SOX11 Interactome Analysis: Implication in Transcriptional Control and Neurogenesis
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Core Transcriptional Regulatory Circuitries in Cancer
Oncogene (2020) 39:6633–6646 https://doi.org/10.1038/s41388-020-01459-w REVIEW ARTICLE Core transcriptional regulatory circuitries in cancer 1 1,2,3 1 2 1,4,5 Ye Chen ● Liang Xu ● Ruby Yu-Tong Lin ● Markus Müschen ● H. Phillip Koeffler Received: 14 June 2020 / Revised: 30 August 2020 / Accepted: 4 September 2020 / Published online: 17 September 2020 © The Author(s) 2020. This article is published with open access Abstract Transcription factors (TFs) coordinate the on-and-off states of gene expression typically in a combinatorial fashion. Studies from embryonic stem cells and other cell types have revealed that a clique of self-regulated core TFs control cell identity and cell state. These core TFs form interconnected feed-forward transcriptional loops to establish and reinforce the cell-type- specific gene-expression program; the ensemble of core TFs and their regulatory loops constitutes core transcriptional regulatory circuitry (CRC). Here, we summarize recent progress in computational reconstitution and biologic exploration of CRCs across various human malignancies, and consolidate the strategy and methodology for CRC discovery. We also discuss the genetic basis and therapeutic vulnerability of CRC, and highlight new frontiers and future efforts for the study of CRC in cancer. Knowledge of CRC in cancer is fundamental to understanding cancer-specific transcriptional addiction, and should provide important insight to both pathobiology and therapeutics. 1234567890();,: 1234567890();,: Introduction genes. Till now, one critical goal in biology remains to understand the composition and hierarchy of transcriptional Transcriptional regulation is one of the fundamental mole- regulatory network in each specified cell type/lineage. -

Table 2. Significant
Table 2. Significant (Q < 0.05 and |d | > 0.5) transcripts from the meta-analysis Gene Chr Mb Gene Name Affy ProbeSet cDNA_IDs d HAP/LAP d HAP/LAP d d IS Average d Ztest P values Q-value Symbol ID (study #5) 1 2 STS B2m 2 122 beta-2 microglobulin 1452428_a_at AI848245 1.75334941 4 3.2 4 3.2316485 1.07398E-09 5.69E-08 Man2b1 8 84.4 mannosidase 2, alpha B1 1416340_a_at H4049B01 3.75722111 3.87309653 2.1 1.6 2.84852656 5.32443E-07 1.58E-05 1110032A03Rik 9 50.9 RIKEN cDNA 1110032A03 gene 1417211_a_at H4035E05 4 1.66015788 4 1.7 2.82772795 2.94266E-05 0.000527 NA 9 48.5 --- 1456111_at 3.43701477 1.85785922 4 2 2.8237185 9.97969E-08 3.48E-06 Scn4b 9 45.3 Sodium channel, type IV, beta 1434008_at AI844796 3.79536664 1.63774235 3.3 2.3 2.75319499 1.48057E-08 6.21E-07 polypeptide Gadd45gip1 8 84.1 RIKEN cDNA 2310040G17 gene 1417619_at 4 3.38875643 1.4 2 2.69163229 8.84279E-06 0.0001904 BC056474 15 12.1 Mus musculus cDNA clone 1424117_at H3030A06 3.95752801 2.42838452 1.9 2.2 2.62132809 1.3344E-08 5.66E-07 MGC:67360 IMAGE:6823629, complete cds NA 4 153 guanine nucleotide binding protein, 1454696_at -3.46081884 -4 -1.3 -1.6 -2.6026947 8.58458E-05 0.0012617 beta 1 Gnb1 4 153 guanine nucleotide binding protein, 1417432_a_at H3094D02 -3.13334396 -4 -1.6 -1.7 -2.5946297 1.04542E-05 0.0002202 beta 1 Gadd45gip1 8 84.1 RAD23a homolog (S. -

The Role of Transcription Factor in the Regulation of Autoimmune Diseases
Chiba Medical J. 97E:17-24, 2021 doi:10.20776/S03035476-97E-2-P17 〔 Chiba Medical Society Award Review 〕 The role of transcription factor in the regulation of autoimmune diseases Akira Suto1,2) 1) Department of Allergy and Clinical Immunology, Graduate School of Medicine, Chiba 260-8670. 2) Institute for Global Prominent Research, Chiba University, Chiba 260-8670. (Received November 14, 2020, Accepted December 9, 2020, Published April 10, 2021.) Abstract IL-21 is produced by Th17 cells and follicular helper T cells. It is an autocrine growth factor and plays critical roles in autoimmune diseases. In autoimmune mouse models, excessive production of IL-21 is associated with the development of lupus-like pathology in Sanroque mice, diabetes in NOD mice, autoimmune lung inflammation in Foxp3-mutant scurfy mice, arthritis in collagen-induced arthritis, and muscle inflammation in experimental autoimmune myositis. IL-21 production is induced by the transcription factor c-Maf following Stat3 activation on stimulation with IL-6. Additionally, Stat3 signaling induces the transcription factor Sox5 that along with c-Maf directly activates the promoter of RORγt, a master regulator of Th17 cells. Moreover, T cell-specific Sox5-deficient mice exhibit decreased Th17 cell differentiation and resistance to experimental autoimmune encephalomyelitis and delayed- type hypersensitivity. Another Sox family gene, Sox12, is expressed in regulatory T cells( Treg) in dextran sulfate sodium-induced colitic mice. T cell receptor-NFAT signaling induces Sox12 expression that further promotes Foxp3 expression in CD4+ T cells. In vivo, Sox12 is involved in the development of peripherally induced Treg cells under inflammatory conditions in an adoptive transfer colitis model. -

Phox2b and Motoneuronal Differentiation
Development 127, 1349-1358 (2000) 1349 Printed in Great Britain © The Company of Biologists Limited 2000 DEV1515 Control of hindbrain motor neuron differentiation by the homeobox gene Phox2b Alexandre Pattyn, Marie-Rose Hirsch, Christo Goridis and Jean-François Brunet* Laboratoire de Génétique et Physiologie du Développement, Developmental Biology Institute of Marseille, CNRS/INSERM/Univ Méditerranée/AP de Marseille, Luminy Case 907, 13288 Marseille Cedex 9, France *Author for correspondence (e-mail: [email protected]) Accepted 11 January; published on WWW 7 March 2000 SUMMARY Motor neurons are a widely studied model of vertebrate both the generic and subtype-specific programs of neurogenesis. They can be subdivided in somatic, branchial motoneuronal differentiation are disrupted at an early and visceral motor neurons. Recent studies on the stage. Most motor neuron precursors die inside the dorsoventral patterning of the rhombencephalon have neuroepithelium while those that emigrate to the mantle implicated the homeobox genes Pax6 and Nkx2.2 in the layer fail to switch on early postmitotic markers and to early divergence of the transcriptional programme of downregulate neuroepithelial markers. Thus, the loss of hindbrain somatic and visceral motor neuronal function of Phox2b in hindbrain motor neurons exemplifies differentiation. We provide genetic evidence that the a novel control point in the generation of CNS neurons. paired-like homeodomain protein Phox2b is required for the formation of all branchial and visceral, but not somatic, Key words: Neurogenesis, Motor neuron, Homeobox gene, motor neurons in the hindbrain. In mice lacking Phox2b, Rhombencephalon, Phox2b, Mouse INTRODUCTION of sm neurons has been studied most extensively and some of the transcriptional cascades controlling their differentiation are The specification of neuronal subtype identity in the beginning to be understood. -

The Evolving Genetic Landscape of Hirschprung Disease: an Update and Review
Review Article Clinics in Surgery Published: 11 Oct, 2017 The Evolving Genetic Landscape of Hirschprung Disease: An Update and Review Amit Kumar Yadav* and Gaurav Chopra Department of Pathology, Vardhman Mahavir Medical College and Safdarjung Hospital, New Delhi, India Abstract Hirschsprung Disease (HD) is a developmental disorder characterized by the complete absence of ganglion cells in the distal gastrointestinal tract. It is the most common cause of functional intestinal obstruction in neonates and children. The aganglionosis is believed to be either due to failure of neural crest cells to migrate, proliferate, differentiate or survive during gut development in the embryonic stage. The incidence of HD is estimated at 1/5000 live births and shows a male predominance. It is usually sporadic, although it can be familial and may be inherited as autosomal dominant or autosomal recessive. In 70% of cases, HD occurs as an isolated trait and in the other 30% it is associated with other congenital malformation syndromes. HD has a complex multifactorial etiology, and genetic factors play a key role in its pathogenesis. Several gene loci appear to be involved. Many of these have been identified by conducting Genome Wide Association (GWAS) studies and recently by Next Generation Sequencing (NGS). These genes encode for receptors, ligands (especially those participating in the RET, EDNRB and Semaphorin signaling pathways), transcriptional factors (PHOX2B & SOX10). These genes are involved in the neural crest cell development and migration that give rise to ganglion cells. Overall, the RET proto-oncogene is considered the major disease causing gene in HD. A greater understanding of the genetic landscape of this disease might pave way for genetic counseling, prenatal and preimplantation diagnosis in the management of HD. -

HAND2 Synergistically Enhances Transcription of Dopamine-ß
Available online at www.sciencedirect.com R Developmental Biology 262 (2003) 183–193 www.elsevier.com/locate/ydbio HAND2 synergistically enhances transcription of dopamine-- hydroxylase in the presence of Phox2a Haiming Xu,a Anthony B. Firulli,b Xiaotong Zhang,a and Marthe J. Howarda,* a Department of Anatomy and Neurobiology, Medical College of Ohio, 3000 Arlington Ave., Toledo, OH 43614, USA b Wells Center for Pediatric Research, James Whitcomb Riley Hospital for Children, 702 Barnhill Drive, Room 2666, Indianapolis, IN 46202-5225, USA Received for publication 28 March 2003, revised 5 June 2003, accepted 5 June 2003 Abstract Noradrenergic neuronal identity and differentiation are controlled by cascades of transcription factors acting downstream of BMP4, including the basic helix–loop–helix DNA binding protein HAND2 and the homeodomain factor Phox2a. Dopamine--hydroxylase (DBH) is the penultimate enzyme required for synthesis of norepinephrine and is thus a noradrenergic cell type-specific marker. We have examined the interaction of HAND2 and Phox2a at the DBH promoter. Using transient transfection of P19 or NT-2 cells, HAND2 is shown to synergistically enhance Phox2a-driven transcriptional activity at the DBH promoter, an effect that is enhanced by cAMP. While mutation of the Phox2a homeodomain binding sites HD1, HD2, and HD3 results in the loss of HAND2/Phox2a transactivation of DBH, it is the interaction of HAND2/Phox2a at the CRE/AP1-HD1/2 domains in the DBH enhancer that are required for synergistic activation by HAND2. We find that HAND2 functions as a transcriptional activator without directly binding to E-box sequences in the DBH promoter, suggesting that HAND2-mediated DBH activity occurs by protein–protein interactions with other transcriptional regulators. -

Defining the Transcriptomic Landscape of the Developing Enteric Nervous System and Its Cellular Environment Sweta Roy-Carson Iowa State University, [email protected]
Genetics, Development and Cell Biology Genetics, Development and Cell Biology Publications 2017 Defining the transcriptomic landscape of the developing enteric nervous system and its cellular environment Sweta Roy-Carson Iowa State University, [email protected] Kevin Natukunda Iowa State University, [email protected] Hsien-chao Chou Iowa State University Narinder Pal Iowa State University Caitlin Farris Iowa State University Follow this and additional works at: https://lib.dr.iastate.edu/gdcb_las_pubs See next page for additional authors Part of the Cell and Developmental Biology Commons, and the Molecular Genetics Commons The ompc lete bibliographic information for this item can be found at https://lib.dr.iastate.edu/ gdcb_las_pubs/202. For information on how to cite this item, please visit http://lib.dr.iastate.edu/ howtocite.html. This Article is brought to you for free and open access by the Genetics, Development and Cell Biology at Iowa State University Digital Repository. It has been accepted for inclusion in Genetics, Development and Cell Biology Publications by an authorized administrator of Iowa State University Digital Repository. For more information, please contact [email protected]. Defining the transcriptomic landscape of the developing enteric nervous system and its cellular environment Abstract Background: Motility and the coordination of moving food through the gastrointestinal tract rely on a complex network of neurons known as the enteric nervous system (ENS). Despite its critical function, many of the molecular mechanisms that direct the development of the ENS and the elaboration of neural network connections remain unknown. The og al of this study was to transcriptionally identify molecular pathways and candidate genes that drive specification, differentiation and the neural circuitry of specific neural progenitors, the phox2b expressing ENS cell lineage, during normal enteric nervous system development. -

Adult Epidermal Notch Activity Induces Dermal Accumulation of T Cells and Neural Crest Derivatives Through Upregulation of Jagged 1 Carrie A
DEVELOPMENT AND STEM CELLS RESEARCH ARTICLE 3569 Development 137, 3569-3579 (2010) doi:10.1242/dev.050310 © 2010. Published by The Company of Biologists Ltd Adult epidermal Notch activity induces dermal accumulation of T cells and neural crest derivatives through upregulation of jagged 1 Carrie A. Ambler1,2,* and Fiona M. Watt1,3,* SUMMARY Notch signalling regulates epidermal differentiation and tumour formation via non-cell autonomous mechanisms that are incompletely understood. This study shows that epidermal Notch activation via a 4-hydroxy-tamoxifen-inducible transgene caused epidermal thickening, focal detachment from the underlying dermis and hair clumping. In addition, there was dermal accumulation of T lymphocytes and stromal cells, some of which localised to the blisters at the epidermal-dermal boundary. The T cell infiltrate was responsible for hair clumping but not for other Notch phenotypes. Notch-induced stromal cells were heterogeneous, expressing markers of neural crest, melanocytes, smooth muscle and peripheral nerve. Although Slug1 expression was expanded in the epidermis, the stromal cells did not arise through epithelial-mesenchymal transition. Epidermal Notch activation resulted in upregulation of jagged 1 in both epidermis and dermis. When Notch was activated in the absence of epidermal jagged 1, jagged 1 was not upregulated in the dermis, and epidermal thickening, blister formation, accumulation of T cells and stromal cells were inhibited. Gene expression profiling revealed that epidermal Notch activation resulted in upregulation of several growth factors and cytokines, including TNF, the expression of which was dependent on epidermal jagged 1. We conclude that jagged 1 is a key mediator of non-cell autonomous Notch signalling in skin. -

REVIEW ARTICLE Pediatric Disorders with Autonomic Dysfunction: What
0031-3998/05/5801-0001 PEDIATRIC RESEARCH Vol. 58, No. 1, 2005 Copyright © 2005 International Pediatric Research Foundation, Inc. Printed in U.S.A. REVIEW ARTICLE Pediatric Disorders with Autonomic Dysfunction: What Role for PHOX2B? CLAUDE GAULTIER, HA TRANG, STÉPHANE DAUGER, AND JORGE GALLEGO INSERM U676 [C.G., S.D., J.G.], Services de Physiologie [C.G., H.T.] et de Réanimation Médicale Pédiatrique [S.D.], Hôpital Robert Debré, 75019 Paris, France ABSTRACT Hirschsprung disease, neuroblastomas, and congenital central crest disorders, international databases of clinical symptoms and hypoventilation syndrome can occur in combination, and familial molecular test results should be established. Furthermore, the cases have been reported in all three conditions. This suggests development of genetic mouse models should help to improve variable expression of a single genetic abnormality as the com- our understanding of the molecular mechanisms underlying neu- mon cause to these neural crest disorders. Because the PHOX2B ral crest disorders. (Pediatr Res 58: 1–6, 2005) gene is pivotal in the development of most relays of the auto- nomic nervous system, including all autonomic neural crest derivatives, it was considered a candidate gene for the above Abbreviations conditions. Recent studies have shown that 1) PHOX2B is the ANS, autonomic nervous system main disease-causing gene for congenital central hypoventilation CCHS, congenital central hypoventilation syndrome syndrome, an autosomal dominant disorder with incomplete HASH-1, human achaete-scute homologous 1 gene penetrance; 2) PHOX2B is the first gene for which germline HSCR, Hirschsprung disease mutations have been demonstrated to predispose to neuroblas- LO-CHS, late-onset central hypoventilation syndrome toma; and 3) Hirschsprung disease was associated with an in- MASH-1, mammalian achaete-scute homologous 1 gene tronic single-nucleotide polymorphism of the PHOX2B gene in a PHOX2B, paired-like homeobox 2b gene case-control study. -
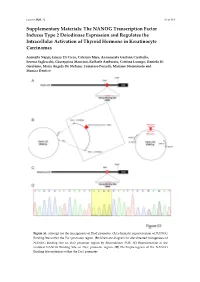
The NANOG Transcription Factor Induces Type 2 Deiodinase Expression and Regulates the Intracellular Activation of Thyroid Hormone in Keratinocyte Carcinomas
Cancers 2020, 12 S1 of S18 Supplementary Materials: The NANOG Transcription Factor Induces Type 2 Deiodinase Expression and Regulates the Intracellular Activation of Thyroid Hormone in Keratinocyte Carcinomas Annarita Nappi, Emery Di Cicco, Caterina Miro, Annunziata Gaetana Cicatiello, Serena Sagliocchi, Giuseppina Mancino, Raffaele Ambrosio, Cristina Luongo, Daniela Di Girolamo, Maria Angela De Stefano, Tommaso Porcelli, Mariano Stornaiuolo and Monica Dentice Figure S1. Strategy for the mutagenesis of Dio2 promoter. (A) Schematic representation of NANOG Binding Site within the Dio2 promoter region. (B) Schematic diagram for site‐directed mutagenesis of NANOG Binding Site on Dio2 promoter region by Recombinant PCR. (C) Representation of the mutated NANOG Binding Site on Dio2 promoter region. (D) Electropherogram of the NANOG Binding Site mutation within the Dio2 promoter. Cancers 2020, 12 S2 of S18 Figure S2. Strategy for the silencing of NANOG expression. (A) Cloning strategies for the generation of NANOG shRNA expression vectors. (B) Electropherograms of the NANOG shRNA sequences cloned into pcDNA3.1 vector. (C) Validation of effective NANOG down-modulation by two different NANOG shRNA vectors was assessed by Western Blot analysis of NANOG expression in BCC cells. (D) Quantification of NANOG protein levels versus Tubulin levels in the same experiment as in C is represented by histograms. Cancers 2020, 12 S3 of S18 Figure S3. The CD34+ cells are characterized by the expression of typical epithelial stemness genes. The mRNA levels of a panel of indicated stemness markers of epidermis were measured by Real Time PCR in the same experiment indicated in figure 3F and G. Cancers 2020, 12 S4 of S18 Figure S4. -

Prenatal Testing Requisition Form
BAYLOR MIRACA GENETICS LABORATORIES SHIP TO: Baylor Miraca Genetics Laboratories 2450 Holcombe, Grand Blvd. -Receiving Dock PHONE: 800-411-GENE | FAX: 713-798-2787 | www.bmgl.com Houston, TX 77021-2024 Phone: 713-798-6555 PRENATAL COMPREHENSIVE REQUISITION FORM PATIENT INFORMATION NAME (LAST,FIRST, MI): DATE OF BIRTH (MM/DD/YY): HOSPITAL#: ACCESSION#: REPORTING INFORMATION ADDITIONAL PROFESSIONAL REPORT RECIPIENTS PHYSICIAN: NAME: INSTITUTION: PHONE: FAX: PHONE: FAX: NAME: EMAIL (INTERNATIONAL CLIENT REQUIREMENT): PHONE: FAX: SAMPLE INFORMATION CLINICAL INDICATION FETAL SPECIMEN TYPE Pregnancy at risk for specific genetic disorder DATE OF COLLECTION: (Complete FAMILIAL MUTATION information below) Amniotic Fluid: cc AMA PERFORMING PHYSICIAN: CVS: mg TA TC Abnormal Maternal Screen: Fetal Blood: cc GESTATIONAL AGE (GA) Calculation for AF-AFP* NTD TRI 21 TRI 18 Other: SELECT ONLY ONE: Abnormal NIPT (attach report): POC/Fetal Tissue, Type: TRI 21 TRI 13 TRI 18 Other: Cultured Amniocytes U/S DATE (MM/DD/YY): Abnormal U/S (SPECIFY): Cultured CVS GA ON U/S DATE: WKS DAYS PARENTAL BLOODS - REQUIRED FOR CMA -OR- Maternal Blood Date of Collection: Multiple Pregnancy Losses LMP DATE (MM/DD/YY): Parental Concern Paternal Blood Date of Collection: Other Indication (DETAIL AND ATTACH REPORT): *Important: U/S dating will be used if no selection is made. Name: Note: Results will differ depending on method checked. Last Name First Name U/S dating increases overall screening performance. Date of Birth: KNOWN FAMILIAL MUTATION/DISORDER SPECIFIC PRENATAL TESTING Notice: Prior to ordering testing for any of the disorders listed, you must call the lab and discuss the clinical history and sample requirements with a genetic counselor. -

FOXA1 Overexpression Mediates Endocrine Resistance by Altering The
FOXA1 overexpression mediates endocrine resistance PNAS PLUS by altering the ER transcriptome and IL-8 expression in ER-positive breast cancer Xiaoyong Fua,b,c, Rinath Jeselsohnd, Resel Pereiraa,b,c, Emporia F. Hollingsworthe, Chad J. Creightonb,f, Fugen Lid, Martin Sheaa,b,f, Agostina Nardonea,b,f, Carmine De Angelisa,b,f, Laura M. Heiserg, Pavana Anurh, Nicholas Wangg, Catherine S. Grassog, Paul T. Spellmanh, Obi L. Griffithi, Anna Tsimelzona,b,f, Carolina Gutierreze, Shixia Huangb,c, Dean P. Edwardsb,c,e, Meghana V. Trivedia,b,j,k, Mothaffar F. Rimawia,b,f, Dolores Lopez-Terradae, Susan G. Hilsenbecka,b,f, Joe W. Grayg, Myles Brownd, C. Kent Osbornea,b,c,f, and Rachel Schiffa,b,c,f,1 aLester and Sue Smith Breast Center, Baylor College of Medicine, Houston, TX 77030; bDan L. Duncan Cancer Center, Baylor College of Medicine, Houston, TX 77030; cDepartment of Molecular and Cellular Biology, Baylor College of Medicine, Houston, TX 77030; dDana–Farber Cancer Institute, Harvard Medical School, Boston, MA 02215; eDepartment of Pathology, Baylor College of Medicine, Houston, TX 77030; fDepartment of Medicine, Baylor College of Medicine, Houston, TX 77030; gDepartment of Biomedical Engineering, Oregon Health and Science University, Portland, OR 97239; hDepartment of Molecular and Medical Genetics, Oregon Health and Science University, Portland, OR 97239; iMcDonnell Genome Institute, Washington University, St. Louis, MO 63108; jDepartment of Pharmacy Practice and Translational Research, University of Houston, Houston, TX 77204; and kDepartment of Pharmacological and Pharmaceutical Sciences, University of Houston, Houston, TX 77204 Edited by Bert W. O’Malley, Baylor College of Medicine, Houston, TX, and approved August 26, 2016 (received for review May 18, 2016) Forkhead box protein A1 (FOXA1) is a pioneer factor of estrogen renders these genomic regions more accessible to other tran- receptor α (ER)–chromatin binding and function, yet its aberration scription factors, such as ER (9), progesterone receptor (PR) in endocrine-resistant (Endo-R) breast cancer is unknown.