABHD8, and luciferase assays indicate six risk alleles increased transactivation of the ADHD8 promoter. Targeted deletion of a region containing risk SNP rs56069439 in a putative enhancer induces ANKLE1 downregulation; and mRNA stability assays indicate functional effects for an ANKLE1 30-UTR SNP. Altogether, these data suggest that multiple SNPs at 19p13 regulate ABHD8 and perhaps ANKLE1 expression, and indicate common mechanisms underlying breast and ovarian cancer risk. -

BABAM1 (H3): 293T Lysate: Sc-128160
SANTA CRUZ BIOTECHNOLOGY, INC. BABAM1 (h3): 293T Lysate: sc-128160 BACKGROUND RECOMMENDED SUPPORT REAGENTS Consisting of around 63 million bases with over 1,400 genes, chromosome 19 To ensure optimal results, the following support reagents are recommended: makes up over 2% of human genomic DNA. Chromosome 19 includes a diver- 1) Western Blotting: use m-IgGκ BP-HRP: sc-516102 or m-IgGκ BP-HRP (Cruz sity of interesting genes and is recognized for having the greatest gene den- Marker): sc-516102-CM (dilution range: 1:1000-1:10000), Cruz Marker™ sity of the human chromosomes. It is the genetic home for a number of im- Molecular Weight Standards: sc-2035, UltraCruz® Blocking Reagent: munoglobulin superfamily members including the killer cell and leukocyte Ig- sc-516214 and Western Blotting Luminol Reagent: sc-2048. like receptors, a number of ICAMs, the CEACAM and PSG family, and Fcα receptors. Key genes for eye color and hair color also map to chromosome DATA 19. Peutz-Jeghers syndrome, spinocerebellar ataxia type 6, the stroke disor- der CADASIL, hypercholesterolemia and Insulin-dependent diabetes have AB been linked to chromosome 19. Translocations with chromosome 19 and 90 K – chromosome 14 can be seen in some lymphoproliferative disorders and typi- 55 K – < BABAM1 cally involve the proto-oncogene BCL3. 43 K – 34 K – REFERENCES 23 K – 1. Zimmermann, W., et al. 1988. Chromosomal localization of the carcinoem- bryonic antigen gene family and differential expression in various tumors. BABAM1 (H-10): sc-398570. Western blot analysis of Cancer Res. 48: 2550-2554. BABAM1 expression in non-transfected: sc-117752 (A) and human BABAM1 transfected: sc-128160 (B) 293T 2. -

DNA Methylation Profiles Correlated to Striped Bass Sperm Fertility L
Woods III et al. BMC Genomics (2018) 19:244 https://doi.org/10.1186/s12864-018-4548-6 RESEARCH ARTICLE Open Access DNA methylation profiles correlated to striped bass sperm fertility L. Curry Woods III1†, Yaokun Li2*†, Yi Ding1†,JiananLiu1, Benjamin J. Reading3,S.AdamFuller4 and Jiuzhou Song1* Abstract Background: Striped bass (Morone saxatilis) spermatozoa are used to fertilize in vitro the eggs of white bass (M. chrysops) to produce the preferred hybrid for the striped bass aquaculture industry. Currently, only one source of domestic striped bass juveniles is available to growers that is not obtained from wild-caught parents and is thus devoid of any genetic improvement in phenotypic traits of importance to aquaculture. Sperm epigenetic modification has been predicted to be associated with fertility, which could switch genes on and off without changing the DNA sequence itself. DNA methylation is one of the most common epigenetic modification types and changes in sperm epigenetics can be correlated to sub-fertility or infertility in male striped bass. The objective of this study was to find the differentially methylated regions (DMRs) between high-fertility and sub-fertility male striped bass, which could potentially regulate the fertility performance. Results: In our present study, we performed DNA methylation analysis of high-fertility and sub-fertility striped bass spermatozoa through MBD-Seq methods. A total of 171 DMRs were discovered in striped bass sperm correlated to fertility. Based on the annotation of these DMRs, we conducted a functional classification analysis and two important groups of genes including the WDR3/UTP12 and GPCR families, were discovered to be related to fertility performance of striped bass. -

The Roles of RNA Polymerase I and III Subunits Polr1a, Polr1c, and Polr1d in Craniofacial Development BY
The roles of RNA Polymerase I and III subunits Polr1a, Polr1c, and Polr1d in craniofacial development BY © 2016 Kristin Emily Noack Watt Submitted to the graduate degree program in Anatomy and Cell Biology and to the Graduate Faculty of The University of Kansas Medical Center in partial fulfillment of the requirements for the degree of Doctor of Philosophy. Paul Trainor, Co-Chairperson Brenda Rongish, Co-Chairperson Brian Andrews Jennifer Gerton Tatjana Piotrowski Russell Swerdlow Date Defended: January 26, 2016 The Dissertation Committee for Kristin Watt certifies that this is the approved version of the following dissertation: The roles of RNA Polymerase I and III subunits Polr1a, Polr1c, and Polr1d in craniofacial development Paul Trainor, Co-Chairperson Brenda Rongish, Co-Chairperson Date approved: February 2, 2016 ii Abstract Craniofacial anomalies account for approximately one-third of all birth defects. Two examples of syndromes associated with craniofacial malformations are Treacher Collins syndrome and Acrofacial Dysostosis, Cincinnati type which have phenotypic overlap including deformities of the eyes, ears, and facial bones. Mutations in TCOF1, POLR1C or POLR1D may cause Treacher Collins syndrome while mutations in POLR1A may cause Acrofacial Dysostosis, Cincinnati type. TCOF1 encodes the nucleolar phosphoprotein Treacle, which functions in rRNA transcription and modification. Previous studies demonstrated that Tcof1 mutations in mice result in reduced ribosome biogenesis and increased neuroepithelial apoptosis. This diminishes the neural crest cell (NCC) progenitor population which contribute to the development of the cranial skeleton. In contrast, apart from being subunits of RNA Polymerases (RNAP) I and/or III, nothing is known about the function of POLR1A, POLR1C, and POLR1D during embryonic and craniofacial development. -

MOCHI Enables Discovery of Heterogeneous Interactome Modules in 3D Nucleome
Downloaded from genome.cshlp.org on October 4, 2021 - Published by Cold Spring Harbor Laboratory Press MOCHI enables discovery of heterogeneous interactome modules in 3D nucleome Dechao Tian1,# , Ruochi Zhang1,# , Yang Zhang1, Xiaopeng Zhu1, and Jian Ma1,* 1Computational Biology Department, School of Computer Science, Carnegie Mellon University, Pittsburgh, PA 15213, USA #These two authors contributed equally *Correspondence: [email protected] Contact To whom correspondence should be addressed: Jian Ma School of Computer Science Carnegie Mellon University 7705 Gates-Hillman Complex 5000 Forbes Avenue Pittsburgh, PA 15213 Phone: +1 (412) 268-2776 Email: [email protected] 1 Downloaded from genome.cshlp.org on October 4, 2021 - Published by Cold Spring Harbor Laboratory Press Abstract The composition of the cell nucleus is highly heterogeneous, with different constituents forming complex interactomes. However, the global patterns of these interwoven heterogeneous interactomes remain poorly understood. Here we focus on two different interactomes, chromatin interaction network and gene regulatory network, as a proof-of-principle, to identify heterogeneous interactome modules (HIMs), each of which represents a cluster of gene loci that are in spatial contact more frequently than expected and that are regulated by the same group of transcription factors. HIM integrates transcription factor binding and 3D genome structure to reflect “transcriptional niche” in the nucleus. We develop a new algorithm MOCHI to facilitate the discovery of HIMs based on network motif clustering in heterogeneous interactomes. By applying MOCHI to five different cell types, we found that HIMs have strong spatial preference within the nucleus and exhibit distinct functional properties. Through integrative analysis, this work demonstrates the utility of MOCHI to identify HIMs, which may provide new perspectives on the interplay between transcriptional regulation and 3D genome organization. -

Human Induced Pluripotent Stem Cell–Derived Podocytes Mature Into Vascularized Glomeruli Upon Experimental Transplantation
BASIC RESEARCH www.jasn.org Human Induced Pluripotent Stem Cell–Derived Podocytes Mature into Vascularized Glomeruli upon Experimental Transplantation † Sazia Sharmin,* Atsuhiro Taguchi,* Yusuke Kaku,* Yasuhiro Yoshimura,* Tomoko Ohmori,* ‡ † ‡ Tetsushi Sakuma, Masashi Mukoyama, Takashi Yamamoto, Hidetake Kurihara,§ and | Ryuichi Nishinakamura* *Department of Kidney Development, Institute of Molecular Embryology and Genetics, and †Department of Nephrology, Faculty of Life Sciences, Kumamoto University, Kumamoto, Japan; ‡Department of Mathematical and Life Sciences, Graduate School of Science, Hiroshima University, Hiroshima, Japan; §Division of Anatomy, Juntendo University School of Medicine, Tokyo, Japan; and |Japan Science and Technology Agency, CREST, Kumamoto, Japan ABSTRACT Glomerular podocytes express proteins, such as nephrin, that constitute the slit diaphragm, thereby contributing to the filtration process in the kidney. Glomerular development has been analyzed mainly in mice, whereas analysis of human kidney development has been minimal because of limited access to embryonic kidneys. We previously reported the induction of three-dimensional primordial glomeruli from human induced pluripotent stem (iPS) cells. Here, using transcription activator–like effector nuclease-mediated homologous recombination, we generated human iPS cell lines that express green fluorescent protein (GFP) in the NPHS1 locus, which encodes nephrin, and we show that GFP expression facilitated accurate visualization of nephrin-positive podocyte formation in -

The Genetic Basis of Malformation of Cortical Development Syndromes: Primary Focus on Aicardi Syndrome
The Genetic Basis of Malformation of Cortical Development Syndromes: Primary Focus on Aicardi Syndrome Thuong Thi Ha B. Sc, M. Bio Neurogenetics Research Group The University of Adelaide Thesis submitted for the degree of Doctor of Philosophy In Discipline of Genetics and Evolution School of Biological Sciences Faculty of Science The University of Adelaide June 2018 Table of Contents Abstract 6 Thesis Declaration 8 Acknowledgements 9 Publications 11 Abbreviations 12 CHAPTER I: Introduction 15 1.1 Overview of Malformations of Cortical Development (MCD) 15 1.2 Introduction into Aicardi Syndrome 16 1.3 Clinical Features of Aicardi Syndrome 20 1.3.1 Epidemiology 20 1.3.2 Clinical Diagnosis 21 1.3.3 Differential Diagnosis 23 1.3.4 Development & Prognosis 24 1.4 Treatment 28 1.5 Pathogenesis of Aicardi Syndrome 29 1.5.1 Prenatal or Intrauterine Disturbances 29 1.5.2 Genetic Predisposition 30 1.6 Hypothesis & Aims 41 1.6.1 Hypothesis 41 1.6.2 Research Aims 41 1.7 Expected Outcomes 42 CHAPTER II: Materials & Methods 43 2.1 Study Design 43 2.1.1 Cohort of Study 44 2.1.2 Inheritance-based Strategy 45 1 2.1.3 Ethics for human and animals 46 2.2 Computational Methods 46 2.2.1 Pre-Processing Raw Reads 46 2.2.2 Sequencing Coverage 47 2.2.3 Variant Discovery 49 2.2.4 Annotating Variants 53 2.2.5 Evaluating Variants 55 2.3 Biological Methods 58 2.3.1 Cell Culture 58 2.3.2 Genomic DNA Sequencing 61 2.3.3 Plasmid cloning 69 2.3.4 RNA, whole exome & Whole Genome Sequencing 74 2.3.5 TOPFlash Assay 75 2.3.6 Western Blot 76 2.3.7 Morpholino Knockdown in Zebrafish 79 CHAPTER III: A mutation in COL4A2 causes autosomal dominant porencephaly with cataracts.