Extract, Transform, Load Framework for the Conversion of Health Databases to OMOP
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Guide to Secure Software Development in Ruby
Fedora Security Team Secure Ruby Development Guide Guide to secure software development in Ruby Ján Rusnačko Secure Ruby Development Guide Fedora Security Team Secure Ruby Development Guide Guide to secure software development in Ruby Edition 1 Author Ján Rusnačko [email protected] Copyright © 2014 Ján Rusnačko. The text of and illustrations in this document are licensed by Red Hat under a Creative Commons Attribution–Share Alike 3.0 Unported license ("CC-BY-SA"). An explanation of CC-BY-SA is available at http://creativecommons.org/licenses/by-sa/3.0/. The original authors of this document, and Red Hat, designate the Fedora Project as the "Attribution Party" for purposes of CC-BY-SA. In accordance with CC-BY-SA, if you distribute this document or an adaptation of it, you must provide the URL for the original version. Red Hat, as the licensor of this document, waives the right to enforce, and agrees not to assert, Section 4d of CC-BY-SA to the fullest extent permitted by applicable law. Red Hat, Red Hat Enterprise Linux, the Shadowman logo, JBoss, MetaMatrix, Fedora, the Infinity Logo, and RHCE are trademarks of Red Hat, Inc., registered in the United States and other countries. For guidelines on the permitted uses of the Fedora trademarks, refer to https://fedoraproject.org/wiki/ Legal:Trademark_guidelines. Linux® is the registered trademark of Linus Torvalds in the United States and other countries. Java® is a registered trademark of Oracle and/or its affiliates. XFS® is a trademark of Silicon Graphics International Corp. or its subsidiaries in the United States and/or other countries. -

WEB2PY Enterprise Web Framework (2Nd Edition)
WEB2PY Enterprise Web Framework / 2nd Ed. Massimo Di Pierro Copyright ©2009 by Massimo Di Pierro. All rights reserved. No part of this publication may be reproduced, stored in a retrieval system, or transmitted in any form or by any means, electronic, mechanical, photocopying, recording, scanning, or otherwise, except as permitted under Section 107 or 108 of the 1976 United States Copyright Act, without either the prior written permission of the Publisher, or authorization through payment of the appropriate per-copy fee to the Copyright Clearance Center, Inc., 222 Rosewood Drive, Danvers, MA 01923, (978) 750-8400, fax (978) 646-8600, or on the web at www.copyright.com. Requests to the Copyright owner for permission should be addressed to: Massimo Di Pierro School of Computing DePaul University 243 S Wabash Ave Chicago, IL 60604 (USA) Email: [email protected] Limit of Liability/Disclaimer of Warranty: While the publisher and author have used their best efforts in preparing this book, they make no representations or warranties with respect to the accuracy or completeness of the contents of this book and specifically disclaim any implied warranties of merchantability or fitness for a particular purpose. No warranty may be created ore extended by sales representatives or written sales materials. The advice and strategies contained herein may not be suitable for your situation. You should consult with a professional where appropriate. Neither the publisher nor author shall be liable for any loss of profit or any other commercial damages, including but not limited to special, incidental, consequential, or other damages. Library of Congress Cataloging-in-Publication Data: WEB2PY: Enterprise Web Framework Printed in the United States of America. -

Original.Pdf
Rails Security Primer I am not a software security expert CVE? Common Vulnerabilities and Exposures Vulnerability A weakness that an attacker can use to exploit a system Exploit A piece of software that exploits a vulnerability to achieve unintended or unanticipated behavior CVE-2012-5664 SQL Injection Vulnerability SQL Injection Vulnerability …but only exploitable if you used Authlogic or find_by_* methods in a certain way A cookie like { "session_id" => "41414141", "user_credentials" => "Phenoelit", "user_credentials_id" => { :select=> " *,\"Phenoelit\" as persistence_token from users -- " } } …would create a query like this User.find_by_id(params[:user_credendtials_id]) …would create a query like this User.find_by_id(params[:user_credendtials_id]) User.find_by_id({:select =>"*,\"Phenoelit\" as persistence_token from users --"}) …would create a query like this User.find_by_id(params[:user_credendtials_id]) User.find_by_id({:select =>"*,\"Phenoelit\" as persistence_token from users --"}) SELECT *,"Phenoelit" as persistence_token from users -- FROM "users" WHERE "users"."id" IS NULL LIMIT 1 Blood in the water… CVE-2013-0155 CVE-2013-0156 CVE-2013-0269 CVE-2013-0333 CVE-2013-0155 "Unsafe Query Generation Risk in Ruby on Rails" def reset_password if (@user = User.find_by_token(params[:token])) @user.reset_password! render :json => 'Success' else render :json => 'Failure' end end # POST to http://localhost:3000/users/ reset_password with "{\"token\":[null]}" CVE-2013-0156 "Multiple vulnerabilities in parameter parsing in Action Pack" Content-Type: -

Azure Forum DK Survey
#msdkpartner #msdkpartner Meeting Ground Rules Please post your questions in the chat – We aim to keep QnA at the end of each session Please mute yourself to ensure a good audio experience during presentations This meeting will be recorded #msdkpartner Today's Agenda 08:30 - 08:35 Welcome 08:35 - 09:15 Best of Build 09:15 - 10:00 Top 5 Reasons to chose azure (vs. on-premise) 10:05 - 10:25 Azure in SMB 10:25 - 10:30 Closing #msdkpartner #msdkpartner Hello! I’m Sherry List Azure Developer Engagement Lead Microsoft You can find me at @SherrryLst | @msdev_dk DevOps with Azure, GitHub, and Azure DevOps 500M apps and microservices will be written in the next five years Source: IDC Developer Velocity 100x 200x 7x 8x faster to set up a more frequent fewer failures on more likely to have dev environment code deployments deployments integrated security Source: DORA / Sonatype GitHub Actions for Azure https://github.com/azure/actions Azure Pipelines AKS & k8s support YAML CI Pipelines YAML CD Pipelines Elastic self-hosted agents Community and Collaboration In modern applications 90% of the code comes Your Code from open source Open Source Most of that code lives on GitHub Sign up for Codespaces Preview today https://github.co/codespaces Security and Compliance 70 Security and Compliance 12 56 10 42 7 LOC (M) LOC 28 5 Security Issues (k) Issues Security 14 2 Lines of code Security threats 0 0 Apr Jul Oct Jan Apr Jul Oct Jan Apr Jul Oct Jan Apr Jul Oct Jan Apr Jul Oct Jan Apr 2015 2015 2015 2016 2016 2016 2016 2017 2017 2017 2017 2018 2018 2018 -

Web Development Frameworks Ruby on Rails VS Google Web Toolkit
Bachelor thesis Web Development Frameworks Ruby on Rails VS Google Web Toolkit Author: Carlos Gallardo Adrián Extremera Supervisor: Welf Löwe Semester: Spring 2011 Course code: 2DV00E SE-391 82 Kalmar / SE-351 95 Växjö Tel +46 (0)772-28 80 00 [email protected] Lnu.se/dfm Abstract Web programming is getting more and more important every day and as a consequence, many new tools are created in order to help developers design and construct applications quicker, easier and better structured. Apart from different IDEs and Technologies, nowadays Web Frameworks are gaining popularity amongst users since they offer a large range of methods, classes, etc. that allow programmers to create and maintain solid Web systems. This research focuses on two different Web Frameworks: Ruby on Rails and Google Web Toolkit and within this document we will examine some of the most important differences between them during a Web development. Keywords web frameworks, Ruby, Rails, Model-View-Controller, web programming, Java, Google Web Toolkit, web development, code lines i List of Figures Figure 2.1. mraible - History of Web Frameworks....................................................4 Figure 2.2. Java BluePrints - MVC Pattern..............................................................6 Figure 2.3. Libros Web - MVC Architecture.............................................................7 Figure 2.4. Ruby on Rails - Logo.............................................................................8 Figure 2.5. Windaroo Consulting Inc - Ruby on Rails Structure.............................10 -

FULLTEXT01.Pdf
UPTEC F 18029 Examensarbete 30 hp Juni 2018 Investigation and Implementation of a Log Management and Analysis Framework for the Treatment Planning System RayStation Elias Norrby Abstract Investigation and Implementation of a Log Management and Analysis Framework for the Treatment Planning System RayStation Elias Norrby Teknisk- naturvetenskaplig fakultet UTH-enheten The purpose of this thesis is to investigate and implement a framework for log management and analysis tailored to the treatment planning system (TPS) Besöksadress: RayStation. A TPS is a highly advanced software package used in radiation Ångströmlaboratoriet Lägerhyddsvägen 1 oncology clinics, and the complexity of the software makes writing robust code Hus 4, Plan 0 challenging. Although the product is tested rigorously during development, bugs are present in released software. The purpose of the the framework is to allow the Postadress: RayStation development team insight into errors encountered in clinics by Box 536 751 21 Uppsala centralizing log file data recorded at clinics around the world. Telefon: A framework based on the Elastic stack, a suite of open-source products, is 018 – 471 30 03 proposed, addressing a set of known issues described as the access problem, the Telefax: processing problem, and the analysis problem. Firstly, log files are stored locally on 018 – 471 30 00 each machine running RayStation, some of which may not be connected to the Internet. Gaining access to the data is further complicated by legal frameworks Hemsida: such as HIPAA and GDPR that put constraints on how clinic data can be handled. http://www.teknat.uu.se/student The framework allows for access to the files while respecting these constraints. -

Fiz: a Component Framework for Web Applications
Fiz: A Component Framework for Web Applications John K. Ousterhout Department of Computer Science Stanford University Abstract Fiz is a framework for developing interactive Web applications. Its overall goal is to raise the level of programming for Web applications, first by providing a set of high-level reusable components that simplify the task of creating interactive Web applications, and second by providing a framework that encourages other people to create addi- tional components. Components in Fiz cover both the front-end of Web applications (managing a browser-based user interface) and the back end (managing the application's data). Fiz makes it possible to create components that encapsulate complex behaviors such as Ajax-based updates, hiding many of the Web's complexities from applica- tion developers. Because of its focus on components, Fiz does not use mechanisms such as templates and model- view-controller in the same way as other frameworks. ger and more useful structures. We will release Fiz in 1 Introduction open-source form and hope to build a user community Although the World-Wide Web was initially conceived that creates an ever-increasing set of interesting com- as a vehicle for delivering and viewing documents, its ponents, which will make it dramatically easier to cre- focus has gradually shifted from documents to applica- ate applications that advance the state-of-the-art in Web tions. Facilities such as Javascript, the Document Ob- interactivity. ject Model (DOM), and Ajax have made it possible to offer sophisticated interactive applications over the The rest of this paper is organized as follows. -

Thomas Holloway Resume
Thomas Holloway Austin, TX / 786-512-2964 / [email protected] nyxtom.dev / github.com/nyxtom / linkedin.com/in/thomas-holloway Senior Software Developer - Netcuras Inc. - Austin, TX - Nov 2015 - Present Technical lead and core contributor for implementing d3.js dashboards designed to solve the problem of infrastructure monitoring without a complicated user experience. ○ Built a dashboard capabilities system to monitor 1000s of devices with automated discovery, metrics aggregation, inferable widgets, plugins and system/company-wide JSON queryable YAML definitions ○ Wrote plugins for Netapp, VMWare, Meraki, MongoDB, Elastic, PostgreSQL, Netflow, Syslog, TCP, and UDP within a distributed collector architecture. ○ To handle Netflow/Syslog traffic I implemented a backlog capable Node.js stream followed by debugging, instrumenting, and profiling with cpu tracing and heap snapshots to maintain consistent throughput and handle network connectivity issues. To provide customers with high level visibility of network traffic, I wrote a number of different visualizations, top lists, and search aggregation queries for the dashboard UI. ○ Integrated Mapbox for visualizing 100s of devices in a hierarchical clustered layout ○ Built a connected graph in d3.js with a force-graph layout to provide visibility of SNMP networks ○ Improved responsiveness of web app from reading through React and Aurelia’s source code to understand and build upon on cooperative scheduling and binding behaviors with cached queries, jsonpath expression templates, virtualized components, and web workers (JavaScript, HTML/CSS, Aurelia, React.js, MongoDB, Redis, TCP/IP, REST, Elastic, D3.js, AJAX, Node.js, Express.js, Python, Go, LevelDB, TCP/IP, Jenkins, Ansible, Shell, Cent OS, VMWare, Netflow, Syslog, UDP, SNMP, JSONPath, Webpack, Babel) Creator - Nuvi.com - Salt Lake City, UT - 2011 - 2014 Launched social media analytics platform for +1000s of brands processing +1000000s of tweets, facebook posts, web articles within high fidelity visualizations, reporting, and dashboard UI. -

AWS App Runner Developer Guide AWS App Runner Developer Guide
AWS App Runner Developer Guide AWS App Runner Developer Guide AWS App Runner: Developer Guide Copyright © Amazon Web Services, Inc. and/or its affiliates. All rights reserved. Amazon's trademarks and trade dress may not be used in connection with any product or service that is not Amazon's, in any manner that is likely to cause confusion among customers, or in any manner that disparages or discredits Amazon. All other trademarks not owned by Amazon are the property of their respective owners, who may or may not be affiliated with, connected to, or sponsored by Amazon. AWS App Runner Developer Guide Table of Contents What is AWS App Runner? .................................................................................................................. 1 Who is App Runner for? .............................................................................................................. 1 Accessing App Runner ................................................................................................................. 1 Pricing for App Runner ............................................................................................................... 2 What's next ............................................................................................................................... 2 Setting up ......................................................................................................................................... 3 Create an AWS account .............................................................................................................. -

Job Description: Riverside Research's Engineering and Support Solutions
Job Description: Riverside Research’s Engineering and Support Solutions Directorate has an opening in the Dayton, Ohio area for a Software Developer. This candidate will have the opportunity to develop a web-based client application and backend server for a highly scalable physics tool while setting direction and leading an assisting support team, with a focus on the use of permissively licensed programming languages and libraries. Job Responsibilities: • Utilizing languages like C and C++ • Working with JavaScript/ECMAscript and its language extensions • User interface development • Creating 3D renderings • Using revision control systems • Other duties as assigned Requirements: • Experience with compiled languages like C and C++ • Experience with Javascript/ECMAscript and its language extensions like dart (dart2js), TypeScript, Google Closure Tools, Google Web Toolkit, Haxe • Experience with user interface development toolkits like Qt, GTK, VTK, etc. • Experience with 3D rendering libraries like OpenGL, OpenSceneGraph, etc. • Experience with revision control systems like git, svn, mercurial, etc. Desired experience: • Experience working in projects comprising >100,000 sloc • Development of browser-centric 3D applications (in particular utilization of WebGL) • Experience combining web client and server backend components into a single thick client application • Development experience on Unix and/or Linux systems • Experience with WebGL • Familiarity with Unix shell scripting and supporting tools like bash, awk, sed • Usage of CAD systems -
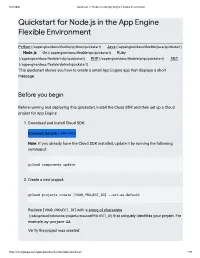
Quicksta for Node.Js in the App Engine Flexible Environment
8/23/2020 Quickstart for Node.js in the App Engine Flexible Environment Quicksta for Node.js in the App Engine Flexible Environment Python (/appengine/docs/exible/python/quickstart) | Java (/appengine/docs/exible/java/quickstart) | Node.js | Go (/appengine/docs/exible/go/quickstart) | Ruby (/appengine/docs/exible/ruby/quickstart) | PHP (/appengine/docs/exible/php/quickstart) | .NET (/appengine/docs/exible/dotnet/quickstart) This quickstart shows you how to create a small App Engine app that displays a short message. Before you begin Before running and deploying this quickstart, install the Cloud SDK and then set up a Cloud project for App Engine: 1. Download and install Cloud SDK: Download the SDK (/sdk/docs) Note: If you already have the Cloud SDK installed, update it by running the following command: gcloud components update 2. Create a new project: gcloud projects create [YOUR_PROJECT_ID] --set-as-default Replace [YOUR_PROJECT_ID] with a string of characters (/sdk/gcloud/reference/projects/create#PROJECT_ID) that uniquely identies your project. For example, my-project-24. Verify the project was created: https://cloud.google.com/appengine/docs/flexible/nodejs/quickstart/ 1/10 8/23/2020 Quickstart for Node.js in the App Engine Flexible Environment gcloud projects describe [YOUR_PROJECT_ID] You see project details that might look like the following: createTime: year-month-hour lifecycleState: ACTIVE name: project-name parent: id: '433637338589' type: organization projectId: project-name-id projectNumber: 499227785679 3. Initialize your App Engine app with your project and choose its region: gcloud app create --project=[YOUR_PROJECT_ID] When prompted, select the region (#before-you-begin) where you want your App Engine application located. -

Arcgis API for Javascript: Building Mobile Web Apps
Building Mobile Apps Matt Driscoll, Lloyd Heberlie @driskull @lheberlie Welcome • Introductions • Expectations • Agenda • Resources Why are we here? Mobile resources in the API Lloyd Heberlie Designing for mobile Lloyd Heberlie Touch aware map Geocoder, LocateButton esri/dijit/Geocoder esri/dijit/LocateButton Popup Mobile esri/dijit/PopupMobile Basemap Gallery esri/dijit/BasemapGallery Basemap Toggle esri/dijit/BasemapToggle Custom components Custom basemap switcher Designing for mobile Lloyd Heberlie Productivity and code management Lloyd Heberlie Setup a developer machine Code quality and verification web server Source Control Code formatting and management Automation and continuous integration Code validation • JSLint • JSHint CSS preprocessors Productivity and code management Lloyd Heberlie Debugging and testing for mobile Lloyd Heberlie Emulators / Simulators Opera Mobile Emulator iOS Simulator * Windows Phone Emulator * Android Emulator Physical devices http://blog.adtile.me/2014/01/08/adtile-device-lab/ Remote debugging • Safari Web Inspector Remote • Google Chrome remote debugging • Adobe Edge Inspect - Synchronized screen capture, device details - Node.js (weinre local) • Web Inspector Remote (weinre) Enable Safari remote web inspection Safari remote debugging Safari remote debugging Chrome remote debugging Chrome remote debugging Adobe Edge Inspect Adobe Edge Inspect Adobe Edge Inspect device_model = LGE Nexus 5 device_res = 1080x1776 device_model = iPhone device_model = iPad orientation = portrait device_res = 1136x640 device_res