Structural Characterization of Bacterial Defense Complex Marko Nedeljković
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Synthesis and Biological Evaluation of Trisindolyl-Cycloalkanes and Bis- Indolyl Naphthalene Small Molecules As Potent Antibacterial and Antifungal Agents
Synthesis and Biological Evaluation of Trisindolyl-Cycloalkanes and Bis- Indolyl Naphthalene Small Molecules as Potent Antibacterial and Antifungal Agents Dissertation Zur Erlangung des akademischen Grades doctor rerum naturalium (Dr. rer. nat.) Vorgelegt der Naturwissenschaftlichen Fakultät I Institut für Pharmazie Fachbereich für Pharmazeutische Chemie der Martin-Luther-Universität Halle-Wittenberg von Kaveh Yasrebi Geboren am 09.14.1987 in Teheran/Iran (Islamische Republik) Gutachter: 1. Prof. Dr. Andreas Hilgeroth (Martin-Luther-Universität Halle-Wittenberg, Germany) 2. Prof. Dr. Sibel Süzen (Ankara Üniversitesi, Turkey) 3. Prof. Dr. Michael Lalk (Ernst-Moritz-Arndt-Universität Greifswald, Germany) Halle (Saale), den 21. Juli 2020 Selbstständigkeitserklärung Hiermit erkläre ich gemäß § 5 (2) b der Promotionsordnung der Naturwissenschaftlichen Fakultät I – Institut für Pharmazie der Martin-Luther-Universität Halle-Wittenberg, dass ich die vorliegende Arbeit selbstständig und ohne Benutzung anderer als der angegebenen Hilfsmittel und Quellen angefertigt habe. Alle Stellen, die wörtlich oder sinngemäß aus Veröffentlichungen entnommen sind, habe ich als solche kenntlich gemacht. Ich erkläre ferner, dass diese Arbeit in gleicher oder ähnlicher Form bisher keiner anderen Prüfbehörde zur Erlangung des Doktorgrades vorgelegt wurde. Halle (Saale), den 21. Juli 2020 Kaveh Yasrebi Acknowledgement This study was carried out from June 2015 to July 2017 in the Research Group of Drug Development and Analysis led by Prof. Dr. Andreas Hilgeroth at the Institute of Pharmacy, Martin-Luther-Universität Halle-Wittenberg. I would like to thank all the people for their participation who supported my work in this way and helped me obtain good results. First of all, I would like to express my gratitude to Prof. Dr. Andreas Hilgeroth for providing me with opportunity to carry out my Ph.D. -

Novel Antimicrobial Agents Inhibiting Lipid II Incorporation Into Peptidoglycan Essay MBB
27 -7-2019 Novel antimicrobial agents inhibiting lipid II incorporation into peptidoglycan Essay MBB Mark Nijland S3265978 Supervisor: Prof. Dr. Dirk-Jan Scheffers Molecular Microbiology University of Groningen Content Abstract..............................................................................................................................................2 1.0 Peptidoglycan biosynthesis of bacteria ........................................................................................3 2.0 Novel antimicrobial agents ...........................................................................................................4 2.1 Teixobactin ...............................................................................................................................4 2.2 tridecaptin A1............................................................................................................................7 2.3 Malacidins ................................................................................................................................8 2.4 Humimycins ..............................................................................................................................9 2.5 LysM ........................................................................................................................................ 10 3.0 Concluding remarks .................................................................................................................... 11 4.0 references ................................................................................................................................. -

Substrate Specificities of Outer Membrane Proteases of the Omptin Family in Escherichia Coli, Salmonella Enterica, and Citrobacter Rodentium
Substrate specificities of outer membrane proteases of the omptin family in Escherichia coli, Salmonella enterica, and Citrobacter rodentium Andrea Portt, McGill University, Montreal October 9th, 2010 A thesis submitted to McGill University in partial fulfillment of the requirements of the degree of Master’s in Microbiology © Andrea Portt 2010 Table of Contents LIST OF ABBREVIATIONS 3 ABSTRACT 6 ACKNOWLEDGEMENTS 7 CONTRIBUTIONS OF AUTHORS 8 LITERATURE REVIEW 9 2. OMPTINS 9 3. OMPTIN SUBSTRATES 13 A. ANTIMICROBIAL PEPTIDES 13 B. BLOOD CLOTTING PROTEINS AND EXTRACELLULAR MATRIX 14 C. COMPLEMENT 16 D. TROPOMYOSIN 17 4. EVOLUTION OF OMPTINS 17 5. OMPTIN REGULATION 20 6. OMPTINS OF PATHOGENIC ENTEROBACTEREACEAE 22 A. Y. PESTIS 22 B. S. ENTERICA 23 C. SHIGELLA 25 D. ATTACHING AND EFFACING BACTERIA 25 INTRODUCTION 28 MATERIALS AND METHODS 29 1. BACTERIAL GROWTH 29 2. BACTERIAL STRAINS AND CONSTRUCTION OF PLASMIDS 29 A. STRAINS 29 B. PLASMIDS 31 3. DISK INHIBITION ASSAYS 34 4. MINIMUM INHIBITORY CONCENTRATION DETERMINATIONS 34 5. PROTEOLYTIC CLEAVAGE OF AMPS BY CELLS EXPRESSING OMPTINS 34 6. OM DISRUPTION ASSAY WITH 1-N-PHENYLNAPHTHYLAMINE 35 7. DEGRADATION OF TROPOMYOSIN BY E. COLI AND C. RODENTIUM STRAINS EXPRESSING OMPTINS 35 8. REAL-TIME QUANTITATIVE PCR 35 9. PURIFICATION OF NATIVE CROP 35 10. EXPRESSION, PURIFICATION, AND REFOLDING OF HIS-PGTE AND HIS-CROP 36 A. EXPRESSION 36 B. PURIFICATION 36 C. REFOLDING 36 11. CLEAVAGE OF C2 FRET SUBSTRATE BY WHOLE CELLS OR PURIFIED OMPTINS 37 A. WHOLE CELLS 37 B. PURIFIED OMPTINS 37 1 RESULTS 38 1. GROWTH INHIBITION OF E. COLI AND C. RODENTIUM STRAINS EXPRESSING OMPTINS BY C18G. -
Parsek Micro410 Lecture
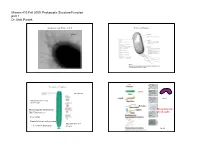
Microm 410 Fall 2009: Prokaryotic Structure/Function part 1 Dr. Matt Parsek Organization of the Prokaryotic Cell Prokaryotic Structures fimbriae Size Range of Prokaryotes bacillus See Table 4.1 (rigid) vibrio Nanobacteria 0.05‐0.2 µm (0.14‐0.2 µm) (flexible) Thiomargarita namibiensis Mycoplasma are 700‐750 µm (Fig. 4.2) pleomorphic green alga Nanochlorum eukaryotum Mycoplasma 0.1‐ ~1-2 µm in diameter 0.3 µm Fig. 4.1 Microm 410 Fall 2009: Prokaryotic Structure/Function part 1 Dr. Matt Parsek Staining cells for Microscopic observation Cell Arrangements Motility- ~80% of prokaryotes are motile streptococcus Staining properties: Gram Stain staphylococcus Fig. 2.3 Gram Stain (1884) (Bacteria) Gram-negative mixed culture Gram-positive Fig. 2.3 and 2.4 Microm 410 Fall 2009: Prokaryotic Structure/Function part 1 Dr. Matt Parsek Functions of the cytoplasmic membrane The phospholipid bi‐layer Fig. 4.9 Fig. 4.4 What is the structure of bacterial phospholipids? Other components of the cytoplasmic membrane Figs. 4.5‐4.6 Microm 410 Fall 2009: Prokaryotic Structure/Function part 1 Dr. Matt Parsek Archaeal membranes can be a lipid monolayer Archaeal phospholipids have an ether linkage Fig. 4.7 Fig. 4.8 Importance of Cell Wall Schematic diagram cell wall • Provides rigidity to cell allowing cell to withstand the large osmotic/ionic Fig. 4.16 changes a bacterium may experience in its environment, and turgor pressure of cytoplasm (conc. of solutes in cytoplasm). Cell lysis • May have a role in shape determination. • Provides a barrier against certain toxic chemical and biological agents. • Site of action of some of the most commonly used antibiotics used to treat bacterial infections (penicillin family). -

Harnessing the Power of Bacteria in Advancing Cancer Treatment
International Journal of Molecular Sciences Review Microbes as Medicines: Harnessing the Power of Bacteria in Advancing Cancer Treatment Shruti S. Sawant, Suyash M. Patil, Vivek Gupta and Nitesh K. Kunda * Department of Pharmaceutical Sciences, College of Pharmacy and Health Sciences, St. John’s University, Jamaica, NY 11439, USA; [email protected] (S.S.S.); [email protected] (S.M.P.); [email protected] (V.G.) * Correspondence: [email protected]; Tel.: +1-718-990-1632 Received: 20 September 2020; Accepted: 11 October 2020; Published: 14 October 2020 Abstract: Conventional anti-cancer therapy involves the use of chemical chemotherapeutics and radiation and are often non-specific in action. The development of drug resistance and the inability of the drug to penetrate the tumor cells has been a major pitfall in current treatment. This has led to the investigation of alternative anti-tumor therapeutics possessing greater specificity and efficacy. There is a significant interest in exploring the use of microbes as potential anti-cancer medicines. The inherent tropism of the bacteria for hypoxic tumor environment and its ability to be genetically engineered as a vector for gene and drug therapy has led to the development of bacteria as a potential weapon against cancer. In this review, we will introduce bacterial anti-cancer therapy with an emphasis on the various mechanisms involved in tumor targeting and tumor suppression. The bacteriotherapy approaches in conjunction with the conventional cancer therapy can be effective in designing novel cancer therapies. We focus on the current progress achieved in bacterial cancer therapies that show potential in advancing existing cancer treatment options and help attain positive clinical outcomes with minimal systemic side-effects. -

Killing of Gram-Negative Bacteria by Lactoferrin and Lysozyme
Killing of gram-negative bacteria by lactoferrin and lysozyme. R T Ellison 3rd, T J Giehl J Clin Invest. 1991;88(4):1080-1091. https://doi.org/10.1172/JCI115407. Research Article Although lactoferrin has antimicrobial activity, its mechanism of action is not full defined. Recently we have shown that the protein alters the Gram-negative outer membrane. As this membrane protects Gram-negative cells from lysozyme, we have studied whether lactoferrin's membrane effect could enhance the antibacterial activity of lysozyme. We have found that while each protein alone is bacteriostatic, together they can be bactericidal for strains of V. cholerae, S. typhimurium, and E. coli. The bactericidal effect is dose dependent, blocked by iron saturation of lactoferrin, and inhibited by high calcium levels, although lactoferrin does not chelate calcium. Using differing media, the effect of lactoferrin and lysozyme can be partially or completely inhibited; the degree of inhibition correlating with media osmolarity. Transmission electron microscopy shows that E. coli cells exposed to lactoferrin and lysozyme at 40 mOsm become enlarged and hypodense, suggesting killing through osmotic damage. Dialysis chamber studies indicate that bacterial killing requires direct contact with lactoferrin, and work with purified LPS suggests that this relates to direct LPS-binding by the protein. As lactoferrin and lysozyme are present together in high levels in mucosal secretions and neutrophil granules, it is probable that their interaction contributes to host defense. Find the latest version: https://jci.me/115407/pdf Killing of Gram-negative Bacteria by Lactofernn and Lysozyme Richard T. Ellison III*" and Theodore J. Giehl *Medical and tResearch Services, Department of Veterans Affairs Medical Center, and Division ofInfectious Diseases, Department ofMedicine, University ofColorado School ofMedicine, Denver, Colorado 80220 Abstract (4). -

Molecular Analysis of Candidate Probiotic Effector Molecules of Lactobacillus Plantarum
Molecular analysis of candidate probiotic effector molecules of Lactobacillus plantarum Daniela Maria Remus Thesis committee Promoter Prof. dr. Michiel Kleerebezem Professor of Bacterial Metagenomics Wageningen University Co-promoter Dr. Peter A. Bron Senior Scientist, NIZO food research BV, Ede Other members Prof. dr. Tjakko Abee, Wageningen University Prof. dr. Pascal Hols, University of Lovain (Lovain la Neuve), Belgium Dr. Philippe Langella, National Institute of Agricultural Research (INRA), Paris, France Prof. dr. Roland J. Siezen, Radboud University, Nijmegen This research was conducted under the auspices of the Graduate School VLAG (Advanced studies in Food Technology, Agrobiotechnology, Nutrition and Health Sciences). Molecular analysis of candidate probiotic effector molecules of Lactobacillus plantarum Daniela Maria Remus Thesis submitted in fulfillment of the requirements for the degree of doctor at Wageningen University by the authority of the Rector Magnificus Prof. dr. M.J. Kropff, in the presence of the Thesis Committee appointed by the Academic Board to be defended in public on Tuesday 09 October 2012 at 4 p.m. in the Aula. Daniela Maria Remus Molecular analysis of candidate probiotic effector molecules of Lactobacillus plantarum Ph.D. Thesis, Wageningen University, Wageningen, The Netherlands (2012) With references and summaries in English and Dutch. ISBN 978-94-6173-373-3 Summary Lactobacilli occupy diverse natural habitats, including dairy products and the mammalian gastrointestinal (GI) tract, and several Lactobacillus strains are marketed as probiotics that can interact with host cells in the GI tract by which they are proposed to beneficially influence the health status of their consumers. The discovery of probiotic effector molecules is instrumental to understand the exact modes of probiotic action, which is required for their controlled, safe, and purpose-directed application. -

Plantaricin KW30
The Molecular and Cellular Characterisation of the First Glycocin: Plantaricin KW30 Judith Stepper 2009 The Molecular and Cellular Characterisation of the First Glycocin: Plantaricin KW30 A thesis presented in partial fulfillment of the requirements for the degree of Doctor of Philosophy in Biochemistry at Massey University, Palmerston North, New Zealand Judith Stepper 2009 “What we know is a drop. What we don’t know is an ocean.” Isaac Newton (1643 – 1727) ABSTRACT Bacteriocins, typically secreted by Gram-positive and -negative bacteria, are ribosomally- synthesised antimicrobial peptides which inhibit the growth of competing bacteria. We have purified a 43 amino acid bacteriocin, plantaricin KW30 (PlnKW30) produced by Lactobacillus plantarum KW30, that has little amino acid sequence similarity to any other characterised bacteriocin. The gene encoding plnKW30 is in a cluster with the genes required for maturation and export of, and immunity to, the bacteriocin. This arrangement of genes is similar to the genomic context of bacteriocin genes in other lactic acid bacteria. The plnKW30 gene cluster comprises six genes encoding a glycosyltransferase, a proteolytic ABC-transporter, two putative thioredoxins, a response regulator and PlnKW30 itself. PlnKW30 was found to possess two unusual post-translational modifications: an O-glycosylated serine and an unprecedented S-glycosylation of the C-terminal cysteine. The modified serine is located on an eight residue loop that is tethered by a disulfide bridge. Both modifications have been identified as N-acetylglucosamines (GlcNAc), making PlnKW30 the first described class IV bacteriocin. A post-translational modification with S-linked GlcNAc is unprecedented in bacteriocins as well as in all genera. -

Detailed Structure and Pathophysiological Roles of the Iga-Albumin Complex in Multiple Myeloma
International Journal of Molecular Sciences Article Detailed Structure and Pathophysiological Roles of the IgA-Albumin Complex in Multiple Myeloma Yuki Kawata 1, Hisashi Hirano 1, Ren Takahashi 1, Yukari Miyano 1, Ayuko Kimura 1, Natsumi Sato 1, Yukio Morita 2, Hirokazu Kimura 1,* and Kiyotaka Fujita 1 1 Department of Health Sciences, Gunma Paz University Graduate School of Health Sciences, 1-7-1, Tonyamachi, Takasaki-shi, Gunma 370-0006, Japan; [email protected] (Y.K.); [email protected] (H.H.); [email protected] (R.T.); [email protected] (Y.M.); [email protected] (A.K.); [email protected] (N.S.); [email protected] (K.F.) 2 Laboratory of Public Health II, Azabu University School of Veterinary Medicine, 1-17-71, Fuchinobe, Chuo-ku, Sagamihara, Kanagawa 252-5201, Japan; [email protected] * Correspondence: [email protected]; Tel.: +81-27-365-3366; Fax: +81-27-388-0386 Abstract: Immunoglobulin A (IgA)-albumin complexes may be associated with pathophysiology of multiple myeloma, although the etiology is not clear. Detailed structural analyses of these protein– protein complexes may contribute to our understanding of the pathophysiology of this disease. We analyzed the structure of the IgA-albumin complex using various electrophoresis, mass spectrom- etry, and in silico techniques. The data based on the electrophoresis and mass spectrometry showed that IgA in the sera of patients was dimeric, linked via the J chain. Only dimeric IgA can bind to albumin molecules leading to IgA-albumin complexes, although both monomeric and dimeric forms of IgA were present in the sera. -

The Bacteriostatic Diglycocylated Bacteriocin Glycocin F Targets A
Copyright is owned by the Author of the thesis. Permission is given for a copy to be downloaded by an individual for the purpose of research and private study only. The thesis may not be reproduced elsewhere without the permission of the Author. The Bacteriostatic Diglycosylated Bacteriocin Glycocin F Targets a Sugar-Specific Transporter A thesis presented in partial fulfilment of the requirements for the degree of Master of Science in Biochemistry at Massey University, Manawatu New Zealand Kelvin Ross Drower 2014 Dedicated to Nana and Pop Abstract The increasing prevalence of antibiotic-resistance bacteria is threatening to end the antibiotic era established following Alexander Fleming's discovery of penicillin in 1928. Over-prescription and misuse of broad-spectrum antibiotics has hastened the development and spread of antibiotic resistance. This, combined with a lack of research and development (R&D) of new antibiotics by major pharmaceutical companies, may lead to a widespread recurrence of 'incurable' bacterial diseases. However while commercial R&D of antibiotics has waned, much research has been carried out to characterise bacteriocins, ribosomally-synthesised antimicrobial polypeptides thought to be produced by virtually all prokaryotes. Although hundreds of bacteriocins have been identified and characterised, only a handful of their cognate receptors on susceptible cells have been identified. Glycocin F is a bacteriostatic diglycosylated 43-amino acid bacteriocin produced by the Gram-positive bacterium Lactobacillus plantarum KW30 that inhibits the growth of a broad range of bacteria. The mechanism of action of glycocin F is unknown, however evidence suggested that glycocin F binds to cells via a N-acetylglucosamine (GlcNAc) specific phosphoenolpyruvate:carbohydrate-phosphotransferase system (PTS) transporter, as had been shown for lactococcin A, lactococcin B and microcin E492 that target a mannose specific PTS transporter. -

Of Penicillin G, Other Penicillins, and a Cephalosporin
GLYCOPEPTIDE TRANSPEPTIDASE AND D-ALANINE CARBOXYPEPTIDASE: PENICILLIN-SENSITIVE ENZYMATIC REACTIONS* BY KAZUO IZAKI, M\ICHIO M\ATSUHASHI, AND JACK L. STROMINGER DEPARTMENT OF PHARMACOLOGY, UNIVERSITY OF WISCONSIN MEDICAL SCHOOL, MADISON Communicated by Clinton L. Woolsey, January 28, 1966 In 1929, Fleming discovered penicillins and observed that this group of substances kills gram-positive bacteria more effectively than gram-negative bacteria (thus im- plying some important physiological difference between the two groups of or- ganisms).1 Subsequent knowledge of the bacterial cell wall made it possible to establish both on physiological and chemical grounds that penicillins are specific and highly selective inhibitors of the biosynthesis of cell walls, both in gram-positive and in gram-negative bacteria.2-6 The reason for the relative insensitivity of gram- negative bacteria to most penicillins has remained obscure. More recent knowledge of the structure and biosynthesis of bacterial cell walls has suggested that, in a terminal reaction in cell wall synthesis, linear glycopeptide strands are cross-linked in a transpeptidation, accompanied by release of the termi- nal D-alanine of the pentapeptide precursor, with formation of a two- or three- dimensional network. Direct chemical analyses of cell walls prepared from cells treated with penicillin7' 8 and isotopic studies of wall biosynthesis9' 10 suggested that penicillin was interfering with this hypothetical glycopeptide cross-linking reaction. In the presence of penicillin, nascent glycopeptide, -

Decision Summary
510(k) SUBSTANTIAL EQUIVALENCE DETERMINATION DECISION SUMMARY A. 510(k) Number: k081249 B. Purpose for Submission: New analyte, controls, and calibrator C. Measurand: Alpha 2 macroglobulin D. Type of Test: Quantitative, Nephelometric E. Applicant: Dade Behring, Inc. F. Proprietary and Established Names: Dimension Vista System A2mac Flex Reagent Cartridge, Dimension Vista System Protein 1 Calibrator, Dimension Vista System. G. Regulatory Information: Regulation Section Classification Product Code Panel 21 CFR 866.5620, Alpha- Class II Alpha-2-Macroglobulin, 82 Immunology 2-macroglobulin Antigen, Antiserum, (IM) immunological test Control (DEB) system. 21 CFR 862.1660, Quality Class I Multi-Analyte Controls, 75 Clinical control material (assayed All Kinds (Assayed) Chemistry and unassayed). (JJY) (CH) 21 CFR 862.1150, Class II Calibrator, Multi- 75 Clinical Calibrator. Analyte Mixture (JIX) Chemistry (CH) H. Intended Use: 1. Intended use(s): Dimension Vista® System A2MAC Flex® Reagent Cartridge The A2MAC assay is an in vitro diagnostic test for the quantitative measurement of alpha2-macroglobulin in human serum and heparinized and EDTA plasma on the Dimension Vista® Systems. Measurements of a2-macroglobulin aid in the diagnosis of blood clotting or blood lysis disorders. Dimension Vista® Protein 1 Calibrator Dimension Vista® Protein 1 Calibrator is an in vitro diagnostic product for the calibration of the Dimension Vista® System for: α1-Acid Glycoprotein (A1AG), α1-Antitrypsin(A1AT), α2-Macroglobulin (A2MAC), β2-Microglobulin (B2MIC), C3 Complement (C3), C4 Complement (C4), Ceruloplasmin(CER), Haptoglobin (HAPT), Hemopexin (HPX), Homocysteine (HCYS), Immunoglobulin A (IGA), Immunoglobulin E (IGE), Immunoglobulin G (IGG, IGG-C*), Immunoglobulin G Subclass 1, (IGG1), Immunoglobulin G Subclass 2 (IGG2), Immunoglobulin G 1 Subclass 3 (IGG3), Immunoglobulin Subclass 4 (IGG4), Immunoglobulin M (IGM), Prealbumin (PREALB), Retinol Binding Protein (RBP), soluble Transferrin Receptor (sTFR) and Transferring (TRF).