1 Single-Cell Mrna Profiling Reveals Heterogeneous Combinatorial Expression of Hoxd Genes During Limb Development Short Title
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

THE ROLE of HOXD10 in the DEVELOPMENT of MOTONEURONS in the POSTERIOR SPINAL CORD by Veeral Shailesh Shah B.S., University of Pi
THE ROLE OF HOXD10 IN THE DEVELOPMENT OF MOTONEURONS IN THE POSTERIOR SPINAL CORD by Veeral Shailesh Shah B.S., University of Pittsburgh, 2000 Submitted to the Graduate Faculty of University of Pittsburgh School of Medicine Department of Neurobiology in partial fulfillment of the requirements for the degree of Doctor of Philosophy University of Pittsburgh 2006 UNIVERSITY OF PITTSBURGH SCHOOL OF MEDICINE DEPARTMENT OF NEUROBIOLOGY This thesis was presented by Veeral Shailesh Shah It was defended on January 17, 2006 and approved by Paula Monaghan-Nichols, Ph. D, CNUP, Neurobiology Debbie Chapman, Ph. D, Biological Sciences Willi Halfter, Ph. D, CNUP, Neurobiology Carl Lagenaur, Ph. D, CNUP, Neurobiology Cynthia Forehand, Ph. D, University of Vermont, Anatomy and Neurobiology Dissertation Advisor: Cynthia Lance-Jones, Ph. D, CNUP, Neurobiology ii Copyright © by Veeral Shailesh Shah 2007 iii THE ROLE OF HOXD10 IN THE DEVELOPMENT OF MOTONEURONS IN THE POSTERIOR SPINAL CORD Veeral Shah University of Pittsburgh, 2007 Hox genes encode anterior-posterior identity during central nervous system development. Few studies have examined Hox gene function at lumbosacral (LS) levels of the spinal cord, where there is extensive information on normal development. Hoxd10 is expressed at high levels in the embryonic LS spinal cord, but not the thoracic (T) spinal cord. To test the hypothesis that restricted expression of Hoxd10 contributes to the attainment of an LS identity, and specifically an LS motoneuron identity, Hoxd10 was ectopically expressed in T segments in chick embryos via in ovo electroporation. Electroporations were carried out at early neural tube stages (stages 13-15) and at the onset of motoneuron differentiation (stages 17-18). -
The NANOG Transcription Factor Induces Type 2 Deiodinase Expression and Regulates the Intracellular Activation of Thyroid Hormone in Keratinocyte Carcinomas
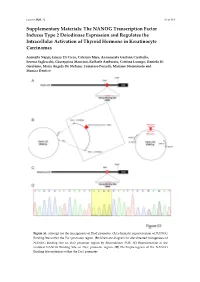
Cancers 2020, 12 S1 of S18 Supplementary Materials: The NANOG Transcription Factor Induces Type 2 Deiodinase Expression and Regulates the Intracellular Activation of Thyroid Hormone in Keratinocyte Carcinomas Annarita Nappi, Emery Di Cicco, Caterina Miro, Annunziata Gaetana Cicatiello, Serena Sagliocchi, Giuseppina Mancino, Raffaele Ambrosio, Cristina Luongo, Daniela Di Girolamo, Maria Angela De Stefano, Tommaso Porcelli, Mariano Stornaiuolo and Monica Dentice Figure S1. Strategy for the mutagenesis of Dio2 promoter. (A) Schematic representation of NANOG Binding Site within the Dio2 promoter region. (B) Schematic diagram for site‐directed mutagenesis of NANOG Binding Site on Dio2 promoter region by Recombinant PCR. (C) Representation of the mutated NANOG Binding Site on Dio2 promoter region. (D) Electropherogram of the NANOG Binding Site mutation within the Dio2 promoter. Cancers 2020, 12 S2 of S18 Figure S2. Strategy for the silencing of NANOG expression. (A) Cloning strategies for the generation of NANOG shRNA expression vectors. (B) Electropherograms of the NANOG shRNA sequences cloned into pcDNA3.1 vector. (C) Validation of effective NANOG down-modulation by two different NANOG shRNA vectors was assessed by Western Blot analysis of NANOG expression in BCC cells. (D) Quantification of NANOG protein levels versus Tubulin levels in the same experiment as in C is represented by histograms. Cancers 2020, 12 S3 of S18 Figure S3. The CD34+ cells are characterized by the expression of typical epithelial stemness genes. The mRNA levels of a panel of indicated stemness markers of epidermis were measured by Real Time PCR in the same experiment indicated in figure 3F and G. Cancers 2020, 12 S4 of S18 Figure S4. -

Role of HOX Genes in Stem Cell Differentiation and Cancer
Thomas Jefferson University Jefferson Digital Commons Kimmel Cancer Center Papers, Presentations, and Grand Rounds Kimmel Cancer Center 7-22-2018 Role of HOX Genes in Stem Cell Differentiation and Cancer. Seema Bhatlekar Helen F. Graham Cancer Center and Research Institute; University of Delaware Jeremy Z Fields CATX Inc. Bruce M. Boman Thomas Jefferson University; Helen F. Graham Cancer Center and Research Institute; University of Delaware; CATX Inc. Follow this and additional works at: https://jdc.jefferson.edu/kimmelgrandrounds Part of the Oncology Commons Let us know how access to this document benefits ouy Recommended Citation Bhatlekar, Seema; Fields, Jeremy Z; and Boman, Bruce M., "Role of HOX Genes in Stem Cell Differentiation and Cancer." (2018). Kimmel Cancer Center Papers, Presentations, and Grand Rounds. Paper 62. https://jdc.jefferson.edu/kimmelgrandrounds/62 This Article is brought to you for free and open access by the Jefferson Digital Commons. The Jefferson Digital Commons is a service of Thomas Jefferson University's Center for Teaching and Learning (CTL). The Commons is a showcase for Jefferson books and journals, peer-reviewed scholarly publications, unique historical collections from the University archives, and teaching tools. The Jefferson Digital Commons allows researchers and interested readers anywhere in the world to learn about and keep up to date with Jefferson scholarship. This article has been accepted for inclusion in Kimmel Cancer Center Papers, Presentations, and Grand Rounds by an authorized administrator of the Jefferson Digital Commons. For more information, please contact: [email protected]. Hindawi Stem Cells International Volume 2018, Article ID 3569493, 15 pages https://doi.org/10.1155/2018/3569493 Review Article Role of HOX Genes in Stem Cell Differentiation and Cancer 1,2 3 1,2,3,4 Seema Bhatlekar , Jeremy Z. -

The Human Gene Connectome As a Map of Short Cuts for Morbid Allele Discovery
The human gene connectome as a map of short cuts for morbid allele discovery Yuval Itana,1, Shen-Ying Zhanga,b, Guillaume Vogta,b, Avinash Abhyankara, Melina Hermana, Patrick Nitschkec, Dror Friedd, Lluis Quintana-Murcie, Laurent Abela,b, and Jean-Laurent Casanovaa,b,f aSt. Giles Laboratory of Human Genetics of Infectious Diseases, Rockefeller Branch, The Rockefeller University, New York, NY 10065; bLaboratory of Human Genetics of Infectious Diseases, Necker Branch, Paris Descartes University, Institut National de la Santé et de la Recherche Médicale U980, Necker Medical School, 75015 Paris, France; cPlateforme Bioinformatique, Université Paris Descartes, 75116 Paris, France; dDepartment of Computer Science, Ben-Gurion University of the Negev, Beer-Sheva 84105, Israel; eUnit of Human Evolutionary Genetics, Centre National de la Recherche Scientifique, Unité de Recherche Associée 3012, Institut Pasteur, F-75015 Paris, France; and fPediatric Immunology-Hematology Unit, Necker Hospital for Sick Children, 75015 Paris, France Edited* by Bruce Beutler, University of Texas Southwestern Medical Center, Dallas, TX, and approved February 15, 2013 (received for review October 19, 2012) High-throughput genomic data reveal thousands of gene variants to detect a single mutated gene, with the other polymorphic genes per patient, and it is often difficult to determine which of these being of less interest. This goes some way to explaining why, variants underlies disease in a given individual. However, at the despite the abundance of NGS data, the discovery of disease- population level, there may be some degree of phenotypic homo- causing alleles from such data remains somewhat limited. geneity, with alterations of specific physiological pathways under- We developed the human gene connectome (HGC) to over- come this problem. -

Interferon Regulatory Factors (IRF) 1 and 8 in the Context of Pathogen Challenge by Chip on Chip and Genome Wide Transcription Profiling
The identification of transcriptional targets of Interferon Regulatory Factors (IRF) 1 and 8 in the context of pathogen challenge by ChIP on Chip and genome wide transcription profiling Oxana Kapoustina August 2009 A thesis submitted for the fulfillment of the Master of Science degree, Dept. of Biochemistry Library and Archives Bibliothèque et Canada Archives Canada Published Heritage Direction du Branch Patrimoine de l’édition 395 Wellington Street 395, rue Wellington Ottawa ON K1A 0N4 Ottawa ON K1A 0N4 Canada Canada Your file Votre référence ISBN: 978-0-494-66143-7 Our file Notre référence ISBN: 978-0-494-66143-7 NOTICE: AVIS: The author has granted a non- L’auteur a accordé une licence non exclusive exclusive license allowing Library and permettant à la Bibliothèque et Archives Archives Canada to reproduce, Canada de reproduire, publier, archiver, publish, archive, preserve, conserve, sauvegarder, conserver, transmettre au public communicate to the public by par télécommunication ou par l’Internet, prêter, telecommunication or on the Internet, distribuer et vendre des thèses partout dans le loan, distribute and sell theses monde, à des fins commerciales ou autres, sur worldwide, for commercial or non- support microforme, papier, électronique et/ou commercial purposes, in microform, autres formats. paper, electronic and/or any other formats. The author retains copyright L’auteur conserve la propriété du droit d’auteur ownership and moral rights in this et des droits moraux qui protège cette thèse. Ni thesis. Neither the thesis nor la thèse ni des extraits substantiels de celle-ci substantial extracts from it may be ne doivent être imprimés ou autrement printed or otherwise reproduced reproduits sans son autorisation. -

Xo PANEL DNA GENE LIST
xO PANEL DNA GENE LIST ~1700 gene comprehensive cancer panel enriched for clinically actionable genes with additional biologically relevant genes (at 400 -500x average coverage on tumor) Genes A-C Genes D-F Genes G-I Genes J-L AATK ATAD2B BTG1 CDH7 CREM DACH1 EPHA1 FES G6PC3 HGF IL18RAP JADE1 LMO1 ABCA1 ATF1 BTG2 CDK1 CRHR1 DACH2 EPHA2 FEV G6PD HIF1A IL1R1 JAK1 LMO2 ABCB1 ATM BTG3 CDK10 CRK DAXX EPHA3 FGF1 GAB1 HIF1AN IL1R2 JAK2 LMO7 ABCB11 ATR BTK CDK11A CRKL DBH EPHA4 FGF10 GAB2 HIST1H1E IL1RAP JAK3 LMTK2 ABCB4 ATRX BTRC CDK11B CRLF2 DCC EPHA5 FGF11 GABPA HIST1H3B IL20RA JARID2 LMTK3 ABCC1 AURKA BUB1 CDK12 CRTC1 DCUN1D1 EPHA6 FGF12 GALNT12 HIST1H4E IL20RB JAZF1 LPHN2 ABCC2 AURKB BUB1B CDK13 CRTC2 DCUN1D2 EPHA7 FGF13 GATA1 HLA-A IL21R JMJD1C LPHN3 ABCG1 AURKC BUB3 CDK14 CRTC3 DDB2 EPHA8 FGF14 GATA2 HLA-B IL22RA1 JMJD4 LPP ABCG2 AXIN1 C11orf30 CDK15 CSF1 DDIT3 EPHB1 FGF16 GATA3 HLF IL22RA2 JMJD6 LRP1B ABI1 AXIN2 CACNA1C CDK16 CSF1R DDR1 EPHB2 FGF17 GATA5 HLTF IL23R JMJD7 LRP5 ABL1 AXL CACNA1S CDK17 CSF2RA DDR2 EPHB3 FGF18 GATA6 HMGA1 IL2RA JMJD8 LRP6 ABL2 B2M CACNB2 CDK18 CSF2RB DDX3X EPHB4 FGF19 GDNF HMGA2 IL2RB JUN LRRK2 ACE BABAM1 CADM2 CDK19 CSF3R DDX5 EPHB6 FGF2 GFI1 HMGCR IL2RG JUNB LSM1 ACSL6 BACH1 CALR CDK2 CSK DDX6 EPOR FGF20 GFI1B HNF1A IL3 JUND LTK ACTA2 BACH2 CAMTA1 CDK20 CSNK1D DEK ERBB2 FGF21 GFRA4 HNF1B IL3RA JUP LYL1 ACTC1 BAG4 CAPRIN2 CDK3 CSNK1E DHFR ERBB3 FGF22 GGCX HNRNPA3 IL4R KAT2A LYN ACVR1 BAI3 CARD10 CDK4 CTCF DHH ERBB4 FGF23 GHR HOXA10 IL5RA KAT2B LZTR1 ACVR1B BAP1 CARD11 CDK5 CTCFL DIAPH1 ERCC1 FGF3 GID4 HOXA11 -

The Posterior HOXD Locus: Its Contribution to Phenotype and Malignancy of Ewing Sarcoma Supplementary Materials
www.impactjournals.com/oncotarget/ Oncotarget, Supplementary Materials 2016 The posterior HOXD locus: Its contribution to phenotype and malignancy of Ewing sarcoma Supplementary Materials SUPPLEMENTARY MATERIALS AND (antisense); si.HOXD11_6 5′-GGCCGAGCGGAUCCU METHODS AAUAUU-3′ (sense) and 5′-AAUAUUAGGAUCCGCU CGGCC-3′ (antisense); si.HOXD13_1 5′-GAGAGUGCC UUACACCAAAUU-3′ (sense) and 5′-UUUGGUGUAAG Cell lines GCACUCUCUU-3′ (antisense); si.HOXD13_2 5′-GAA Osteosarcoma cell lines (HOS, MG-63, SaOS-2 and CCUAUCUGAGAGACAAUU-3′ (sense) and 5′-UUGUC U2 OS) were kindly provided by Jan Smida and Michaela UCUCAGAUAGGUUCGU-3′ (antisense); si.HOXD13_3 Nathrath, Institute of Pathology and Radiation Biology 5′-CAGUAUAAAGGGACUUGAAGC-3′ (sense) and (Neuherberg, Germany) and ES line EW7 by Olivier 5′-GCUUCAAGUCCCUUUAUACUG-3′ (antisense); si. Delattre, Institut Curie (Paris, France). A673 was purchased TCF4_3 5′-CAGCUGUUUGGUCUAGAAATT-3′ (sense) from ATCC (LGC Standards GmbH). The other ES lines and 5′-UUUCUAGACCAAACAGCUGTG-3′ (antisense); (MHH-ES1, RD-ES, SK-ES1, SK-N-MC and TC-71) si.TCF4_5 5′-CGACAAGAAGGAUAUCAAATT-3′ and neuroblastoma lines (SH-SY5Y and SIMA) were (sense) and 5′-UUUGAUAUCCUUCUUGUCGTC-3′ obtained from the German Collection of Microorganisms (antisense) and si.control 5′-UUCUCCGAACGUGUC and Cell Cultures (DSMZ). Mesenchymal stem cells L87 ACGU-3′ (sense) and 5′-ACGUGACACGUUCGGAG and V54.2, kindly provided by Peter Nelson (Medizinische AA-3′ (antisense). Klinik und Poliklinik IV, München, Germany), were immortalized with SV40 large T-antigen [1]. Retrovirus Short hairpin RNA coding oligonucleotides packaging cell line PT67 was from Takara Bio Europe/ Clontech. DKK2 (sh.DKK2) 5′-GATCCGGGGATTT GCTATCATAATATTCAAGAGATATTATGATAGCAAA Small interfering RNAs used TCCCCTTTTTTCTAGAG-3′ (sense) and 5′-AATTCTCT AGAAAAAAGGGGATTTGCTATCATAATATCTCTTG siRNAs for EWS-FLI1 were synthesized at MWG AATATTATGATAGCAAATCCCCG-3′ (antisense); Biotech and correspond to published sequences [2]. -

BMC Biology Biomed Central
BMC Biology BioMed Central Research article Open Access Classification and nomenclature of all human homeobox genes PeterWHHolland*†1, H Anne F Booth†1 and Elspeth A Bruford2 Address: 1Department of Zoology, University of Oxford, South Parks Road, Oxford, OX1 3PS, UK and 2HUGO Gene Nomenclature Committee, European Bioinformatics Institute (EMBL-EBI), Wellcome Trust Genome Campus, Hinxton, Cambridgeshire, CB10 1SA, UK Email: Peter WH Holland* - [email protected]; H Anne F Booth - [email protected]; Elspeth A Bruford - [email protected] * Corresponding author †Equal contributors Published: 26 October 2007 Received: 30 March 2007 Accepted: 26 October 2007 BMC Biology 2007, 5:47 doi:10.1186/1741-7007-5-47 This article is available from: http://www.biomedcentral.com/1741-7007/5/47 © 2007 Holland et al; licensee BioMed Central Ltd. This is an Open Access article distributed under the terms of the Creative Commons Attribution License (http://creativecommons.org/licenses/by/2.0), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited. Abstract Background: The homeobox genes are a large and diverse group of genes, many of which play important roles in the embryonic development of animals. Increasingly, homeobox genes are being compared between genomes in an attempt to understand the evolution of animal development. Despite their importance, the full diversity of human homeobox genes has not previously been described. Results: We have identified all homeobox genes and pseudogenes in the euchromatic regions of the human genome, finding many unannotated, incorrectly annotated, unnamed, misnamed or misclassified genes and pseudogenes. -

Hox5 Interacts with Plzf to Restrict Shh Expression in the Developing Forelimb
Hox5 interacts with Plzf to restrict Shh expression in the developing forelimb Ben Xua,1, Steven M. Hrycaja, Daniel C. McIntyrea,2, Nicholas C. Bakera, Jun K. Takeuchib, Lucie Jeannottec, Zachary B. Gaberd, Bennett G. Novitchd, and Deneen M. Wellika,3 aDepartment of Internal Medicine, Division of Molecular Medicine and Genetics, University of Michigan, Ann Arbor, MI 48109; bCardiovascular Regeneration Institute of Molecular and Cellular Biosciences, University of Tokyo, Tokyo 113-0032, Japan; cCentre de Recherche en Cancérologie de l’Université Laval, Centre Hospitalier Universitaire de Québec, Québec, Canada G1R 2J6; and dDepartment of Neurobiology, Eli and Edythe Broad Center of Regenerative Medicine and Stem Cell Research, David Geffen School of Medicine at UCLA, Los Angeles, CA 90095 Edited by Clifford J. Tabin, Harvard Medical School, Boston, MA, and approved October 18, 2013 (received for review August 8, 2013) To date, only the five most posterior groups of Hox genes, Hox9– expression in limb AP patterning (12, 13). Although misexpression Hox13, have demonstrated loss-of-function roles in limb pattern- of more anterior Hox genes in mice reportedly affects limb patterning ing. Individual paralog groups control proximodistal patterning of (16), no loss-of-function mutants of anterior, non–abdominal B the limb skeletal elements. Hox9 genes also initiate the onset of (AbdB)-related genes have demonstrated defects in the patterning of Hand2 expression in the posterior forelimb compartment, and col- limb skeletal elements. Moreover, no HoxB or HoxC group genes had lectively, the posterior HoxA/D genes maintain posterior Sonic been shown to play a role in forelimb development until a report by Hedgehog (Shh) expression. -

Variations in Microrna-25 Expression Influence the Severity of Diabetic
BASIC RESEARCH www.jasn.org Variations in MicroRNA-25 Expression Influence the Severity of Diabetic Kidney Disease † † † Yunshuang Liu,* Hongzhi Li,* Jieting Liu,* Pengfei Han, Xuefeng Li, He Bai,* Chunlei Zhang,* Xuelian Sun,* Yanjie Teng,* Yufei Zhang,* Xiaohuan Yuan,* Yanhui Chu,* and Binghai Zhao* *Heilongjiang Key Laboratory of Anti-Fibrosis Biotherapy, Medical Research Center, Heilongjiang, People’s Republic of China; and †Clinical Laboratory of Hong Qi Hospital, Mudanjiang Medical University, Heilongjiang, People’s Republic of China ABSTRACT Diabetic nephropathy is characterized by persistent albuminuria, progressive decline in GFR, and second- ary hypertension. MicroRNAs are dysregulated in diabetic nephropathy, but identification of the specific microRNAs involved remains incomplete. Here, we show that the peripheral blood from patients with diabetes and the kidneys of animals with type 1 or 2 diabetes have low levels of microRNA-25 (miR-25) compared with those of their nondiabetic counterparts. Furthermore, treatment with high glucose decreased the expression of miR-25 in cultured kidney cells. In db/db mice, systemic administration of an miR-25 agomir repressed glomerular fibrosis and reduced high BP. Notably, knockdown of miR-25 in normal mice by systemic administration of an miR-25 antagomir resulted in increased proteinuria, extracellular matrix accumulation, podocyte foot process effacement, and hypertension with renin-angiotensin system activation. However, excessive miR-25 did not cause kidney dysfunction in wild-type mice. RNA sequencing showed the alteration of miR-25 target genes in antagomir-treated mice, including the Ras-related gene CDC42. In vitro,cotrans- fection with the miR-25 antagomir repressed luciferase activity from a reporter construct containing the CDC42 39 untranslated region. -

Identification and Validation of Candidate Epigenetic Biomarkers in Lung Adenocarcinoma Received: 01 June 2016 Iben Daugaard1, Diana Dominguez1, Tina E
www.nature.com/scientificreports OPEN Identification and validation of candidate epigenetic biomarkers in lung adenocarcinoma Received: 01 June 2016 Iben Daugaard1, Diana Dominguez1, Tina E. Kjeldsen1, Lasse S. Kristensen1, Henrik Hager2, Accepted: 23 September 2016 Tomasz K. Wojdacz1,3,* & Lise Lotte Hansen1,* Published: 26 October 2016 Lung cancer is the number one cause of cancer-related deaths worldwide. DNA methylation is an epigenetic mechanism that regulates gene expression, and disease-specific methylation changes can be targeted as biomarkers. We have compared the genome-wide methylation pattern in tumor and tumor- adjacent normal lung tissue from four lung adenocarcinoma (LAC) patients using DNA methylation microarrays and identified 74 differentially methylated regions (DMRs). Eighteen DMRs were selected for validation in a cohort comprising primary tumors from 52 LAC patients and tumor-adjacent normal lung tissue from 32 patients by methylation-sensitive high resolution melting (MS-HRM) analysis. Significant increases in methylation were confirmed for 15 DMRs associated with the genes and genomic regions: OSR1, SIM1, GHSR, OTX2, LOC648987, HIST1H3E, HIST1H3G/HIST1H2BI, HIST1H2AJ/ HIST1H2BM, HOXD10, HOXD3, HOXB3/HOXB4, HOXA3, HOXA5, Chr1(q21.1).A, and Chr6(p22.1). In particular the OSR1, SIM1 and HOXB3/HOXB4 regions demonstrated high potential as biomarkers in LAC. For OSR1, hypermethylation was detected in 47/48 LAC cases compared to 1/31 tumor-adjacent normal lung samples. Similarly, 45/49 and 36/48 LAC cases compared to 3/31 and 0/31 tumor-adjacent normal lung samples showed hypermethylation of the SIM1 and HOXB3/HOXB4 regions, respectively. In conclusion, this study has identified and validated 15 DMRs that can be targeted as biomarkers in LAC. -

Supplementary File Table S1
Supplementary file Table S1 Grouping of Homeobox genes according to their main known function. Anatomical Structure Morphogenesis EN1, HOXC10, HOXC13, HOXD3, LBX1, SIX2, SIX4 Organ Morphogenesis CDX1, CDX2, HOXA11, HOXA13, ISL1, LHX1, PAX3, PDHX, PITX2, PITX3, PROX1, SIX6 Body Pattern Formation ALX3, EMX2, HHEX, HOXA11, HOXA2, HOXA4, HOXA5, HOXA6, HOXB1, HOXB5, HOXB6, HOXC5, HOXD10, HOXD8, LMX1B, PITX2 Ectoderm Development PROX1, VAX2 Endoderm Development HOXC11 Brain & Nervous System Development Brain Development ALX1, DLX2, EMX2 Nervous System Development: ARX, DLX5, DLX6, HOXD10, LBX1, LHX1, OTP, PAX3, PHOX2A, PHOX2B Skeletal Development: ALX3, ALX4, DLX3, DLX5, DLX6, EN1, HOXA11, HOXA13, HOXA2, HOXB6, HOXD10, HOXD13, MSX2 Muscle Development: BARX2, MKX, SIRT1, SIRT2, SIX1 Other Homeobox Genes Involved In BARX1, CDX4, CUX1, DLX1, EMX1, EN2, Multicellular Organismal HOXA1, HOXA7, HOXA9, HOXB13, HOXB2, Development: HOXB3, HOXB4, HOXB7, HOXB8, HOXB9, HOXC12, HOXC8, HOXC9, HOXD1, HOXD11, HOXD12, HOXD9, ISL2, LBX2, LMX1A, MEIS1, NKX3-1, OTX1, TLX1, VAX1, VSX1, VSX2 Homeobox Genes Involved In Cell ARX, EMX2, HHEX, HLX, HOPX, LBX1, LHX1, Differentiation: LMX1B, MIXL1, OTP, PHOX2A, SIRT1, VSX2 Other Genes: PHTF1, SIRT3, SIRT6, SIRT7, ZHX1, ZHX2 Homeobox genes include two subsets of genes coding for transcription factors involved in multiple functions. The clustered HOX genes are indicated in bold. Supplementary file Figure S2 5’ Spatial collinearity 3’ HOXA Chr. 7p15.3 HOXB Chr. 17q21.3 HOXC Chr. 12q13.3 HOXD Chr. 2q31 13 12 11 10 9 8 7 6 5 4 3 2 1 Paralogous HOX groups Distribution of the 39 human HOX genes in four clusters located in different chromosomal regions*. Blue indicates anterior HOX genes. Yellow, paralogy group 3 Hox genes, green and purple indicatete central HOX genes and Red the posterior HOX genes.