Deciphering Binding Patterns of Therapeutic Antibodies with Immune Cells
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Antigen –Antibody Interaction By: Dr
TDC 5TH SEM MAJOR: PAPER 5.3 ANTIGEN –ANTIBODY INTERACTION BY: DR. LUNA PHUKAN Antigen-antibody interaction, or antigen-antibody reaction, is a specific chemical interaction between antibodies produced by B cells of the white blood cells and antigens during immune reaction. The antigens and antibodies combine by a process called agglutination Antigen-antibody interaction, or antigen-antibody reaction, is a specific chemical interaction between antibodies produced by B cells of the white blood cells and antigens during immune reaction. The antigens and antibodies combine by a process called agglutination. It is the fundamental reaction in the body by which the body is protected from complex foreign molecules, such as pathogens and their chemical toxins. In the blood, the antigens are specifically and with high affinity bound by antibodies to form an antigen- antibody complex. The immune complex is then transported to cellular systems where it can be destroyed or deactivated. The first correct description of the antigen-antibody reaction was given by Richard J. Goldberg at the University of Wisconsin in 1952. It came to be known as "Goldberg's theory" (of antigen-antibody reaction) There are several types of antibodies and antigens, and each antibody is capable of binding only to a specific antigen. The specificity of the binding is due to specific chemical constitution of each antibody. The antigenic determinant or epitope is recognized by the paratope of the antibody, situated at the variable region of the polypeptide chain. The variable region in turn has hyper-variable regions which are unique amino acid sequences in each antibody. -

Complement Peptide C3a Receptor 1 Promotes Optic Nerve Degeneration in DBA/2J Mice Jeffrey M
Harder et al. Journal of Neuroinflammation (2020) 17:336 https://doi.org/10.1186/s12974-020-02011-z RESEARCH Open Access Complement peptide C3a receptor 1 promotes optic nerve degeneration in DBA/2J mice Jeffrey M. Harder1, Pete A. Williams1,2 , Catherine E. Braine1,3, Hongtian S. Yang1, Jocelyn M. Thomas1, Nicole E. Foxworth1, Simon W. M. John1,4,5* and Gareth R. Howell1,6,7* Abstract Background: The risk of glaucoma increases significantly with age and exposure to elevated intraocular pressure, two factors linked with neuroinflammation. The complement cascade is a complex immune process with many bioactive end-products, including mediators of inflammation. Complement cascade activation has been shown in glaucoma patients and models of glaucoma. However, the function of complement-mediated inflammation in glaucoma is largely untested. Here, the complement peptide C3a receptor 1 was genetically disrupted in DBA/2J mice, an ocular hypertensive model of glaucoma, to test its contribution to neurodegeneration. Methods: A null allele of C3ar1 was backcrossed into DBA/2J mice. Development of iris disease, ocular hypertension, optic nerve degeneration, retinal ganglion cell activity, loss of RGCs, and myeloid cell infiltration in C3ar1-deficient and sufficient DBA/2J mice were compared across multiple ages. RNA sequencing was performed on microglia from primary culture to determine global effects of C3ar1 on microglia gene expression. Results: Deficiency in C3ar1 lowered the risk of degeneration in ocular hypertensive mice without affecting intraocular pressure elevation at 10.5 months of age. Differences were found in the percentage of mice affected, but not in individual characteristics of disease progression. -

WO 2016/147053 Al 22 September 2016 (22.09.2016) P O P C T
(12) INTERNATIONAL APPLICATION PUBLISHED UNDER THE PATENT COOPERATION TREATY (PCT) (19) World Intellectual Property Organization International Bureau (10) International Publication Number (43) International Publication Date WO 2016/147053 Al 22 September 2016 (22.09.2016) P O P C T (51) International Patent Classification: (71) Applicant: RESVERLOGIX CORP. [CA/CA]; 300, A61K 31/551 (2006.01) A61P 37/02 (2006.01) 4820 Richard Road Sw, Calgary, AB, T3E 6L1 (CA). A61K 31/517 (2006.01) C07D 239/91 (2006.01) (72) Inventors: WASIAK, Sylwia; 431 Whispering Water (21) International Application Number: Trail, Calgary, AB, T3Z 3V1 (CA). KULIKOWSKI, PCT/IB20 16/000443 Ewelina, B.; 31100 Swift Creek Terrace, Calgary, AB, T3Z 0B7 (CA). HALLIDAY, Christopher, R.A.; 403 (22) International Filing Date: 138-18th Avenue SE, Calgary, AB, T2G 5P9 (CA). GIL- 10 March 2016 (10.03.2016) HAM, Dean; 249 Scenic View Close NW, Calgary, AB, (25) Filing Language: English T3L 1Y5 (CA). (26) Publication Language: English (81) Designated States (unless otherwise indicated, for every kind of national protection available): AE, AG, AL, AM, (30) Priority Data: AO, AT, AU, AZ, BA, BB, BG, BH, BN, BR, BW, BY, 62/132,572 13 March 2015 (13.03.2015) US BZ, CA, CH, CL, CN, CO, CR, CU, CZ, DE, DK, DM, 62/264,768 8 December 2015 (08. 12.2015) US DO, DZ, EC, EE, EG, ES, FI, GB, GD, GE, GH, GM, GT, [Continued on nextpage] (54) Title: COMPOSITIONS AND THERAPEUTIC METHODS FOR THE TREATMENT OF COMPLEMENT-ASSOCIATED DISEASES (57) Abstract: The invention comprises methods of modulating the complement cascade in a mammal and for treating and/or preventing diseases and disorders as sociated with the complement pathway by administering a compound of Formula I or Formula II, such as, for example, 2-(4-(2-hydroxyethoxy)-3,5-dimethylphenyl)- 5,7-dimethoxyquinazolin-4(3H)-one or a pharmaceutically acceptable salt thereof. -

Active Complement Component 1, Q
APD208Hu01 200µg Active Complement Component 1, Q Subcomponent B (C1qB) Organism Species: Homo sapiens (Human) Instruction manual FOR RESEARCH USE ONLY NOT FOR USE IN CLINICAL DIAGNOSTIC PROCEDURES 1th Edition (Apr, 2016) [ PROPERTIES ] Source: Prokaryotic expression. Host: E. coli Residues: Gln28~Ala253 Tags: Two N-terminal Tags, His-tag and SUMO-tag Purity: >94% Endotoxin Level: <1.0EU per 1μg (determined by the LAL method). Buffer Formulation: 20mM Tris, 150mM NaCl, pH8.0, containing 0.05% sarcosyl and 5% trehalose. Applications: Cell culture; Activity Assays. (May be suitable for use in other assays to be determined by the end user.) Predicted isoelectric point: 8.9 Predicted Molecular Mass: 37.5kDa Accurate Molecular Mass: 38kDa as determined by SDS-PAGE reducing conditions. [ USAGE ] Reconstitute in 20mM Tris, 150mM NaCl (pH8.0) to a concentration of 0.1-1.0 mg/mL. Do not vortex. [ STORAGE AND STABILITY ] Storage: Avoid repeated freeze/thaw cycles. Store at 2-8oC for one month. Aliquot and store at -80oC for 12 months. Stability Test: The thermal stability is described by the loss rate. The loss rate was determined by accelerated thermal degradation test, that is, incubate the protein at 37oC for 48h, and no obvious degradation and precipitation were observed. The loss rate is less than 5% within the expiration date under appropriate storage condition. [ SEQUENCE ] [ ACTIVITY ] The complement component 1q (or simply C1q) is a protein complex involved in the complement system, which is part of the innate immune system. C1q together with C1r and C1s form the C1 complex. Antibodies of the adaptive immune system can bind antigen, forming an antigen-antibody complex. -

High-Throughput Proteomic Profiling of the Fish Liver Following Bacterial
Causey et al. BMC Genomics (2018) 19:719 https://doi.org/10.1186/s12864-018-5092-0 RESEARCH ARTICLE Open Access High-throughput proteomic profiling of the fish liver following bacterial infection Dwight R Causey1, Moritz A N Pohl1, David A Stead2, Samuel A M Martin1, Christopher J Secombes1 and Daniel J Macqueen1* Abstract Background: High-throughput proteomics was used to determine the role of the fish liver in defense responses to bacterial infection. This was done using a rainbow trout (Oncorhynchus mykiss) model following infection with Aeromonas salmonicida, the causative agent of furunculosis. The vertebrate liver has multifaceted functions in innate immunity, metabolism, and growth; we hypothesize this tissue serves a dual role in supporting host defense in parallel to metabolic adjustments that promote effectiveimmunefunction.Whilepaststudieshavereported mRNA responses to A. salmonicida in salmonids, the impact of bacterial infection on the liver proteome remains uncharacterized in fish. Results: Rainbow trout were injected with A. salmonicida or PBS (control) and liver extracted 48 h later for analysis on a hybrid quadrupole-Orbitrap mass spectrometer. A label-free method was used for protein abundance profiling, which revealed a strong innate immune response along with evidence to support parallel rewiring of metabolic and growth systems. 3076 proteins were initially identified against all proteins (n = 71,293 RefSeq proteins) annotated in a single high-quality rainbow trout reference genome, of which 2433 were maintained for analysis post-quality filtering. Among the 2433 proteins, 109 showed significant differential abundance following A. salmonicida challenge, including many upregulated complement system and acute phase response proteins, in addition to molecules with putative functions that may support metabolic re-adjustments. -

Subdominant CD8 T-Cell Epitopes Account for Protection Against Cytomegalovirus Independent of Immunodominationᰔ† Rafaela Holtappels,1* Christian O
JOURNAL OF VIROLOGY, June 2008, p. 5781–5796 Vol. 82, No. 12 0022-538X/08/$08.00ϩ0 doi:10.1128/JVI.00155-08 Copyright © 2008, American Society for Microbiology. All Rights Reserved. Subdominant CD8 T-Cell Epitopes Account for Protection against Cytomegalovirus Independent of Immunodominationᰔ† Rafaela Holtappels,1* Christian O. Simon,1 Michael W. Munks,2‡ Doris Thomas,1 Petra Deegen,1 Birgit Ku¨hnapfel,1 Torsten Da¨ubner,1 Simone F. Emde,1 Ju¨rgen Podlech,1 Natascha K. A. Grzimek,1 Silke A. Oehrlein-Karpi,1 Ann B. Hill,2 and Matthias J. Reddehase1 Institute for Virology, Johannes Gutenberg University, 55131 Mainz, Germany,1 and Department of Molecular Microbiology and Immunology, Oregon Health and Science University, Portland, Oregon 972392 Received 22 January 2008/Accepted 17 March 2008 Cytomegalovirus (CMV) infection continues to be a complication in recipients of hematopoietic stem cell transplantation (HSCT). Preexisting donor immunity is recognized as a favorable prognostic factor for the reconstitution of protective antiviral immunity mediated primarily by CD8 T cells. Furthermore, adoptive transfer of CMV-specific memory CD8 T (CD8-TM) cells is a therapeutic option for preventing CMV disease in HSCT recipients. Given the different CMV infection histories of donor and recipient, a problem may arise from an antigenic mismatch between the CMV variant that has primed donor immunity and the CMV variant acquired by the recipient. Here, we have used the BALB/c mouse model of CMV infection in the immunocom- promised host to evaluate the importance of donor-recipient CMV matching in immundominant epitopes (IDEs). For this, we generated the murine CMV (mCMV) recombinant virus mCMV-⌬IDE, in which the two memory repertoire IDEs, the IE1-derived peptide 168-YPHFMPTNL-176 presented by the major histocom- patibility complex class I (MHC-I) molecule Ld and the m164-derived peptide 257-AGPPRYSRI-265 presented d by the MHC-I molecule D , are both functionally deleted. -

Deficiency of Complement Component 5 Ameliorates Glaucoma in DBA/2J Mice
Rowan University Rowan Digital Works Faculty Scholarship for the College of Science & Mathematics College of Science & Mathematics 6-27-2013 Deficiency of complement component 5 ameliorates glaucoma in DBA/2J mice Gareth R. Howell Ileana Soto Reyes Rowan University, [email protected] Margaret Ryan Leah C. Graham Richard S. Smith See next page for additional authors Follow this and additional works at: https://rdw.rowan.edu/csm_facpub Part of the Neuroscience and Neurobiology Commons Recommended Citation Howell, G. R., Soto, I., Ryan, M., Graham, L. C., Smith, R. S., & John, S. W. (2013). Deficiency of complement component 5 ameliorates glaucoma in DBA/2J mice. Journal of Neuroinflammation, 10, 76-2094-10-76. This Article is brought to you for free and open access by the College of Science & Mathematics at Rowan Digital Works. It has been accepted for inclusion in Faculty Scholarship for the College of Science & Mathematics by an authorized administrator of Rowan Digital Works. Authors Gareth R. Howell, Ileana Soto Reyes, Margaret Ryan, Leah C. Graham, Richard S. Smith, and Simon W.M. John This article is available at Rowan Digital Works: https://rdw.rowan.edu/csm_facpub/24 Howell et al. Journal of Neuroinflammation 2013, 10:76 JOURNAL OF http://www.jneuroinflammation.com/content/10/1/76 NEUROINFLAMMATION RESEARCH Open Access Deficiency of complement component 5 ameliorates glaucoma in DBA/2J mice Gareth R Howell1*†, Ileana Soto1†, Margaret Ryan1, Leah C Graham1, Richard S Smith1 and Simon WM John1,2,3* Abstract Background: Glaucoma is an age-related neurodegenerative disorder involving the loss of retinal ganglion cells (RGCs), which results in blindness. -
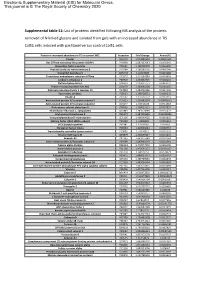
Supplemental Table S1
Electronic Supplementary Material (ESI) for Molecular Omics. This journal is © The Royal Society of Chemistry 2020 Supplemental table S1: List of proteins identified following MS analysis of the proteins removed of N-linked glycans and isolated from gels with an increased abundance in TIS Cal51 cells induced with paclitaxel versus control Cal51 cells. Protein in increased abundance in TIS vs control WCL Accession Fold Change Anova (P) Plectin Q15149 1.073855593 0.00691631 Ras GTPase-activating-like protein IQGAP1 P46940 1.087337643 0.0176342 Elongation factor1-gamma P26641 1.138709703 0.0116496 Peptidyl-prolyl cis-transisomerase B P23284 1.188383105 0.0436246 Dipeptidyl peptidase 3 Q9NY33 1.20163605 0.0215448 Transitional endoplasmic reticulum ATPase P55072 1.214194884 0.0449691 Carbonic anhydrase 2 P00918 1.232852325 0.0158141 Clathrin heavy chain 1 Q00610 1.239621773 0.0463237 Protein transport protein Sec 31A O94979 1.263565104 0.0284155 Aldo-ketoreductase family 1 member C1 Q04828 1.282092186 0.0324406 Spermidine synthase P19623 1.298728621 0.0196232 Plastin-3 P13797 1.310756772 0.0161319 Actin-related protein 2/3 complex subunit 5 O15511 1.333483524 0.00476923 Actin-related protein 2/3 complex subunit 2 O15144 1.35416168 0.0411018 Proteasome subunit alpha type-5 P28066 1.358015551 0.0337657 Thioredoxin reductase 1, cytoplasmic Q16881 1.383670089 0.0235472 Acyl-protein thioesterase 2 O95372 1.387415589 0.00233899 Isoaspartylpeptidase/L-asparaginase Q7L266 1.408149002 0.0319602 Splicing factor U2AF 65kDa subunit P26368 1.41489991 0.0256619 -

PLATELIA™ TOXO Igg AVIDITY 72842
PLATELIA™ TOXO IgG AVIDITY 48 72842 DETERMINATION OF ANTI-TOXOPLASMA GONDII IgG AVIDITY IN HUMAN SERUM BY ENZYME IMMUNOASSAY 881130 - 2013/11 Table of Content 1- INTENDED USE ...............................................................................3 2- CLINICAL VALUE ............................................................................3 3- PRINCIPLE ......................................................................................4 4- PRODUCT INFORMATION ..............................................................5 5- WARNINGS AND PRECAUTIONS ..................................................5 6- SAMPLES ........................................................................................7 7- ASSAY PROCEDURE ......................................................................7 8- INTERPRETATION OF RESULTS .................................................10 9- PERFORMANCES .........................................................................11 10- LIMITATIONS OF THE PROCEDURE ...........................................13 11- QUALITY CONTROL OF THE MANUFACTURER .........................13 12- REFERENCES ...............................................................................14 2 [EN] 1. INTENDED USE Platelia™ TOXO IgG AVIDITY is an immuno-enzyme assay for determination of avidity of anti-T. gondii IgG antibodies in human serum. The Platelia™ TOXO IgG AVIDITY assay should be used in association with the Platelia™ TOXO IgG kit (Ref. 72840). 2. CLINICAL VALUE T. gondii is a protozoan causing infection in numerous -
Enzyme Immunoassays for the Diagnostics of Toxoplasmosis
INFECTIOUS SEROLOGY – PARASITOLOGY – Toxoplasma gondii FOLLOW US FOLLOW US Toxoplasma gondii Enzyme immunoassays for the diagnostics of toxoplasmosis ELISA and IMMUNOBLOT kits are optimized and validated for detection of IgA, IgE, IgG and IgM antibodies in human serum and plasma BIOVENDOR.GROUP INFECTIOUS SEROLOGY – PARASITOLOGY – Toxoplasma gondii Introduction Diagnosis of infection Toxoplasmosis is a widespread parasitic disease cau- Diagnosis of the disease is based on epidemiological sed by protozoan Toxoplasma gondii – a parasite with anamnesis, clinical manifestation and laboratory tests. a complicated life cycle consisting of several morpho- Direct detection of the parasite is not available for rou- logically different stadia. Primary hosts are members of tine diagnostics. Serology is the most important tool for the feline family. Humans and most warm-blooded ani- laboratory diagnostics of toxoplasmosis. mals can be infected by either primarily infected food – Screening – determination of total antibodies by (insufficiently heat-treated meat) or by ingestion of oo- complement fixation test (CFT) cysts (secondary contaminated food or contaminated – Determination of specific IgA, IgE, IgM, IgG fingers, objects, etc.). antibodies and IgG avidity by ELISA and confirmation Acquired toxoplasmosis in immunocompetent indi- of results by Immunoblot viduals is usually asymptomatic or can manifest itself with flu-like symptoms (subfebrility, fatigue, lymphade- nopathy, muscle aches) and has no lasting ill effects. Severe life-threatening infections (encephalitis, hepati- tis, chorioretinitis, myocarditis, generalized form of the disease) may develop in immunocompromised patients usually because of a reactivation of a latent infection. Congenital toxoplasmosis is caused by transmission of infection from mother to foetus and it might result in severe damages of the foetus (brain calcification, hydrocepha- lus, vision disorders, mental affections), still birth or abortion. -

In Kidney Injury
antioxidants Review Regulation of Complement Activation by Heme Oxygenase-1 (HO-1) in Kidney Injury Maria G. Detsika 1,* and Elias A. Lianos 2,3 1 First Department of Critical Care Medicine & Pulmonary Services, GP Livanos and M. Simou Laboratories, National & Kapodistrian University of Athens, Medical School, Evangelismos Hospital, 10675 Athens, Greece 2 Thorax Foundation, Research Center of Intensive Care and Emergency Thoracic Medicine, 10675 Athens, Greece; [email protected] 3 Veterans Affairs Medical Center and Virginia Tech, Carilion School of Medicine, 1970 Roanoke Blvd, Salem, VA 24153, USA * Correspondence: [email protected]; Tel.: +30-210-723552; Fax: +30-210-7239127 Abstract: Heme oxygenase is a cytoprotective enzyme with strong antioxidant and anti-apoptotic properties. Its cytoprotective role is mainly attributed to its enzymatic activity, which involves the degradation of heme to biliverdin with simultaneous release of carbon monoxide (CO). Recent studies uncovered a new cytoprotective role for heme oxygenase-1 (HO-1) by identifying a regulatory role on the complement control protein decay-accelerating factor. This is a key complement regulatory protein preventing dysregulation or overactivation of complement cascades that can cause kidney injury. Cell-specific targeting of HO-1 induction may, therefore, be a novel approach to attenuate complement-dependent forms of kidney disease. Keywords: heme; heme oxygenase-1 (HO-1); complement; kidney injury 1. Introduction Citation: Detsika, M.G.; Lianos, E.A. Although the role of heme, in various cellular processes, such as gene transcription Regulation of Complement and translation and cellular differentiation, proliferation, and apoptosis, has been known Activation by Heme Oxygenase-1 for decades, the role of the heme-degrading enzyme heme oxygenase-1 (HO-1) only gained (HO-1) in Kidney Injury. -

Complement Expression in the Retina Is Not Influenced by Short-Term
City University of New York (CUNY) CUNY Academic Works Publications and Research Kingsborough Community College 2014 Complement Expression in the Retina is not Influenced yb Short- term Pressure Elevation Konstantin Astafurov SUNY Downstate Medical Center Cecilia Q. Dong SUNY Downstate Medical Center Lampros Panagis SUNY Downstate Medical Center Gautam Kamtham Mount Sinai School of Medicine Lizhen Ren SUNY Downstate Medical Center See next page for additional authors How does access to this work benefit ou?y Let us know! More information about this work at: https://academicworks.cuny.edu/kb_pubs/46 Discover additional works at: https://academicworks.cuny.edu This work is made publicly available by the City University of New York (CUNY). Contact: [email protected] Authors Konstantin Astafurov, Cecilia Q. Dong, Lampros Panagis, Gautam Kamtham, Lizhen Ren, Anna Rozenboym, Tarique D. Perera, Jeremy D. Coplan, and John Danias This article is available at CUNY Academic Works: https://academicworks.cuny.edu/kb_pubs/46 Molecular Vision 2014; 20:140-152 <http://www.molvis.org/molvis/v20/140> © 2014 Molecular Vision Received 25 February 2013 | Accepted 28 January 2014 | Published 31 January 2014 Complement expression in the retina is not influenced by short- term pressure elevation Konstantin Astafurov,1 Cecilia Q. Dong,2 Lampros Panagis,1 Gautam Kamthan,3 Lizhen Ren,1 Anna Rozenboym,4 Tarique D. Perera,5 Jeremy D. Coplan,6 John Danias1,2 1Department of Cell Biology, SUNY Downstate Medical Center, Brooklyn, NY; 2Department of Ophthalmology, SUNY Downstate Medical Center, Brooklyn, NY; 3Department of Ophthalmology, Mount Sinai School of Medicine, New York, NY; 4Department of Biological Sciences, CUNY Kingsborough Community College, Brooklyn, NY; 5Department of Psychiatry, Columbia University Medical Center and New York State Psychiatric Institute, New York, NY; 6Department of Psychiatry, SUNY Downstate Medical Center, Brooklyn, NY Purpose: To determine whether short-term pressure elevation affects complement gene expression in the retina in vitro and in vivo.