Altered Mrna Expression Levels of the Major Components of Sphingolipid Metabolism, Ceramide Synthases and Their Clinical Implication in Colorectal Cancer
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

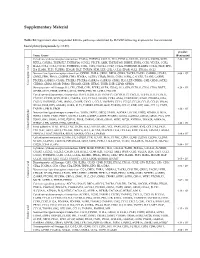
Supplemental Figure 1. Vimentin
Double mutant specific genes Transcript gene_assignment Gene Symbol RefSeq FDR Fold- FDR Fold- FDR Fold- ID (single vs. Change (double Change (double Change wt) (single vs. wt) (double vs. single) (double vs. wt) vs. wt) vs. single) 10485013 BC085239 // 1110051M20Rik // RIKEN cDNA 1110051M20 gene // 2 E1 // 228356 /// NM 1110051M20Ri BC085239 0.164013 -1.38517 0.0345128 -2.24228 0.154535 -1.61877 k 10358717 NM_197990 // 1700025G04Rik // RIKEN cDNA 1700025G04 gene // 1 G2 // 69399 /// BC 1700025G04Rik NM_197990 0.142593 -1.37878 0.0212926 -3.13385 0.093068 -2.27291 10358713 NM_197990 // 1700025G04Rik // RIKEN cDNA 1700025G04 gene // 1 G2 // 69399 1700025G04Rik NM_197990 0.0655213 -1.71563 0.0222468 -2.32498 0.166843 -1.35517 10481312 NM_027283 // 1700026L06Rik // RIKEN cDNA 1700026L06 gene // 2 A3 // 69987 /// EN 1700026L06Rik NM_027283 0.0503754 -1.46385 0.0140999 -2.19537 0.0825609 -1.49972 10351465 BC150846 // 1700084C01Rik // RIKEN cDNA 1700084C01 gene // 1 H3 // 78465 /// NM_ 1700084C01Rik BC150846 0.107391 -1.5916 0.0385418 -2.05801 0.295457 -1.29305 10569654 AK007416 // 1810010D01Rik // RIKEN cDNA 1810010D01 gene // 7 F5 // 381935 /// XR 1810010D01Rik AK007416 0.145576 1.69432 0.0476957 2.51662 0.288571 1.48533 10508883 NM_001083916 // 1810019J16Rik // RIKEN cDNA 1810019J16 gene // 4 D2.3 // 69073 / 1810019J16Rik NM_001083916 0.0533206 1.57139 0.0145433 2.56417 0.0836674 1.63179 10585282 ENSMUST00000050829 // 2010007H06Rik // RIKEN cDNA 2010007H06 gene // --- // 6984 2010007H06Rik ENSMUST00000050829 0.129914 -1.71998 0.0434862 -2.51672 -

Implications in Parkinson's Disease
Journal of Clinical Medicine Review Lysosomal Ceramide Metabolism Disorders: Implications in Parkinson’s Disease Silvia Paciotti 1,2 , Elisabetta Albi 3 , Lucilla Parnetti 1 and Tommaso Beccari 3,* 1 Laboratory of Clinical Neurochemistry, Department of Medicine, University of Perugia, Sant’Andrea delle Fratte, 06132 Perugia, Italy; [email protected] (S.P.); [email protected] (L.P.) 2 Section of Physiology and Biochemistry, Department of Experimental Medicine, University of Perugia, Sant’Andrea delle Fratte, 06132 Perugia, Italy 3 Department of Pharmaceutical Sciences, University of Perugia, Via Fabretti, 06123 Perugia, Italy; [email protected] * Correspondence: [email protected] Received: 29 January 2020; Accepted: 20 February 2020; Published: 21 February 2020 Abstract: Ceramides are a family of bioactive lipids belonging to the class of sphingolipids. Sphingolipidoses are a group of inherited genetic diseases characterized by the unmetabolized sphingolipids and the consequent reduction of ceramide pool in lysosomes. Sphingolipidoses include several disorders as Sandhoff disease, Fabry disease, Gaucher disease, metachromatic leukodystrophy, Krabbe disease, Niemann Pick disease, Farber disease, and GM2 gangliosidosis. In sphingolipidosis, lysosomal lipid storage occurs in both the central nervous system and visceral tissues, and central nervous system pathology is a common hallmark for all of them. Parkinson’s disease, the most common neurodegenerative movement disorder, is characterized by the accumulation and aggregation of misfolded α-synuclein that seem associated to some lysosomal disorders, in particular Gaucher disease. This review provides evidence into the role of ceramide metabolism in the pathophysiology of lysosomes, highlighting the more recent findings on its involvement in Parkinson’s disease. Keywords: ceramide metabolism; Parkinson’s disease; α-synuclein; GBA; GLA; HEX A-B; GALC; ASAH1; SMPD1; ARSA * Correspondence [email protected] 1. -

Molecular Profile of Tumor-Specific CD8+ T Cell Hypofunction in a Transplantable Murine Cancer Model
Downloaded from http://www.jimmunol.org/ by guest on September 25, 2021 T + is online at: average * The Journal of Immunology , 34 of which you can access for free at: 2016; 197:1477-1488; Prepublished online 1 July from submission to initial decision 4 weeks from acceptance to publication 2016; doi: 10.4049/jimmunol.1600589 http://www.jimmunol.org/content/197/4/1477 Molecular Profile of Tumor-Specific CD8 Cell Hypofunction in a Transplantable Murine Cancer Model Katherine A. Waugh, Sonia M. Leach, Brandon L. Moore, Tullia C. Bruno, Jonathan D. Buhrman and Jill E. Slansky J Immunol cites 95 articles Submit online. Every submission reviewed by practicing scientists ? is published twice each month by Receive free email-alerts when new articles cite this article. Sign up at: http://jimmunol.org/alerts http://jimmunol.org/subscription Submit copyright permission requests at: http://www.aai.org/About/Publications/JI/copyright.html http://www.jimmunol.org/content/suppl/2016/07/01/jimmunol.160058 9.DCSupplemental This article http://www.jimmunol.org/content/197/4/1477.full#ref-list-1 Information about subscribing to The JI No Triage! Fast Publication! Rapid Reviews! 30 days* Why • • • Material References Permissions Email Alerts Subscription Supplementary The Journal of Immunology The American Association of Immunologists, Inc., 1451 Rockville Pike, Suite 650, Rockville, MD 20852 Copyright © 2016 by The American Association of Immunologists, Inc. All rights reserved. Print ISSN: 0022-1767 Online ISSN: 1550-6606. This information is current as of September 25, 2021. The Journal of Immunology Molecular Profile of Tumor-Specific CD8+ T Cell Hypofunction in a Transplantable Murine Cancer Model Katherine A. -

Environmental Influences on Endothelial Gene Expression
ENDOTHELIAL CELL GENE EXPRESSION John Matthew Jeff Herbert Supervisors: Prof. Roy Bicknell and Dr. Victoria Heath PhD thesis University of Birmingham August 2012 University of Birmingham Research Archive e-theses repository This unpublished thesis/dissertation is copyright of the author and/or third parties. The intellectual property rights of the author or third parties in respect of this work are as defined by The Copyright Designs and Patents Act 1988 or as modified by any successor legislation. Any use made of information contained in this thesis/dissertation must be in accordance with that legislation and must be properly acknowledged. Further distribution or reproduction in any format is prohibited without the permission of the copyright holder. ABSTRACT Tumour angiogenesis is a vital process in the pathology of tumour development and metastasis. Targeting markers of tumour endothelium provide a means of targeted destruction of a tumours oxygen and nutrient supply via destruction of tumour vasculature, which in turn ultimately leads to beneficial consequences to patients. Although current anti -angiogenic and vascular targeting strategies help patients, more potently in combination with chemo therapy, there is still a need for more tumour endothelial marker discoveries as current treatments have cardiovascular and other side effects. For the first time, the analyses of in-vivo biotinylation of an embryonic system is performed to obtain putative vascular targets. Also for the first time, deep sequencing is applied to freshly isolated tumour and normal endothelial cells from lung, colon and bladder tissues for the identification of pan-vascular-targets. Integration of the proteomic, deep sequencing, public cDNA libraries and microarrays, delivers 5,892 putative vascular targets to the science community. -

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

743914V1.Full.Pdf
bioRxiv preprint doi: https://doi.org/10.1101/743914; this version posted August 24, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. 1 Cross-talks of glycosylphosphatidylinositol biosynthesis with glycosphingolipid biosynthesis 2 and ER-associated degradation 3 4 Yicheng Wang1,2, Yusuke Maeda1, Yishi Liu3, Yoko Takada2, Akinori Ninomiya1, Tetsuya 5 Hirata1,2,4, Morihisa Fujita3, Yoshiko Murakami1,2, Taroh Kinoshita1,2,* 6 7 1Research Institute for Microbial Diseases, Osaka University, Suita, Osaka 565-0871, Japan 8 2WPI Immunology Frontier Research Center, Osaka University, Suita, Osaka 565-0871, 9 Japan 10 3Key Laboratory of Carbohydrate Chemistry and Biotechnology, Ministry of Education, 11 School of Biotechnology, Jiangnan University, Wuxi, Jiangsu 214122, China 12 4Current address: Center for Highly Advanced Integration of Nano and Life Sciences (G- 13 CHAIN), Gifu University, 1-1 Yanagido, Gifu-City, Gifu 501-1193, Japan 14 15 *Correspondence and requests for materials should be addressed to T.K. (email: 16 [email protected]) 17 18 19 Glycosylphosphatidylinositol (GPI)-anchored proteins and glycosphingolipids interact with 20 each other in the mammalian plasma membranes, forming dynamic microdomains. How their 21 interaction starts in the cells has been unclear. Here, based on a genome-wide CRISPR-Cas9 22 genetic screen for genes required for GPI side-chain modification by galactose in the Golgi 23 apparatus, we report that b1,3-galactosyltransferase 4 (B3GALT4), also called GM1 24 ganglioside synthase, additionally functions in transferring galactose to the N- 25 acetylgalactosamine side-chain of GPI. -

Understanding the Molecular Pathobiology of Acid Ceramidase Deficiency
Understanding the Molecular Pathobiology of Acid Ceramidase Deficiency By Fabian Yu A thesis submitted in conformity with the requirements for the degree of Doctor of Philosophy Institute of Medical Science University of Toronto © Copyright by Fabian PS Yu 2018 Understanding the Molecular Pathobiology of Acid Ceramidase Deficiency Fabian Yu Doctor of Philosophy Institute of Medical Science University of Toronto 2018 Abstract Farber disease (FD) is a devastating Lysosomal Storage Disorder (LSD) caused by mutations in ASAH1, resulting in acid ceramidase (ACDase) deficiency. ACDase deficiency manifests along a broad spectrum but in its classical form patients die during early childhood. Due to the scarcity of cases FD has largely been understudied. To circumvent this, our lab previously generated a mouse model that recapitulates FD. In some case reports, patients have shown signs of visceral involvement, retinopathy and respiratory distress that may lead to death. Beyond superficial descriptions in case reports, there have been no in-depth studies performed to address these conditions. To improve the understanding of FD and gain insights for evaluating future therapies, we performed comprehensive studies on the ACDase deficient mouse. In the visual system, we reported presence of progressive uveitis. Further tests revealed cellular infiltration, lipid buildup and extensive retinal pathology. Mice developed retinal dysplasia, impaired retinal response and decreased visual acuity. Within the pulmonary system, lung function tests revealed a decrease in lung compliance. Mice developed chronic lung injury that was contributed by cellular recruitment, and vascular leakage. Additionally, we report impairment to lipid homeostasis in the lungs. ii To understand the liver involvement in FD, we characterized the pathology and performed transcriptome analysis to identify gene and pathway changes. -

Targeting Lysophosphatidic Acid in Cancer: the Issues in Moving from Bench to Bedside
View metadata, citation and similar papers at core.ac.uk brought to you by CORE provided by IUPUIScholarWorks cancers Review Targeting Lysophosphatidic Acid in Cancer: The Issues in Moving from Bench to Bedside Yan Xu Department of Obstetrics and Gynecology, Indiana University School of Medicine, 950 W. Walnut Street R2-E380, Indianapolis, IN 46202, USA; [email protected]; Tel.: +1-317-274-3972 Received: 28 August 2019; Accepted: 8 October 2019; Published: 10 October 2019 Abstract: Since the clear demonstration of lysophosphatidic acid (LPA)’s pathological roles in cancer in the mid-1990s, more than 1000 papers relating LPA to various types of cancer were published. Through these studies, LPA was established as a target for cancer. Although LPA-related inhibitors entered clinical trials for fibrosis, the concept of targeting LPA is yet to be moved to clinical cancer treatment. The major challenges that we are facing in moving LPA application from bench to bedside include the intrinsic and complicated metabolic, functional, and signaling properties of LPA, as well as technical issues, which are discussed in this review. Potential strategies and perspectives to improve the translational progress are suggested. Despite these challenges, we are optimistic that LPA blockage, particularly in combination with other agents, is on the horizon to be incorporated into clinical applications. Keywords: Autotaxin (ATX); ovarian cancer (OC); cancer stem cell (CSC); electrospray ionization tandem mass spectrometry (ESI-MS/MS); G-protein coupled receptor (GPCR); lipid phosphate phosphatase enzymes (LPPs); lysophosphatidic acid (LPA); phospholipase A2 enzymes (PLA2s); nuclear receptor peroxisome proliferator-activated receptor (PPAR); sphingosine-1 phosphate (S1P) 1. -
Supplementary Material
Supplementary Material Table S1: Significant downregulated KEGGs pathways identified by DAVID following exposure to five cinnamon- based phenylpropanoids (p < 0.05). p-value Term: Genes (Benjamini) Cytokine-cytokine receptor interaction: FASLG, TNFSF14, CXCL11, IL11, FLT3LG, CCL3L1, CCL3L3, CXCR6, XCR1, 2.43 × 105 RTEL1, CSF2RA, TNFRSF17, TNFRSF14, CCNL2, VEGFB, AMH, TNFRSF10B, INHBE, IFNB1, CCR3, VEGFA, CCR2, IL12A, CCL1, CCL3, CXCL5, TNFRSF25, CCR1, CSF1, CX3CL1, CCL7, CCL24, TNFRSF1B, IL12RB1, CCL21, FIGF, EPO, IL4, IL18R1, FLT1, TGFBR1, EDA2R, HGF, TNFSF8, KDR, LEP, GH2, CCL13, EPOR, XCL1, IFNA16, XCL2 Neuroactive ligand-receptor interaction: OPRM1, THRA, GRIK1, DRD2, GRIK2, TACR2, TACR1, GABRB1, LPAR4, 9.68 × 105 GRIK5, FPR1, PRSS1, GNRHR, FPR2, EDNRA, AGTR2, LTB4R, PRSS2, CNR1, S1PR4, CALCRL, TAAR5, GABRE, PTGER1, GABRG3, C5AR1, PTGER3, PTGER4, GABRA6, GABRA5, GRM1, PLG, LEP, CRHR1, GH2, GRM3, SSTR2, Chlorogenic acid Chlorogenic CHRM3, GRIA1, MC2R, P2RX2, TBXA2R, GHSR, HTR2C, TSHR, LHB, GLP1R, OPRD1 Hematopoietic cell lineage: IL4, CR1, CD8B, CSF1, FCER2, GYPA, ITGA2, IL11, GP9, FLT3LG, CD38, CD19, DNTT, 9.29 × 104 GP1BB, CD22, EPOR, CSF2RA, CD14, THPO, EPO, HLA-DRA, ITGA2B Cytokine-cytokine receptor interaction: IL6ST, IL21R, IL19, TNFSF15, CXCR3, IL15, CXCL11, TGFB1, IL11, FLT3LG, CXCL10, CCR10, XCR1, RTEL1, CSF2RA, IL21, CCNL2, VEGFB, CCR8, AMH, TNFRSF10C, IFNB1, PDGFRA, EDA, CXCL5, TNFRSF25, CSF1, IFNW1, CNTFR, CX3CL1, CCL5, TNFRSF4, CCL4, CCL27, CCL24, CCL25, CCL23, IFNA6, IFNA5, FIGF, EPO, AMHR2, IL2RA, FLT4, TGFBR2, EDA2R, -

Supplementary Table S4. FGA Co-Expressed Gene List in LUAD
Supplementary Table S4. FGA co-expressed gene list in LUAD tumors Symbol R Locus Description FGG 0.919 4q28 fibrinogen gamma chain FGL1 0.635 8p22 fibrinogen-like 1 SLC7A2 0.536 8p22 solute carrier family 7 (cationic amino acid transporter, y+ system), member 2 DUSP4 0.521 8p12-p11 dual specificity phosphatase 4 HAL 0.51 12q22-q24.1histidine ammonia-lyase PDE4D 0.499 5q12 phosphodiesterase 4D, cAMP-specific FURIN 0.497 15q26.1 furin (paired basic amino acid cleaving enzyme) CPS1 0.49 2q35 carbamoyl-phosphate synthase 1, mitochondrial TESC 0.478 12q24.22 tescalcin INHA 0.465 2q35 inhibin, alpha S100P 0.461 4p16 S100 calcium binding protein P VPS37A 0.447 8p22 vacuolar protein sorting 37 homolog A (S. cerevisiae) SLC16A14 0.447 2q36.3 solute carrier family 16, member 14 PPARGC1A 0.443 4p15.1 peroxisome proliferator-activated receptor gamma, coactivator 1 alpha SIK1 0.435 21q22.3 salt-inducible kinase 1 IRS2 0.434 13q34 insulin receptor substrate 2 RND1 0.433 12q12 Rho family GTPase 1 HGD 0.433 3q13.33 homogentisate 1,2-dioxygenase PTP4A1 0.432 6q12 protein tyrosine phosphatase type IVA, member 1 C8orf4 0.428 8p11.2 chromosome 8 open reading frame 4 DDC 0.427 7p12.2 dopa decarboxylase (aromatic L-amino acid decarboxylase) TACC2 0.427 10q26 transforming, acidic coiled-coil containing protein 2 MUC13 0.422 3q21.2 mucin 13, cell surface associated C5 0.412 9q33-q34 complement component 5 NR4A2 0.412 2q22-q23 nuclear receptor subfamily 4, group A, member 2 EYS 0.411 6q12 eyes shut homolog (Drosophila) GPX2 0.406 14q24.1 glutathione peroxidase -

Identification of Potential Key Genes and Pathway Linked with Sporadic Creutzfeldt-Jakob Disease Based on Integrated Bioinformatics Analyses
medRxiv preprint doi: https://doi.org/10.1101/2020.12.21.20248688; this version posted December 24, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted medRxiv a license to display the preprint in perpetuity. All rights reserved. No reuse allowed without permission. Identification of potential key genes and pathway linked with sporadic Creutzfeldt-Jakob disease based on integrated bioinformatics analyses Basavaraj Vastrad1, Chanabasayya Vastrad*2 , Iranna Kotturshetti 1. Department of Biochemistry, Basaveshwar College of Pharmacy, Gadag, Karnataka 582103, India. 2. Biostatistics and Bioinformatics, Chanabasava Nilaya, Bharthinagar, Dharwad 580001, Karanataka, India. 3. Department of Ayurveda, Rajiv Gandhi Education Society`s Ayurvedic Medical College, Ron, Karnataka 562209, India. * Chanabasayya Vastrad [email protected] Ph: +919480073398 Chanabasava Nilaya, Bharthinagar, Dharwad 580001 , Karanataka, India NOTE: This preprint reports new research that has not been certified by peer review and should not be used to guide clinical practice. medRxiv preprint doi: https://doi.org/10.1101/2020.12.21.20248688; this version posted December 24, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted medRxiv a license to display the preprint in perpetuity. All rights reserved. No reuse allowed without permission. Abstract Sporadic Creutzfeldt-Jakob disease (sCJD) is neurodegenerative disease also called prion disease linked with poor prognosis. The aim of the current study was to illuminate the underlying molecular mechanisms of sCJD. The mRNA microarray dataset GSE124571 was downloaded from the Gene Expression Omnibus database. Differentially expressed genes (DEGs) were screened. -

Cholesterol-Dependent Increases in Glucosylceramide
Neuropharmacology 110 (2016) 458e469 Contents lists available at ScienceDirect Neuropharmacology journal homepage: www.elsevier.com/locate/neuropharm Cholesterol-dependent increases in glucosylceramide synthase activity in Niemann-Pick disease type C model cells: Abnormal trafficking of endogenously formed ceramide metabolites by inhibition of the enzyme Naohiro Hashimoto 1, Ikiru Matsumoto 1, Hiromasa Takahashi, Hitomi Ashikawa, * Hiroyuki Nakamura , Toshihiko Murayama Laboratory of Chemical Pharmacology, Graduate School of Pharmaceutical Sciences, Chiba University, Chiba 260-8675, Japan article info abstract Article history: Sphingolipids such as sphingomyelin and glycosphingolipids (GSLs) derived from glucosylceramide Received 1 February 2016 (GlcCer), in addition to cholesterol, accumulate in cells/neurons in Niemann-Pick disease type C (NPC). Received in revised form The activities of acid sphingomyelinase and lysosomal glucocerebrosidase (GCase), which degrade 10 August 2016 sphingomyelin and GlcCer, respectively, are down-regulated in NPC cells, however, changes in GlcCer Accepted 12 August 2016 synthase activity have not yet been elucidated. We herein demonstrated for the first time that GlcCer Available online 15 August 2016 synthase activity for the fluorescent ceramide, 4-nitrobenzo-2-oxa-1,3-diazole-labeled C6-ceramide À À (NBD-ceramide) increased in intact NPC1( / ) cells and cell lysates without affecting the protein levels. Keywords: (À/À) fl Niemann-Pick disease type C In NBD-ceramide-labeled NPC1 cells, NBD- uorescence preferentially accumulated in the Golgi Cholesterol complex and vesicular specks in the cytoplasm 40 and 150 min, respectively, after labeling, while a Glucosylceramide synthase treatment for 48 h with the GlcCer synthase inhibitors, N-butyldeoxynojirimycin (NB-DNJ) and 1-phenyl- Sphingomyelin 2-palmitoylamino-3-morpholino-1-propanol, accelerated the appearance of vesicular specks emitting À À Vesicular trafficking NBD-fluorescence within 40 min.