Analysis of BMP4 and BMP7 Signaling in Breast Cancer Cells Unveils Time
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Epidermal Growth Factor Receptor (EGFR) Mutation Analysis, Gene
Peraldo-Neia et al. BMC Cancer 2011, 11:31 http://www.biomedcentral.com/1471-2407/11/31 RESEARCHARTICLE Open Access Epidermal Growth Factor Receptor (EGFR) mutation analysis, gene expression profiling and EGFR protein expression in primary prostate cancer Caterina Peraldo-Neia1,2*, Giorgia Migliardi1, Maurizia Mello-Grand2, Filippo Montemurro3, Raffaella Segir2, Ymera Pignochino1, Giuliana Cavalloni1, Bruno Torchio4, Luciano Mosso4, Giovanna Chiorino2, Massimo Aglietta1,3 Abstract Background: Activating mutations of the epidermal growth factor receptor (EGFR) confer sensitivity to the tyrosine kinase inhibitors (TKi), gefitinib and erlotinib. We analysed EGFR expression, EGFR mutation status and gene expression profiles of prostate cancer (PC) to supply a rationale for EGFR targeted therapies in this disease. Methods: Mutational analysis of EGFR TK domain (exons from 18 to 21) and immunohistochemistry for EGFR were performed on tumour tissues derived from radical prostatectomy from 100 PC patients. Gene expression profiling using oligo-microarrays was also carried out in 51 of the PC samples. Results: EGFR protein overexpression (EGFRhigh) was found in 36% of the tumour samples, and mutations were found in 13% of samples. Patients with EGFRhigh tumours experienced a significantly increased risk of biochemical relapse (hazard ratio-HR 2.52, p=0.02) compared with patients with tumours expressing low levels of EGFR (EGFRlow). Microarray analysis did not reveal any differences in gene expression between EGFRhigh and EGFRlow tumours. Conversely, in EGFRhigh tumours, we were able to identify a 79 gene signature distinguishing mutated from non-mutated tumours. Additionally, 29 genes were found to be differentially expressed between mutated/ EGFRhigh (n=3) and mutated/EGFRlow tumours (n=5). -

Supplementary Table S4. FGA Co-Expressed Gene List in LUAD
Supplementary Table S4. FGA co-expressed gene list in LUAD tumors Symbol R Locus Description FGG 0.919 4q28 fibrinogen gamma chain FGL1 0.635 8p22 fibrinogen-like 1 SLC7A2 0.536 8p22 solute carrier family 7 (cationic amino acid transporter, y+ system), member 2 DUSP4 0.521 8p12-p11 dual specificity phosphatase 4 HAL 0.51 12q22-q24.1histidine ammonia-lyase PDE4D 0.499 5q12 phosphodiesterase 4D, cAMP-specific FURIN 0.497 15q26.1 furin (paired basic amino acid cleaving enzyme) CPS1 0.49 2q35 carbamoyl-phosphate synthase 1, mitochondrial TESC 0.478 12q24.22 tescalcin INHA 0.465 2q35 inhibin, alpha S100P 0.461 4p16 S100 calcium binding protein P VPS37A 0.447 8p22 vacuolar protein sorting 37 homolog A (S. cerevisiae) SLC16A14 0.447 2q36.3 solute carrier family 16, member 14 PPARGC1A 0.443 4p15.1 peroxisome proliferator-activated receptor gamma, coactivator 1 alpha SIK1 0.435 21q22.3 salt-inducible kinase 1 IRS2 0.434 13q34 insulin receptor substrate 2 RND1 0.433 12q12 Rho family GTPase 1 HGD 0.433 3q13.33 homogentisate 1,2-dioxygenase PTP4A1 0.432 6q12 protein tyrosine phosphatase type IVA, member 1 C8orf4 0.428 8p11.2 chromosome 8 open reading frame 4 DDC 0.427 7p12.2 dopa decarboxylase (aromatic L-amino acid decarboxylase) TACC2 0.427 10q26 transforming, acidic coiled-coil containing protein 2 MUC13 0.422 3q21.2 mucin 13, cell surface associated C5 0.412 9q33-q34 complement component 5 NR4A2 0.412 2q22-q23 nuclear receptor subfamily 4, group A, member 2 EYS 0.411 6q12 eyes shut homolog (Drosophila) GPX2 0.406 14q24.1 glutathione peroxidase -

Alzheimer's Disease
VU Research Portal Spot the difference Bossers, C.A.M. 2009 document version Publisher's PDF, also known as Version of record Link to publication in VU Research Portal citation for published version (APA) Bossers, C. A. M. (2009). Spot the difference: microarray analysis of gene expression changes in Alzheimer's and Parkinson's Disease. General rights Copyright and moral rights for the publications made accessible in the public portal are retained by the authors and/or other copyright owners and it is a condition of accessing publications that users recognise and abide by the legal requirements associated with these rights. • Users may download and print one copy of any publication from the public portal for the purpose of private study or research. • You may not further distribute the material or use it for any profit-making activity or commercial gain • You may freely distribute the URL identifying the publication in the public portal ? Take down policy If you believe that this document breaches copyright please contact us providing details, and we will remove access to the work immediately and investigate your claim. E-mail address: [email protected] Download date: 11. Oct. 2021 Spot the difference: microarray analysis of gene expression changes in Alzheimer’s and Parkinson’s Disease The research described in this thesis was conducted at the Netherlands Institute for Neuroscience, an Institute of the Royal Netherlands Academy of Arts and Sci- ences, Amsterdam, The Netherlands. The research was financially supported by a collaboration between the Netherlands Institute for Neuroscience and Solvay Phar- maceuticals. Leescommissie: dr. -

Downloadable Code for Forms the Data Analysis and Renders a Visual Display
Open Access Software2005KemmeretVolume al. 6, Issue 12, Article R106 Ulysses - an application for the projection of molecular interactions comment across species Danielle Kemmer*†, Yong Huang‡, Sohrab P Shah‡¥, Jonathan Lim†, Jochen Brumm†, Macaire MS Yuen‡, John Ling‡, Tao Xu‡, Wyeth W Wasserman†§ and BF Francis Ouellette‡§¶ * † Addresses: Center for Genomics and Bioinformatics, Karolinska Institutet, 171 77 Stockholm, Sweden. Centre for Molecular Medicine and reviews Therapeutics, University of British Columbia, Vancouver V5Z 4H4, BC, Canada. ‡UBC Bioinformatics Centre, University of British Columbia, Vancouver V6T 1Z4, BC, Canada. §Department of Medical Genetics, University of British Columbia, Vancouver, BC, Canada. ¶Michael Smith Laboratories, University of British Columbia, Vancouver V6T 1Z4, BC, Canada. ¥Department of Computer Science, University of British Columbia, Vancouver V6T 1Z4, BC, Canada. Correspondence: Wyeth W Wasserman. E-mail: [email protected] Published: 2 December 2005 Received: 23 February 2005 Revised: 3 August 2005 reports Genome Biology 2005, 6:R106 (doi:10.1186/gb-2005-6-12-r106) Accepted: 8 November 2005 The electronic version of this article is the complete one and can be found online at http://genomebiology.com/2005/6/12/R106 © 2005 Kemmer et al.; licensee BioMed Central Ltd. This is an Open Access article distributed under the terms of the Creative Commons Attribution License (http://creativecommons.org/licenses/by/2.0), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited. deposited research Projecting<p>Ulysses, molecular a new software interactions for the across parallel species analysis and display of protein interactions detected in various species, is described.</p> Abstract We developed Ulysses as a user-oriented system that uses a process called Interolog Analysis for the parallel analysis and display of protein interactions detected in various species. -

Mouse Dok6 Conditional Knockout Project (CRISPR/Cas9)
https://www.alphaknockout.com Mouse Dok6 Conditional Knockout Project (CRISPR/Cas9) Objective: To create a Dok6 conditional knockout Mouse model (C57BL/6J) by CRISPR/Cas-mediated genome engineering. Strategy summary: The Dok6 gene (NCBI Reference Sequence: NM_001039173 ; Ensembl: ENSMUSG00000073514 ) is located on Mouse chromosome 18. 8 exons are identified, with the ATG start codon in exon 1 and the TGA stop codon in exon 8 (Transcript: ENSMUST00000097495). Exon 3 will be selected as conditional knockout region (cKO region). Deletion of this region should result in the loss of function of the Mouse Dok6 gene. To engineer the targeting vector, homologous arms and cKO region will be generated by PCR using BAC clone RP23-356C16 as template. Cas9, gRNA and targeting vector will be co-injected into fertilized eggs for cKO Mouse production. The pups will be genotyped by PCR followed by sequencing analysis. Note: Exon 3 starts from about 17.62% of the coding region. The knockout of Exon 3 will result in frameshift of the gene. The size of intron 2 for 5'-loxP site insertion: 38566 bp, and the size of intron 3 for 3'-loxP site insertion: 63053 bp. The size of effective cKO region: ~615 bp. The cKO region does not have any other known gene. Page 1 of 7 https://www.alphaknockout.com Overview of the Targeting Strategy Wildtype allele gRNA region 5' gRNA region 3' 1 3 8 Targeting vector Targeted allele Constitutive KO allele (After Cre recombination) Legends Exon of mouse Dok6 Homology arm cKO region loxP site Page 2 of 7 https://www.alphaknockout.com Overview of the Dot Plot Window size: 10 bp Forward Reverse Complement Sequence 12 Note: The sequence of homologous arms and cKO region is aligned with itself to determine if there are tandem repeats. -

Mapiranje I Funkcionalna Karakterizacija Porodice Receptora Tirozin Kinaze Erbb2
Mapiranje i funkcionalna karakterizacija porodice receptora tirozin kinaze ErbB2 Vučetić, Adriana Master's thesis / Diplomski rad 2017 Degree Grantor / Ustanova koja je dodijelila akademski / stručni stupanj: University of Zagreb, Faculty of Science / Sveučilište u Zagrebu, Prirodoslovno-matematički fakultet Permanent link / Trajna poveznica: https://urn.nsk.hr/urn:nbn:hr:217:150116 Rights / Prava: In copyright Download date / Datum preuzimanja: 2021-10-06 Repository / Repozitorij: Repository of Faculty of Science - University of Zagreb University of Zagreb Faculty of Science Department of Biology Adriana Vučetić Mapping and functional characterization of the ErbB receptor tyrosine kinase family using the Mammalian Membrane Two-Hybrid Assay Master thesis Zagreb, 2017. This thesis was performed in Stagljar Lab, Department of Biochemistry and Department of Molecular Genetics, Faculty of Medicine, University of Toronto, Toronto, Canada under supervision of Dr. Igor Štagljar, Prof. Thesis is submitted on evaluation to Department of Biology, Faculty of Science, University of Zagreb, Zagreb, Croatia in order to acquire academic title of Master of Molecular Biology. Acknowledgments I would like to express my gratitude to my parents and family for their unconditional love and support, for believing in me and motivating me. I would like to thank Dr. Igor Štagljar and Educational Foundation Igor Štagljar for giving me this amazing opportunity to work and live in Toronto for seven months. I am grateful to Dr. Laura Riley and Dr. Damjan Franjević who helped me plan my stay in Canada and deal with obligations to University of Zagreb and University of Toronto. I am thankful for each member in Stagljar lab, especially Dr. Punit Saraon for sharing his knowledge and training me, and Shivanthy Pathmanathan for huge moral support and being a great colleague and friend. -

Dual Specificity Phosphatases from Molecular Mechanisms to Biological Function
International Journal of Molecular Sciences Dual Specificity Phosphatases From Molecular Mechanisms to Biological Function Edited by Rafael Pulido and Roland Lang Printed Edition of the Special Issue Published in International Journal of Molecular Sciences www.mdpi.com/journal/ijms Dual Specificity Phosphatases Dual Specificity Phosphatases From Molecular Mechanisms to Biological Function Special Issue Editors Rafael Pulido Roland Lang MDPI • Basel • Beijing • Wuhan • Barcelona • Belgrade Special Issue Editors Rafael Pulido Roland Lang Biocruces Health Research Institute University Hospital Erlangen Spain Germany Editorial Office MDPI St. Alban-Anlage 66 4052 Basel, Switzerland This is a reprint of articles from the Special Issue published online in the open access journal International Journal of Molecular Sciences (ISSN 1422-0067) from 2018 to 2019 (available at: https: //www.mdpi.com/journal/ijms/special issues/DUSPs). For citation purposes, cite each article independently as indicated on the article page online and as indicated below: LastName, A.A.; LastName, B.B.; LastName, C.C. Article Title. Journal Name Year, Article Number, Page Range. ISBN 978-3-03921-688-8 (Pbk) ISBN 978-3-03921-689-5 (PDF) c 2019 by the authors. Articles in this book are Open Access and distributed under the Creative Commons Attribution (CC BY) license, which allows users to download, copy and build upon published articles, as long as the author and publisher are properly credited, which ensures maximum dissemination and a wider impact of our publications. The book as a whole is distributed by MDPI under the terms and conditions of the Creative Commons license CC BY-NC-ND. Contents About the Special Issue Editors .................................... -
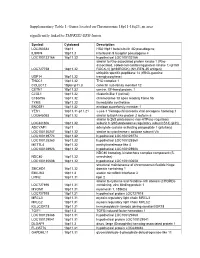
Supplementary Table 1: Genes Located on Chromosome 18P11-18Q23, an Area Significantly Linked to TMPRSS2-ERG Fusion
Supplementary Table 1: Genes located on Chromosome 18p11-18q23, an area significantly linked to TMPRSS2-ERG fusion Symbol Cytoband Description LOC260334 18p11 HSA18p11 beta-tubulin 4Q pseudogene IL9RP4 18p11.3 interleukin 9 receptor pseudogene 4 LOC100132166 18p11.32 hypothetical LOC100132166 similar to Rho-associated protein kinase 1 (Rho- associated, coiled-coil-containing protein kinase 1) (p160 LOC727758 18p11.32 ROCK-1) (p160ROCK) (NY-REN-35 antigen) ubiquitin specific peptidase 14 (tRNA-guanine USP14 18p11.32 transglycosylase) THOC1 18p11.32 THO complex 1 COLEC12 18pter-p11.3 collectin sub-family member 12 CETN1 18p11.32 centrin, EF-hand protein, 1 CLUL1 18p11.32 clusterin-like 1 (retinal) C18orf56 18p11.32 chromosome 18 open reading frame 56 TYMS 18p11.32 thymidylate synthetase ENOSF1 18p11.32 enolase superfamily member 1 YES1 18p11.31-p11.21 v-yes-1 Yamaguchi sarcoma viral oncogene homolog 1 LOC645053 18p11.32 similar to BolA-like protein 2 isoform a similar to 26S proteasome non-ATPase regulatory LOC441806 18p11.32 subunit 8 (26S proteasome regulatory subunit S14) (p31) ADCYAP1 18p11 adenylate cyclase activating polypeptide 1 (pituitary) LOC100130247 18p11.32 similar to cytochrome c oxidase subunit VIc LOC100129774 18p11.32 hypothetical LOC100129774 LOC100128360 18p11.32 hypothetical LOC100128360 METTL4 18p11.32 methyltransferase like 4 LOC100128926 18p11.32 hypothetical LOC100128926 NDC80 homolog, kinetochore complex component (S. NDC80 18p11.32 cerevisiae) LOC100130608 18p11.32 hypothetical LOC100130608 structural maintenance -

Vast Human-Specific Delay in Cortical Ontogenesis Associated With
Supplementary information Extension of cortical synaptic development distinguishes humans from chimpanzees and macaques Supplementary Methods Sample collection We used prefrontal cortex (PFC) and cerebellar cortex (CBC) samples from postmortem brains of 33 human (aged 0-98 years), 14 chimpanzee (aged 0-44 years) and 44 rhesus macaque individuals (aged 0-28 years) (Table S1). Human samples were obtained from the NICHD Brain and Tissue Bank for Developmental Disorders at the University of Maryland, USA, the Netherlands Brain Bank, Amsterdam, Netherlands and the Chinese Brain Bank Center, Wuhan, China. Informed consent for use of human tissues for research was obtained in writing from all donors or their next of kin. All subjects were defined as normal by forensic pathologists at the corresponding brain bank. All subjects suffered sudden death with no prolonged agonal state. Chimpanzee samples were obtained from the Yerkes Primate Center, GA, USA, the Anthropological Institute & Museum of the University of Zürich-Irchel, Switzerland and the Biomedical Primate Research Centre, Netherlands (eight Western chimpanzees, one Central/Eastern and five of unknown origin). Rhesus macaque samples were obtained from the Suzhou Experimental Animal Center, China. All non-human primates used in this study suffered sudden deaths for reasons other than their participation in this study and without any relation to the tissue used. CBC dissections were made from the cerebellar cortex. PFC dissections were made from the frontal part of the superior frontal gyrus. All samples contained an approximately 2:1 grey matter to white matter volume ratio. RNA microarray hybridization RNA isolation, hybridization to microarrays, and data preprocessing were performed as described previously (Khaitovich et al. -

Genome-Wide Association Studies Identifying Functionally Related Genes and Intragenic Regions in Small Sample Studies Knut M
Rockefeller University Digital Commons @ RU Krueger Laboratory Laboratories and Research 2013 From Single-SNP to Wide-Locus: Genome-Wide Association Studies Identifying Functionally Related Genes and Intragenic Regions in Small Sample Studies Knut M. Wittkowski Vikas Sonakya Tingting Song Martin P. Seybold Mehdi Keddache See next page for additional authors Follow this and additional works at: http://digitalcommons.rockefeller.edu/krueger_laboratory Part of the Life Sciences Commons Recommended Citation Pharmacogenomics (2013), 14 (4), p. 391-401 This Article is brought to you for free and open access by the Laboratories and Research at Digital Commons @ RU. It has been accepted for inclusion in Krueger Laboratory by an authorized administrator of Digital Commons @ RU. For more information, please contact [email protected]. Authors Knut M. Wittkowski, Vikas Sonakya, Tingting Song, Martin P. Seybold, Mehdi Keddache, and Martina Durner This article is available at Digital Commons @ RU: http://digitalcommons.rockefeller.edu/krueger_laboratory/1 WITTKOWSKI KM et al. Pharmacogenomics (2013) 14 (4), 391–401 -1- From Single-SNP to Wide-Locus: genome-wide association studies identifying functionally related genes and intragenic regions in small sample studies KNUT M. WITTKOWSKI 1) , VIKAS SONAKYA 1) , TINGTING SONG 1) , MARTIN P. SEYBOLD 2), MEHDI KEDDACHE 3), MARTINA DURNER 4) 1) The Rockefeller University, Center for Clinical and Translational Science 1230 York Ave Box 322, New York, NY 10021, U.S.A. [email protected] 2) Stuttgart University, Institut für Formale Methoden der Informatik Universitaetstrasse 38, D-70569 Stuttgart, Germany 3) Cincinnati Children's Hospital Medical Center, 3333 Burnet Avenue, Cincinnati, Ohio 45229-3039 4) Mount Sinai School of Medicine, Department of Psychiatry, One Gustave Levy Place, Box 1230, New York, NY 10029 Abstract: Background: Genome Wide Association Studies (GWAS) have had limited success when ap- plied to complex diseases. -

Transcriptome Profiling Reveals the Complexity of Pirfenidone Effects in IPF
ERJ Express. Published on August 30, 2018 as doi: 10.1183/13993003.00564-2018 Early View Original article Transcriptome profiling reveals the complexity of pirfenidone effects in IPF Grazyna Kwapiszewska, Anna Gungl, Jochen Wilhelm, Leigh M. Marsh, Helene Thekkekara Puthenparampil, Katharina Sinn, Miroslava Didiasova, Walter Klepetko, Djuro Kosanovic, Ralph T. Schermuly, Lukasz Wujak, Benjamin Weiss, Liliana Schaefer, Marc Schneider, Michael Kreuter, Andrea Olschewski, Werner Seeger, Horst Olschewski, Malgorzata Wygrecka Please cite this article as: Kwapiszewska G, Gungl A, Wilhelm J, et al. Transcriptome profiling reveals the complexity of pirfenidone effects in IPF. Eur Respir J 2018; in press (https://doi.org/10.1183/13993003.00564-2018). This manuscript has recently been accepted for publication in the European Respiratory Journal. It is published here in its accepted form prior to copyediting and typesetting by our production team. After these production processes are complete and the authors have approved the resulting proofs, the article will move to the latest issue of the ERJ online. Copyright ©ERS 2018 Copyright 2018 by the European Respiratory Society. Transcriptome profiling reveals the complexity of pirfenidone effects in IPF Grazyna Kwapiszewska1,2, Anna Gungl2, Jochen Wilhelm3†, Leigh M. Marsh1, Helene Thekkekara Puthenparampil1, Katharina Sinn4, Miroslava Didiasova5, Walter Klepetko4, Djuro Kosanovic3, Ralph T. Schermuly3†, Lukasz Wujak5, Benjamin Weiss6, Liliana Schaefer7, Marc Schneider8†, Michael Kreuter8†, Andrea Olschewski1, -

Genome-Wide Pharmacogenomic Analysis of the Response to Interferon Beta Therapy in Multiple Sclerosis
ORIGINAL CONTRIBUTION Genome-Wide Pharmacogenomic Analysis of the Response to Interferon Beta Therapy in Multiple Sclerosis Esther Byun, MD; Stacy J. Caillier, BSc; Xavier Montalban, MD; Pablo Villoslada, MD, PhD; Oscar Ferna´ndez, MD; David Brassat, MD; Manuel Comabella, MD, PhD; Joanne Wang, MPH; Lisa F. Barcellos, PhD; Sergio E. Baranzini, PhD; Jorge R. Oksenberg, PhD Objective: To identify promising candidate genes linked Results: A multianalytical approach detected signifi- to interindividual differences in the efficacy of inter- cant associations between several SNPs and treatment re- feron beta therapy. Recombinant interferon beta therapy sponse, which were validated by individual DNA geno- is widely used to reduce disease activity in multiple scle- typing on an independent platform. After the validation rosis (MS). However, up to 50% of patients continue to stage was complete, 81 additional individuals were added have relapses and worsening disability despite therapy. to the analysis to increase power. We found that respond- ers and nonresponders had significantly different geno- Design: We used a genome-wide pharmacogenomic ap- type frequencies for SNPs located in many genes, includ- proach to identify single-nucleotide polymorphism (SNP) ing glypican 5, collagen type XXV α1, hyaluronan allelic differences associated with interferon beta therapy response. proteoglycan link protein, calpastatin, and neuronal PAS domain protein 3. Setting: Four collaborating centers in the Mediterra- nean Basin. Data Coordination Center at the University Conclusions: The reported results address the ques- of California, San Francisco. tion of genetic heterogeneity in MS and the response to immunotherapy by analysis of the correlation between Patients: A cohort of 206 patients with relapsing- different genotypes and clinical response to interferon remitting MS followed up prospectively for 2 years after beta therapy.