SET Domain Containing Protein 5 (SETD5) Enhances Tumor Cell
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Rare Variant Analysis of Human and Rodent Obesity Genes in Individuals with Severe Childhood Obesity Received: 11 November 2016 Audrey E
www.nature.com/scientificreports OPEN Rare Variant Analysis of Human and Rodent Obesity Genes in Individuals with Severe Childhood Obesity Received: 11 November 2016 Audrey E. Hendricks1,2, Elena G. Bochukova3,4, Gaëlle Marenne1, Julia M. Keogh3, Neli Accepted: 10 April 2017 Atanassova3, Rebecca Bounds3, Eleanor Wheeler1, Vanisha Mistry3, Elana Henning3, Published: xx xx xxxx Understanding Society Scientific Group*, Antje Körner5,6, Dawn Muddyman1, Shane McCarthy1, Anke Hinney7, Johannes Hebebrand7, Robert A. Scott8, Claudia Langenberg8, Nick J. Wareham8, Praveen Surendran9, Joanna M. Howson9, Adam S. Butterworth9,10, John Danesh1,9,10, EPIC-CVD Consortium*, Børge G Nordestgaard11,12, Sune F Nielsen11,12, Shoaib Afzal11,12, SofiaPa padia3, SofieAshford 3, Sumedha Garg3, Glenn L. Millhauser13, Rafael I. Palomino13, Alexandra Kwasniewska3, Ioanna Tachmazidou1, Stephen O’Rahilly3, Eleftheria Zeggini1, UK10K Consortium*, Inês Barroso1,3 & I. Sadaf Farooqi3 Obesity is a genetically heterogeneous disorder. Using targeted and whole-exome sequencing, we studied 32 human and 87 rodent obesity genes in 2,548 severely obese children and 1,117 controls. We identified 52 variants contributing to obesity in 2% of cases including multiple novel variants in GNAS, which were sometimes found with accelerated growth rather than short stature as described previously. Nominally significant associations were found for rare functional variants inBBS1 , BBS9, GNAS, MKKS, CLOCK and ANGPTL6. The p.S284X variant in ANGPTL6 drives the association signal (rs201622589, MAF~0.1%, odds ratio = 10.13, p-value = 0.042) and results in complete loss of secretion in cells. Further analysis including additional case-control studies and population controls (N = 260,642) did not support association of this variant with obesity (odds ratio = 2.34, p-value = 2.59 × 10−3), highlighting the challenges of testing rare variant associations and the need for very large sample sizes. -

Elucidating Biological Roles of Novel Murine Genes in Hearing Impairment in Africa
Preprints (www.preprints.org) | NOT PEER-REVIEWED | Posted: 19 September 2019 doi:10.20944/preprints201909.0222.v1 Review Elucidating Biological Roles of Novel Murine Genes in Hearing Impairment in Africa Oluwafemi Gabriel Oluwole,1* Abdoulaye Yal 1,2, Edmond Wonkam1, Noluthando Manyisa1, Jack Morrice1, Gaston K. Mazanda1 and Ambroise Wonkam1* 1Division of Human Genetics, Department of Pathology, Faculty of Health Sciences, University of Cape Town, Observatory, Cape Town, South Africa. 2Department of Neurology, Point G Teaching Hospital, University of Sciences, Techniques and Technology, Bamako, Mali. *Correspondence to: [email protected]; [email protected] Abstract: The prevalence of congenital hearing impairment (HI) is highest in Africa. Estimates evaluated genetic causes to account for 31% of HI cases in Africa, but the identification of associated causative genes mutations have been challenging. In this study, we reviewed the potential roles, in humans, of 38 novel genes identified in a murine study. We gathered information from various genomic annotation databases and performed functional enrichment analysis using online resources i.e. genemania and g.proflier. Results revealed that 27/38 genes are express mostly in the brain, suggesting additional cognitive roles. Indeed, HERC1- R3250X had been associated with intellectual disability in a Moroccan family. A homozygous 216-bp deletion in KLC2 was found in two siblings of Egyptian descent with spastic paraplegia. Up to 27/38 murine genes have link to at least a disease, and the commonest mode of inheritance is autosomal recessive (n=8). Network analysis indicates that 20 other genes have intermediate and biological links to the novel genes, suggesting their possible roles in HI. -

Stanford University
Stanford University Stanford, California Reports on Federal Awards in Accordance with the Uniform Guidance August 31, 2019 EIN: 94-1156365 Stanford University Stanford, California Reports on Federal Awards in Accordance with the Uniform Guidance August 31, 2019 Page(s) I. Financial Report of Independent Auditors .................................................................................................. 1–2 Consolidated Financial Statements............................................................................................. 3–6 Notes to Consolidated Financial Statements ............................................................................. 7–77 Schedule of Expenditures of Federal Awards Part A, Award Expenditure Detail ................................................................................. 78–132 Part B, Summary of Program Clusters ........................................................................ 133–191 Part C, Federal Loan Program Year End Balances ............................................................. 192 Notes to Schedule of Expenditures of Federal Awards ............................................................... 193 II. Internal Control and Compliance Report of Independent Auditors on Internal Control Over Financial Reporting and on Compliance and Other Matters Based on an Audit of Financial Statements Performed in Accordance with Government Auditing Standards ...................................... 194-195 Report of Independent Auditors on Compliance with Requirements That Could Have -

A Sleeping Beauty Transposon-Mediated Screen Identifies Murine Susceptibility Genes for Adenomatous Polyposis Coli (Apc)-Dependent Intestinal Tumorigenesis
A Sleeping Beauty transposon-mediated screen identifies murine susceptibility genes for adenomatous polyposis coli (Apc)-dependent intestinal tumorigenesis Timothy K. Starra,1, Patricia M. Scottb, Benjamin M. Marshb, Lei Zhaob, Bich L. N. Thanb, M. Gerard O’Sullivana,c, Aaron L. Sarverd, Adam J. Dupuye, David A. Largaespadaa, and Robert T. Cormierb,1 aDepartment of Genetics, Cell Biology and Development, Center for Genome Engineering, Masonic Cancer Center, University of Minnesota, Minneapolis, MN 55455; bDepartment of Biochemistry and Molecular Biology, University of Minnesota Medical School, Duluth, MN 55812; cDepartment of Veterinary Population Medicine, College of Veterinary Medicine, University of Minnesota, St. Paul, MN 55108; dDepartment of Biostatistics and Informatics, Masonic Cancer Center, University of Minnesota, Minneapolis, MN 55455; and eDepartment of Anatomy and Cell Biology, University of Iowa, Iowa City, IA 52242 Edited* by William F. Dove, University of Wisconsin, Madison, WI, and approved March 2, 2011 (received for review December 1, 2010) Min Min It is proposed that a progressive series of mutations and epigenetic conducted the screen in mice carrying the Apc allele. Apc events leads to human colorectal cancer (CRC) and metastasis. mice harbor a T→A nonsense mutation in the Apc gene (4, 5) Furthermore, data from resequencing of the coding regions of that results in a truncated protein product that is unable to bind human CRC suggests that a relatively large number of mutations β-catenin and promote its degradation, thus leading to abnormal occur in individual human CRC, most at low frequency. The levels of β-catenin protein and up-regulation of β-catenin target functional role of these low-frequency mutations in CRC, and genes such as cyclin D1 (Ccnd1) and myelocytomatosis oncogene specifically how they may cooperate with high-frequency muta- (C-Myc). -
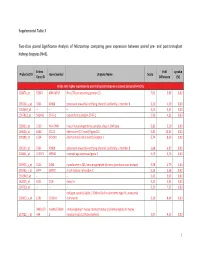
Supplemental Table 3 Two-Class Paired Significance Analysis of Microarrays Comparing Gene Expression Between Paired
Supplemental Table 3 Two‐class paired Significance Analysis of Microarrays comparing gene expression between paired pre‐ and post‐transplant kidneys biopsies (N=8). Entrez Fold q‐value Probe Set ID Gene Symbol Unigene Name Score Gene ID Difference (%) Probe sets higher expressed in post‐transplant biopsies in paired analysis (N=1871) 218870_at 55843 ARHGAP15 Rho GTPase activating protein 15 7,01 3,99 0,00 205304_s_at 3764 KCNJ8 potassium inwardly‐rectifying channel, subfamily J, member 8 6,30 4,50 0,00 1563649_at ‐‐ ‐‐ ‐‐ 6,24 3,51 0,00 1567913_at 541466 CT45‐1 cancer/testis antigen CT45‐1 5,90 4,21 0,00 203932_at 3109 HLA‐DMB major histocompatibility complex, class II, DM beta 5,83 3,20 0,00 204606_at 6366 CCL21 chemokine (C‐C motif) ligand 21 5,82 10,42 0,00 205898_at 1524 CX3CR1 chemokine (C‐X3‐C motif) receptor 1 5,74 8,50 0,00 205303_at 3764 KCNJ8 potassium inwardly‐rectifying channel, subfamily J, member 8 5,68 6,87 0,00 226841_at 219972 MPEG1 macrophage expressed gene 1 5,59 3,76 0,00 203923_s_at 1536 CYBB cytochrome b‐245, beta polypeptide (chronic granulomatous disease) 5,58 4,70 0,00 210135_s_at 6474 SHOX2 short stature homeobox 2 5,53 5,58 0,00 1562642_at ‐‐ ‐‐ ‐‐ 5,42 5,03 0,00 242605_at 1634 DCN decorin 5,23 3,92 0,00 228750_at ‐‐ ‐‐ ‐‐ 5,21 7,22 0,00 collagen, type III, alpha 1 (Ehlers‐Danlos syndrome type IV, autosomal 201852_x_at 1281 COL3A1 dominant) 5,10 8,46 0,00 3493///3 IGHA1///IGHA immunoglobulin heavy constant alpha 1///immunoglobulin heavy 217022_s_at 494 2 constant alpha 2 (A2m marker) 5,07 9,53 0,00 1 202311_s_at -

Genes Essential for Embryonic Stem Cells Are Associated with Neurodevelopmental Disorders
Downloaded from genome.cshlp.org on September 25, 2021 - Published by Cold Spring Harbor Laboratory Press Resource Genes essential for embryonic stem cells are associated with neurodevelopmental disorders Shahar Shohat and Sagiv Shifman Department of Genetics, The Institute of Life Sciences, The Hebrew University of Jerusalem, Jerusalem 9190401, Israel Mouse embryonic stem cells (mESCs) are key components in generating mouse models for human diseases and performing basic research on pluripotency, yet the number of genes essential for mESCs is still unknown. We performed a genome-wide screen for essential genes in mESCs and compared it to screens in human cells. We found that essential genes are enriched for basic cellular functions, are highly expressed in mESCs, and tend to lack paralog genes. We discovered that genes that are essential specifically in mESCs play a role in pathways associated with their pluripotent state. We show that 29.5% of human genes intolerant to loss-of-function mutations are essential in mouse or human ESCs, and that the human phenotypes most significantly associated with genes essential for ESCs are neurodevelopmental. Our results provide insights into essential genes in the mouse, the pathways which govern pluripotency, and suggest that many genes associated with neurodevelop- mental disorders are essential at very early embryonic stages. [Supplemental material is available for this article.] Essential genes are required for organism survival or development. Results Recent advances in CRISPR technology have enabled the identifi- cation of essential genes in multiple human cancer cell lines CRISPR screen in mouse embryonic stem cells (Wang et al. 2015, 2017; Tzelepis et al. -

Setd5 Haploinsufficiency Alters Neuronal Network Connectivity And
Moore et al. Translational Psychiatry (2019) 9:24 https://doi.org/10.1038/s41398-018-0344-y Translational Psychiatry ARTICLE Open Access Setd5 haploinsufficiency alters neuronal network connectivity and leads to autistic- like behaviors in mice Spencer M. Moore 1,2,JasonS.Seidman1,3, Jacob Ellegood 4,5,RichardGao6, Alex Savchenko2,TyD.Troutman3, Yohei Abe3, Josh Stender3, Daehoon Lee7,SicongWang2, Bradley Voytek6,8,9,10, Jason P. Lerch5,11, Hoonkyo Suh 7, Christopher K Glass3 and Alysson R. Muotri 2,3,10,12,13,14 Abstract SETD5, a gene linked to intellectual disability (ID) and autism spectrum disorder (ASD), is a member of the SET-domain family and encodes a putative histone methyltransferase (HMT). To date, the mechanism by which SETD5 haploinsufficiency causes ASD/ID remains an unanswered question. Setd5 is the highly conserved mouse homolog, and although the Setd5 null mouse is embryonic lethal, the heterozygote is viable. Morphological tracing and multielectrode array was used on cultured cortical neurons. MRI was conducted of adult mouse brains and immunohistochemistry of juvenile mouse brains. RNA-Seq was used to investigate gene expression in the developing cortex. Behavioral assays were conducted on adult mice. Setd5+/− cortical neurons displayed significantly reduced synaptic density and neuritic outgrowth in vitro, with corresponding decreases in network activity and synchrony by electrophysiology. A specific subpopulation of fetal Setd5+/− cortical neurons showed altered gene expression of neurodevelopment-related genes. Setd5+/− animals manifested several autism-like behaviors, including hyperactivity, +/− 1234567890():,; 1234567890():,; 1234567890():,; 1234567890():,; cognitive deficit, and altered social interactions. Anatomical differences were observed in Setd5 adult brains, accompanied by a deficit of deep-layer cortical neurons in the developing brain. -

SUPPLEMENTARY DATA Page Supplementary Text …………………………………………………………………………………
SUPPLEMENTARY DATA Page Supplementary Text …………………………………………………………………………………. 2 Discovery Cohort Descriptions ……………………………………………………………………. 2 Translational Cohort Descriptions …………………………………………………………………. 6 Clinical characteristics of the Translation cohorts ………………………………………………… 7 References …………………………………………………………………………………………. 8 Supplementary Table 1. Pearson’s correlation matrix for T2D-related quantitative traits in the Discovery cohorts. A) Euglycemic clamp cohorts including MACAD, HTN-IR, NIDDM-Athero and B) FSIGT cohorts including IRAS, IRAS Family, BetaGene, TRIPOD. …………………………...... 10 Supplementary Table 2. Top Discovery hits (P<1.00x10‐4) from the GUARDIAN Consortium. SNPs with evidence of association are tabulated by trait (a. MCRI, b. SI+M, c. SI, d. M, e. AIRg, f. DI, g. SG) in order of genomic position. ...…………………………………………………………...... 11 Supplementary Table 3. Top Discovery hits from the GUARDIAN Consortium ordered by trait. … 41 Supplementary Table 4. Top Discovery hits from the GUARDIAN Consortium ordered by trait with conditional analyses performed at loci with multiple signals of association or in close proximity to previously identified T2D loci. ……………………………………………………………………... 42 Supplementary Table 5. Association with reported type 2 diabetes (T2D) susceptibility loci (a. MCRI, b. SI+M, c. SI, d. M, e. AIRg, f. DI, and g. SG) in GUARDIAN Hispanic Americans (nmax=3,787). Index SNPs reaching genome‐wide significance (P<5.0E‐08) for association with T2D were selected from the NHGRI GWAS catalog. ……………………………………………………… 43 Supplementary Table 6. All SNPs from GWAS of T2D-related quantitative traits with Translation to T2D in Mexican-origin cohorts ordered by trait from the Discovery stage. ……………………….. 65 Supplementary Table 7. Top regions from GWAS of T2D-related quantitative traits with translation to T2D in Mexican-origin. ………………………………………………………………… 86 Supplementary Table 8. -

Setd5 Is Essential for Mammalian Development and the Co-Transcriptional Regulation of Histone Acetylation Anna B
© 2016. Published by The Company of Biologists Ltd | Development (2016) 143, 4595-4607 doi:10.1242/dev.141465 STEM CELLS AND REGENERATION RESEARCH ARTICLE Setd5 is essential for mammalian development and the co-transcriptional regulation of histone acetylation Anna B. Osipovich1,2, Rama Gangula1, Pedro G. Vianna1 and Mark A. Magnuson1,2,* ABSTRACT H3 and H4 has long been associated with transcriptionally active SET domain-containing proteins play a vital role in regulating genes and open chromatin, more recent studies have established that gene expression during development through modifications in histone methylations are also associated with actively transcribed chromatin structure. Here we show that SET domain-containing genes, including histone H3 methylation at lysines 4, 36 and 79 5(Setd5) is divergently transcribed with Gt(ROSA26)Sor,is (H3K4me, H3K36me and H3K79me) and histone H4 methylation necessary for mammalian development, and interacts with the at lysine 20 (H4K20me) (Smolle and Workman, 2013). Studies in PAF1 co-transcriptional complex and other proteins. Setd5-deficient yeast have shown that these histone modifications occur co- mouse embryos exhibit severe defects in neural tube formation, transcriptionally and require coordination between RNAP II, somitogenesis and cardiac development, have aberrant elongation complexes and regulators of histone modification vasculogenesis in embryos, yolk sacs and placentas, and die (Tanny, 2014). between embryonic day 10.5 and 11.5. Setd5-deficient embryonic Polymerase-associated factor 1 complex (PAF1C), which stem cells have impaired cellular proliferation, increased apoptosis, contains Paf1, Ctr9, Leo1, Cdc73 and Rtf1 as core components, defective cell cycle progression, a diminished ability to differentiate colocalizes to coding regions of genes and is involved in multiple into cardiomyocytes and greatly perturbed gene expression. -
HDAC3 Complex Determines Enhancer Activation for Adipogenesis
Spatiotemporal dynamics of SETD5-containing NCoR-HDAC3 complex determines enhancer activation for adipogenesis Yoshihiro Matsumura Division of Metabolic Medicine, RCAST, The University of Tokyo Ryo Ito Tohoku University Graduate School of Medicine Ayumu Yajima The University of Tokyo Rei Yamaguchi Tohoku University Graduate School of Medicine Kenta Magoori The University of Tokyo Toshiya Tanaka The University of Tokyo https://orcid.org/0000-0003-1224-7898 Takeshi Kawamura The University of Tokyo Hiroyuki Hirakawa The University of Tokyo Hitomi Fujihashi The University of Tokyo Yohei Abe The University of Tokyo Ryo Nakaki The University of Tokyo Aoi Uchida Division of Metabolic Medicine, RCAST, The University of Tokyo Shogo Yamamoto Research Center for Advanced Science and Technology, The University of Tokyo https://orcid.org/0000-0002-5564-2567 Satoshi Ota The University of Tokyo Shuichi Tsutsumi University of Tokyo Shin-ichi Inoue Tohoku University Hiroshi Kimura Tokyo Institute of Technology https://orcid.org/0000-0003-0854-083X Yoichiro Wada University of Tokyo Takeshi Inagaki The University of Tokyo Tatsuhiko Kodama University of Tokyo Timothy Osborne Institute for Fundamental Biomedical Research Hiroyuki Aburatani University of Tokyo https://orcid.org/0000-0003-0438-1544 Koichi Node Saga University https://orcid.org/0000-0002-2534-0939 Juro Sakai ( [email protected] ) Division of Metabolic Medicine, RCAST, The University of Tokyo Article Keywords: adipogenesis, cell-type specic gene expression Posted Date: February 25th, 2021 -
Thesis, I Have Studied Rare Monogenic Novel Neurodevelopmental Disorders Associated with ID
APPLYING ANIMAL MODELLING TO UNDERSTAND RARE NOVEL NEURODEVELOPMENTAL DISORDERS ASSOCIATED WITH INTELLECTUAL DISABILITY Maria Levitin St. John’s College University of Cambridge Wellcome Sanger Institute This dissertation is submitted for the degree of Doctor of Philosophy. September 2017 Applying animal modelling to understand rare novel neurodevelopmental disorders associated with intellectual disability Maria Levitin Intellectual disability (ID) is categorised by a significant reduction in cognitive function and adaptive abilities that begin in childhood. ID is part of a heterogeneous group of neurodevelopmental conditions associated with impairment in developmental domains and a cause of particularly adverse socioeconomic impact worldwide. There have been many recent advances in identifying causative genetic mutations in previously unexplained ID cases. With these advances comes an increasing demand for understanding mechanisms underpinning these pathogenic pathways. In this PhD thesis, I have studied rare monogenic novel neurodevelopmental disorders associated with ID. The objective of the thesis was to model a subset of mutations associated with novel neurodevelopmental disorders in mice to demonstrate a causal link between mutation and phenotype and to further understand the mechanisms by which these mutations result in human neurodevelopmental disorders. In order to achieve this, I adopted a multi-phase approach. Firstly, I designed a phenotyping platform, by combining behavioural and cognitive tests with morphometric brain analysis and genome-wide transcriptional analysis. I then used this approach to study KPTN-related syndrome, a novel developmental disorder that to date has not been characterised in mice, successfully recapitulating the main phenotypes described in the patients. Moreover, I gained further insight into the underlying pathogenic mechanisms associated with the disorder, opening the possibility of a therapy that could treat some aspects of cognitive and morphological impairments identified in the patients with KPTN-related syndrome. -

Loss-Of-Function Variants of SETD5 Cause Intellectual Disability and the Core Phenotype of Microdeletion 3P25.3 Syndrome
European Journal of Human Genetics (2015) 23, 753–760 & 2015 Macmillan Publishers Limited All rights reserved 1018-4813/15 www.nature.com/ejhg ARTICLE Loss-of-function variants of SETD5 cause intellectual disability and the core phenotype of microdeletion 3p25.3 syndrome Alma Kuechler1,11, Alexander M Zink2,11, Thomas Wieland3, Hermann-Josef Lu¨decke1, Kirsten Cremer2, Leonardo Salviati4, Pamela Magini5, Kimia Najafi6, Christiane Zweier7, Johanna Christina Czeschik1, Stefan Aretz2, Sabine Endele7, Federica Tamburrino8, Claudia Pinato4, Maurizio Clementi4, Jasmin Gundlach3, Carina Maylahn2, Laura Mazzanti8, Eva Wohlleber2, Thomas Schwarzmayr3, Roxana Kariminejad6, Avner Schlessinger9, Dagmar Wieczorek1, Tim M Strom3,12, Gaia Novarino10,12 and Hartmut Engels*,2,12 Intellectual disability (ID) has an estimated prevalence of 2–3%. Due to its extreme heterogeneity, the genetic basis of ID remains elusive in many cases. Recently, whole exome sequencing (WES) studies revealed that a large proportion of sporadic cases are caused by de novo gene variants. To identify further genes involved in ID, we performed WES in 250 patients with unexplained ID and their unaffected parents and included exomes of 51 previously sequenced child–parents trios in the analysis. Exome analysis revealed de novo intragenic variants in SET domain-containing 5 (SETD5) in two patients. One patient carried a nonsense variant, and the other an 81 bp deletion located across a splice-donor site. Chromosomal microarray diagnostics further identified four de novo non-recurrent microdeletions encompassing SETD5. CRISPR/Cas9 mutation modelling of the two intragenic variants demonstrated nonsense-mediated decay of the resulting transcripts, pointing to a loss-of-function (LoF) and haploinsufficiency as the common disease-causing mechanism of intragenic SETD5 sequence variants and SETD5-containing microdeletions.