OLFML3) Is a Candidate for Severe
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Role of Amylase in Ovarian Cancer Mai Mohamed University of South Florida, [email protected]
University of South Florida Scholar Commons Graduate Theses and Dissertations Graduate School July 2017 Role of Amylase in Ovarian Cancer Mai Mohamed University of South Florida, [email protected] Follow this and additional works at: http://scholarcommons.usf.edu/etd Part of the Pathology Commons Scholar Commons Citation Mohamed, Mai, "Role of Amylase in Ovarian Cancer" (2017). Graduate Theses and Dissertations. http://scholarcommons.usf.edu/etd/6907 This Dissertation is brought to you for free and open access by the Graduate School at Scholar Commons. It has been accepted for inclusion in Graduate Theses and Dissertations by an authorized administrator of Scholar Commons. For more information, please contact [email protected]. Role of Amylase in Ovarian Cancer by Mai Mohamed A dissertation submitted in partial fulfillment of the requirements for the degree of Doctor of Philosophy Department of Pathology and Cell Biology Morsani College of Medicine University of South Florida Major Professor: Patricia Kruk, Ph.D. Paula C. Bickford, Ph.D. Meera Nanjundan, Ph.D. Marzenna Wiranowska, Ph.D. Lauri Wright, Ph.D. Date of Approval: June 29, 2017 Keywords: ovarian cancer, amylase, computational analyses, glycocalyx, cellular invasion Copyright © 2017, Mai Mohamed Dedication This dissertation is dedicated to my parents, Ahmed and Fatma, who have always stressed the importance of education, and, throughout my education, have been my strongest source of encouragement and support. They always believed in me and I am eternally grateful to them. I would also like to thank my brothers, Mohamed and Hussien, and my sister, Mariam. I would also like to thank my husband, Ahmed. -

Appendix 2. Significantly Differentially Regulated Genes in Term Compared with Second Trimester Amniotic Fluid Supernatant
Appendix 2. Significantly Differentially Regulated Genes in Term Compared With Second Trimester Amniotic Fluid Supernatant Fold Change in term vs second trimester Amniotic Affymetrix Duplicate Fluid Probe ID probes Symbol Entrez Gene Name 1019.9 217059_at D MUC7 mucin 7, secreted 424.5 211735_x_at D SFTPC surfactant protein C 416.2 206835_at STATH statherin 363.4 214387_x_at D SFTPC surfactant protein C 295.5 205982_x_at D SFTPC surfactant protein C 288.7 1553454_at RPTN repetin solute carrier family 34 (sodium 251.3 204124_at SLC34A2 phosphate), member 2 238.9 206786_at HTN3 histatin 3 161.5 220191_at GKN1 gastrokine 1 152.7 223678_s_at D SFTPA2 surfactant protein A2 130.9 207430_s_at D MSMB microseminoprotein, beta- 99.0 214199_at SFTPD surfactant protein D major histocompatibility complex, class II, 96.5 210982_s_at D HLA-DRA DR alpha 96.5 221133_s_at D CLDN18 claudin 18 94.4 238222_at GKN2 gastrokine 2 93.7 1557961_s_at D LOC100127983 uncharacterized LOC100127983 93.1 229584_at LRRK2 leucine-rich repeat kinase 2 HOXD cluster antisense RNA 1 (non- 88.6 242042_s_at D HOXD-AS1 protein coding) 86.0 205569_at LAMP3 lysosomal-associated membrane protein 3 85.4 232698_at BPIFB2 BPI fold containing family B, member 2 84.4 205979_at SCGB2A1 secretoglobin, family 2A, member 1 84.3 230469_at RTKN2 rhotekin 2 82.2 204130_at HSD11B2 hydroxysteroid (11-beta) dehydrogenase 2 81.9 222242_s_at KLK5 kallikrein-related peptidase 5 77.0 237281_at AKAP14 A kinase (PRKA) anchor protein 14 76.7 1553602_at MUCL1 mucin-like 1 76.3 216359_at D MUC7 mucin 7, -

Alzheimer's Disease
VU Research Portal Spot the difference Bossers, C.A.M. 2009 document version Publisher's PDF, also known as Version of record Link to publication in VU Research Portal citation for published version (APA) Bossers, C. A. M. (2009). Spot the difference: microarray analysis of gene expression changes in Alzheimer's and Parkinson's Disease. General rights Copyright and moral rights for the publications made accessible in the public portal are retained by the authors and/or other copyright owners and it is a condition of accessing publications that users recognise and abide by the legal requirements associated with these rights. • Users may download and print one copy of any publication from the public portal for the purpose of private study or research. • You may not further distribute the material or use it for any profit-making activity or commercial gain • You may freely distribute the URL identifying the publication in the public portal ? Take down policy If you believe that this document breaches copyright please contact us providing details, and we will remove access to the work immediately and investigate your claim. E-mail address: [email protected] Download date: 11. Oct. 2021 Spot the difference: microarray analysis of gene expression changes in Alzheimer’s and Parkinson’s Disease The research described in this thesis was conducted at the Netherlands Institute for Neuroscience, an Institute of the Royal Netherlands Academy of Arts and Sci- ences, Amsterdam, The Netherlands. The research was financially supported by a collaboration between the Netherlands Institute for Neuroscience and Solvay Phar- maceuticals. Leescommissie: dr. -

Human Induced Pluripotent Stem Cell–Derived Podocytes Mature Into Vascularized Glomeruli Upon Experimental Transplantation
BASIC RESEARCH www.jasn.org Human Induced Pluripotent Stem Cell–Derived Podocytes Mature into Vascularized Glomeruli upon Experimental Transplantation † Sazia Sharmin,* Atsuhiro Taguchi,* Yusuke Kaku,* Yasuhiro Yoshimura,* Tomoko Ohmori,* ‡ † ‡ Tetsushi Sakuma, Masashi Mukoyama, Takashi Yamamoto, Hidetake Kurihara,§ and | Ryuichi Nishinakamura* *Department of Kidney Development, Institute of Molecular Embryology and Genetics, and †Department of Nephrology, Faculty of Life Sciences, Kumamoto University, Kumamoto, Japan; ‡Department of Mathematical and Life Sciences, Graduate School of Science, Hiroshima University, Hiroshima, Japan; §Division of Anatomy, Juntendo University School of Medicine, Tokyo, Japan; and |Japan Science and Technology Agency, CREST, Kumamoto, Japan ABSTRACT Glomerular podocytes express proteins, such as nephrin, that constitute the slit diaphragm, thereby contributing to the filtration process in the kidney. Glomerular development has been analyzed mainly in mice, whereas analysis of human kidney development has been minimal because of limited access to embryonic kidneys. We previously reported the induction of three-dimensional primordial glomeruli from human induced pluripotent stem (iPS) cells. Here, using transcription activator–like effector nuclease-mediated homologous recombination, we generated human iPS cell lines that express green fluorescent protein (GFP) in the NPHS1 locus, which encodes nephrin, and we show that GFP expression facilitated accurate visualization of nephrin-positive podocyte formation in -

Persistent Transcription-Blocking DNA Lesions Trigger Somatic Growth Attenuation Associated with Longevity
ARTICLES Persistent transcription-blocking DNA lesions trigger somatic growth attenuation associated with longevity George A. Garinis1,2, Lieneke M. Uittenboogaard1, Heike Stachelscheid3,4, Maria Fousteri5, Wilfred van Ijcken6, Timo M. Breit7, Harry van Steeg8, Leon H. F. Mullenders5, Gijsbertus T. J. van der Horst1, Jens C. Brüning4,9, Carien M. Niessen3,9,10, Jan H. J. Hoeijmakers1 and Björn Schumacher1,9,11 The accumulation of stochastic DNA damage throughout an organism’s lifespan is thought to contribute to ageing. Conversely, ageing seems to be phenotypically reproducible and regulated through genetic pathways such as the insulin-like growth factor-1 (IGF-1) and growth hormone (GH) receptors, which are central mediators of the somatic growth axis. Here we report that persistent DNA damage in primary cells from mice elicits changes in global gene expression similar to those occurring in various organs of naturally aged animals. We show that, as in ageing animals, the expression of IGF-1 receptor and GH receptor is attenuated, resulting in cellular resistance to IGF-1. This cell-autonomous attenuation is specifically induced by persistent lesions leading to stalling of RNA polymerase II in proliferating, quiescent and terminally differentiated cells; it is exacerbated and prolonged in cells from progeroid mice and confers resistance to oxidative stress. Our findings suggest that the accumulation of DNA damage in transcribed genes in most if not all tissues contributes to the ageing-associated shift from growth to somatic maintenance that triggers stress resistance and is thought to promote longevity. Ageing represents the progressive functional decline that is exempted levels as a result of pituitary dysfunction (Snell and Ames mice) — have an from evolutionary selection because it largely occurs after reproduc- extended lifespan17–20. -

Arginine to Glutamine Mutation in Olfactomedin Like 3 (OLFML3) Is a Candidate for Severe
bioRxiv preprint doi: https://doi.org/10.1101/321406; this version posted July 25, 2018. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. 1 Arginine to glutamine mutation in olfactomedin like 3 (OLFML3) is a candidate for severe 2 goniodysgenesis and glaucoma in the Border Collie dog breed. 3 4 Carys A Pugh*, Lindsay L Farrell* 1, Ailsa J Carlisle*, Stephen J Bush*†, Violeta Trejo- 5 Reveles*, Oswald Matika*, Arne de Kloet‡, Caitlin Walsh‡, Stephen C Bishop* 2, James GD 6 Prendergast*, Jeffrey J Schoenebeck*, Joe Rainger*, Kim M Summers*§ 7 8 * The Roslin Institute, Easter Bush EH25 9RG, United Kingdom 9 † Nuffield Department of Clinical Medicine, University of Oxford, John Radcliffe Hospital, 10 Oxford OX3 9DU, United Kingdom 11 ‡ Animal Genetics, 1336 Timberlane Rd, Tallahassee, FL 32312, USA 12 § Mater Research Institute-UQ, Translational Research Institute, Brisbane, Qld 4012, 13 Australia 14 1 Present address: British Columbia Centre on Substance Abuse, St Paul’s Hospital, 15 Vancouver, BC V6Z 1Y6, Canada 16 2 Deceased 17 18 1 bioRxiv preprint doi: https://doi.org/10.1101/321406; this version posted July 25, 2018. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. 19 Running title: OLFML3 mutation in Border Collie glaucoma 20 21 Key words: Glaucoma, goniodysgenesis, olfactomedin like 3, Border Collie, eye development 22 23 Corresponding authors: 24 Dr Carys Pugh, The Roslin Institute, University of Edinburgh, Easter Bush EH25 9RG, UK 25 Phone: +441316519221 26 Email: [email protected] 27 Professor Kim M Summers, Mater Research Institute-UQ, Translational Research Institute, 28 37 Kent St, Woolloongabba, QLD 4102, Australia 29 Phone: +61734437655 30 Email: [email protected] 2 bioRxiv preprint doi: https://doi.org/10.1101/321406; this version posted July 25, 2018. -

Targeting Breakthroughs - Harnessing Cures October 9-12, 2014 • Grand Hyatt Shanghai • Pudong, Shanghai, P.R
New Horizons in Cancer Research: Targeting Breakthroughs - Harnessing Cures October 9-12, 2014 • Grand Hyatt Shanghai • Pudong, Shanghai, P.R. China Poster Session A – Friday, October 10, 2014, 12:30-3:00 p.m. A02 Therapeutic targeting of the histone demethylase KDM4 subfamily in breast cancer. Qin Ye, Andreana Holowatyj, Jack Wu, Hui Liu, Zeng-Quan Yang. Department of Oncology, Karmanos Cancer Institute, Wayne State University, Detroit, MI, USA. Breast cancer is a heterogeneous disease, consisting of tumors with varying pathologic and molecular characteristics. The primary biological subtypes of breast cancer include estrogen receptor–positive (ER+) tumors (luminal A and B), tumors that are human epidermal growth factor receptor 2 (HER2) –enriched, and tumors that are ER/PR-negative (basal-like). These molecular determinants have significant effects on disease pathophysiology, clinical outcome, and treatment response. ER+ tumors are generally associated with better clinical prognosis, whereas basal-like tumors are associated with higher rates of metastasis and death. The treatment for basal breast cancer consists of standard chemotherapy regimens, as no effective molecularly targeted therapy has been developed. New therapeutic options are likely to result from a growing understanding of both genetic and epigenetic abnormalities that participate in the different types of breast cancer, which helps to identify new subtype-specific targets for therapy. Aberrant histone lysine methylation, controlled by histone lysine methyltransferases and demethylases, plays significant roles in cancer initiation and progression. Previously, we demonstrated that histone demethylase KDM4C (also known as GASC1 and JMJD2C) is significantly amplified and overexpressed in aggressive basal-type breast cancer. The KDM4C/GASC1 protein belongs to the KDM4 family of histone demethylases, in which KDM4A, B, and C share more than 50% percent of sequence identity. -
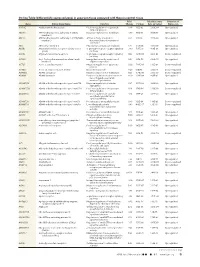
Differentially Expressed Genes in Aneurysm Tissue Compared With
On-line Table: Differentially expressed genes in aneurysm tissue compared with those in control tissue Fold False Discovery Direction of Gene Entrez Gene Name Function Change P Value Rate (q Value) Expression AADAC Arylacetamide deacetylase Positive regulation of triglyceride 4.46 1.33E-05 2.60E-04 Up-regulated catabolic process ABCA6 ATP-binding cassette, subfamily A (ABC1), Integral component of membrane 3.79 9.15E-14 8.88E-12 Up-regulated member 6 ABCC3 ATP-binding cassette, subfamily C (CFTR/MRP), ATPase activity, coupled to 6.63 1.21E-10 7.33E-09 Up-regulated member 3 transmembrane movement of substances ABI3 ABI family, member 3 Peptidyl-tyrosine phosphorylation 6.47 2.47E-05 4.56E-04 Up-regulated ACKR1 Atypical chemokine receptor 1 (Duffy blood G-protein–coupled receptor signaling 3.80 7.95E-10 4.18E-08 Up-regulated group) pathway ACKR2 Atypical chemokine receptor 2 G-protein–coupled receptor signaling 0.42 3.29E-04 4.41E-03 Down-regulated pathway ACSM1 Acyl-CoA synthetase medium-chain family Energy derivation by oxidation of 9.87 1.70E-08 6.52E-07 Up-regulated member 1 organic compounds ACTC1 Actin, ␣, cardiac muscle 1 Negative regulation of apoptotic 0.30 7.96E-06 1.65E-04 Down-regulated process ACTG2 Actin, ␥2, smooth muscle, enteric Blood microparticle 0.29 1.61E-16 2.36E-14 Down-regulated ADAM33 ADAM domain 33 Integral component of membrane 0.23 9.74E-09 3.95E-07 Down-regulated ADAM8 ADAM domain 8 Positive regulation of tumor necrosis 4.69 2.93E-04 4.01E-03 Up-regulated factor (ligand) superfamily member 11 production ADAMTS18 -

Supplemental Table 1 Proteins Only Present in VLDL Fraction Entrez Gene Name Coagulation System Extrinsic Prothrombin Activation
Supplemental Table 1 Proteins only present in VLDL fraction Intrinsic Extrinsic Prothrombi Entrez Gene Name Coagulation Prothrombin n Cellular System Activation Activation location 1 ADP-ribosylation factor interacting protein 1 - - - Cytoplasm 2 afamin - - - Extracellular 3 alpha-1-B glycoprotein - - - Extracellular 4 alpha-2-macroglobulin + - - Extracellular 5 alpha-2-macroglobulin-like 1 - - - Cytoplasm 6 angiotensinogen (serpin peptidase inhibitor, clade A, member 8) - - - Extracellular 7 angiotensinogen (serpin peptidase inhibitor, clade A, member 8) - - - Extracellular 8 apolipoprotein B - - - Extracellular 9 apolipoprotein B - - - Extracellular 10 apolipoprotein E - - - Extracellular 11 apolipoprotein E - - - Extracellular 12 apolipoprotein E - - - Extracellular 13 ArfGAP with GTPase domain, ankyrin repeat and PH domain 6 - - - Other 14 ATP-binding cassette, sub-family A (ABC1), member 1 - - - Plasma Membrane 15 ATP-binding cassette, sub-family C (CFTR/MRP), member 12 - - - Cytoplasm 16 attractin - - - Extracellular 17 Bardet-Biedl syndrome 4 - - - Cytoplasm 18 carboxypeptidase N, polypeptide 2 - - - Extracellular 19 caspase 8, apoptosis-related cysteine peptidase - - - Nucleus 20 Cbl proto-oncogene, E3 ubiquitin protein ligase - - - Nucleus 21 ceruloplasmin (ferroxidase) - - - Extracellular 22 chemokine (C-X-C motif) ligand 2 - - - Extracellular 23 cholinergic receptor, muscarinic 2 - - - Plasma 1 Membrane 24 chromosome 6 open reading frame 163 - - - Other 25 clathrin, heavy chain (Hc) - - - Plasma Membrane 26 coagulation factor -

Using IEG's to Uncover Pathways for Spatial Learning in The
Using IEG’s to uncover pathways for spatial learning in the rat A thesis submitted for the degree of doctor of philosophy Katherine L. Shires Department of Psychology/Biosciences, Cardiff University July 2006 UMI Number: U584126 All rights reserved INFORMATION TO ALL USERS The quality of this reproduction is dependent upon the quality of the copy submitted. In the unlikely event that the author did not send a complete manuscript and there are missing pages, these will be noted. Also, if material had to be removed, a note will indicate the deletion. Dissertation Publishing UMI U584126 Published by ProQuest LLC 2013. Copyright in the Dissertation held by the Author. Microform Edition © ProQuest LLC. All rights reserved. This work is protected against unauthorized copying under Title 17, United States Code. ProQuest LLC 789 East Eisenhower Parkway P.O. Box 1346 Ann Arbor, Ml 48106-1346 Declaration This work has not previously been accepted in substance for any degree and is not concurrently submittedTr! candidature for any degree. Signed ...................................... Date ............. Statement 1 This thesis is being ed in partial fulfillment of the requirements for the degree of PhD Signed.. Date ijfi.J Statement 2 This thesis is the result of my own independent work/investigation, except where otherwise stated. Other SourceSafe acknowledged by footnotes giving explicit references. Signed... .............................. Date............... P . j l . J . 0 .7........ Statement 3 I hereby give consent for my thesis, if accepted, to be available for photocopying and for inter-library loan, and for the title and summary to be made available to outside organisation Abstract This thesis concentrates on the induction of the immediate early genes zif268 and c-fos in a functional and dysfunctional brain network. -

Gene Expression Profiling of Paired Ovarian Tumors Obtained Prior to and Following Adjuvant Chemotherapy: Molecular Signatures of Chemoresistant Tumors
5-24 1/6/06 10:25 Page 5 INTERNATIONAL JOURNAL OF ONCOLOGY 29: 5-24, 2006 5 Gene expression profiling of paired ovarian tumors obtained prior to and following adjuvant chemotherapy: Molecular signatures of chemoresistant tumors SYLVAIN L'ESPÉRANCE1,4, ION POPA2,4, MAGDALENA BACHVAROVA4, MARIE PLANTE3,4, NANCY PATTEN5, LIN WU5, BERNARD TÊTU2,4 and DIMCHO BACHVAROV1,4 1Department of Medicine, 2Division of Pathology, 3Department of Gynecologic Oncology, Faculty of Medicine, Laval University, Québec G1K 7P4; 4Cancer Research Centre, Hôpital L'Hotel-Dieu de Québec, Centre Hospitalier Universitaire de Québec, 9 rue McMahon, Québec G1R 2J6, Canada; 5Pharmacogenetics Department, Roche Molecular Systems, Inc., 4300 Hacienda Drive, Pleasanton, CA 94588, USA Received January 18, 2006; Accepted March 27, 2006 Abstract. Chemotherapy (CT) resistance in ovarian cancer is the existence of two distinct expression signatures of chemo- related to multiple factors, and assessment of these factors is resistant tumors, which was further confirmed by assessment necessary for the development of new drugs and therapeutic of some genetic (p53 gene mutation status) and clinical para- regimens. In an effort to identify such determinants, we meters (CT regimens). Our data suggest that intrinsic and evaluated the expression of approximately 21,000 genes using acquired chemoresistant phenotypes of post-CT tumors may be DNA microarray screening in paired tumor samples taken attributed to the combined action of different factors implicated prior to and after CT treatment from 6 patients with pre- in mechanisms of chemoresistance, tumor invasion/progression dominantly advanced stage, high-grade epithelial ovarian and control of cell proliferation. This type of molecular cancer. -
Suppl. Fig. Legends
Supplementary Figures. Supplementary Figure 1. Extent of lesions. a. Extent of smallest (dark gray), average (medium gray), and largest (light gray) lesions in animals treated as indicated 4 weeks after the induction of focal ischemia. b, Quantitation of lesion volume shows that neither treatment alters stroke volume (controls are shown for comparison and are from concurrently run animals reported in ref. 24). Supplementary Figure 2. Combinatorial treatment promotes axon rewiring on the intact side of the spinal cord. a. Low-power camera lucida drawing showing area in the cervical spinal cord contralateral to the undamaged hemisphere with BDA-labeled axon in close proximity to a ventral horn motorneuron in lamina IX. b, c. Photomicrograph (b) and camera lucida drawing (c) of labeled axon shown in a. Quantitation of BDA-labeled axons in lamina IX per mm in the cervical enlargement. *: Differences between groups significant at P < 0.05. Supplementary Figure 3. Sequence homology between the scrambled NEP1-40 peptide and myelin-associated glycoprotein (MAG). Top: Sequence of NEP1-40 and the scrambled peptide. Bottom: Alignment of the scrambled peptide and a region of MAG sharing 12/32 (37%) identities, and 17/32 (53%) positives. Gaps = 8/32 (25%). The probability of this similarity occurring by chance is estimated as 7 x 10-4 (PSI-BlastP of Scrambled NEP vs mouse NR databank at NCBI Blast). Zai et al., Suppl. Fig. 2 Supplementary Table 1 Stereotaxic coordinates for BDA injections Injection site Rostro-caudal Medio-lateral 1 0.12 -0.23 2 0.07 -0.18 3 0.02 -0.18 4 0.02 -0.39 5 -0.03 -0.29 6 -0.03 -0.15 7 -0.03 -0.41 8 -0.08 -0.23 9 -0.08 -0.31 10 -0.13 -0.25 11 -0.13 -0.33 12 -0.18 -0.23 13 -0.18 -0.31 14 -0.23 -0.23 15 -0.23 -0.31 16 -0.28 -0.25 17 -0.28 -0.33 18 -0.33 -0.31 Figures are in cm relative to Bregma Supplementary Table 2.