OMERO Documentation Release 5.2.5
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Sun Glassfish Enterprise Server V3 Scripting Framework Guide
Sun GlassFish Enterprise Server v3 Scripting Framework Guide Sun Microsystems, Inc. 4150 Network Circle Santa Clara, CA 95054 U.S.A. Part No: 820–7697–11 December 2009 Copyright 2009 Sun Microsystems, Inc. 4150 Network Circle, Santa Clara, CA 95054 U.S.A. All rights reserved. Sun Microsystems, Inc. has intellectual property rights relating to technology embodied in the product that is described in this document. In particular, and without limitation, these intellectual property rights may include one or more U.S. patents or pending patent applications in the U.S. and in other countries. U.S. Government Rights – Commercial software. Government users are subject to the Sun Microsystems, Inc. standard license agreement and applicable provisions of the FAR and its supplements. This distribution may include materials developed by third parties. Parts of the product may be derived from Berkeley BSD systems, licensed from the University of California. UNIX is a registered trademark in the U.S. and other countries, exclusively licensed through X/Open Company, Ltd. Sun, Sun Microsystems, the Sun logo, the Solaris logo, the Java Coffee Cup logo, docs.sun.com, Enterprise JavaBeans, EJB, GlassFish, J2EE, J2SE, Java Naming and Directory Interface, JavaBeans, Javadoc, JDBC, JDK, JavaScript, JavaServer, JavaServer Pages, JMX, JRE, JSP,JVM, MySQL, NetBeans, OpenSolaris, SunSolve, Sun GlassFish, ZFS, Java, and Solaris are trademarks or registered trademarks of Sun Microsystems, Inc. or its subsidiaries in the U.S. and other countries. All SPARC trademarks are used under license and are trademarks or registered trademarks of SPARC International, Inc. in the U.S. and other countries. -

(Cont'd) Current Trends
Scripting vs Systems Programming Languages (cont’d) Designed for gluing Designed for building Does application implement complex algorithms and data applications : flexibility applications : efficiency structures? Does application process large data sets (>10,000 items)? Interpreted Compiled Are application functions well-defined, fixed? Dynamic variable creation Variable declaration If yes, consider a system programming language. Data and code integrated : Data and code separated : meta-programming cannot create/run code on Is the main task to connect components, legacy apps? supported the fly Does the application manipulate a variety of things? Does the application have a GUI? Dynamic typing (typeless) Static typing Are the application's functions evolving rapidly? Examples: PERL, Tcl, Examples: PL/1, Ada, Must the application be extensible? Python, Ruby, Scheme, Java, C/C++, C#, etc Does the application do a lot of string manipulation? Visual Basic, etc If yes, consider a scripting language. cs480 (Prasad) LSysVsScipt 1 cs480 (Prasad) LSysVsScipt 2 Current Trends Jython (for convenient access to Java APIs) Hybrid Languages : Scripting + Systems Programming I:\tkprasad\cs480>jython – Recent JVM-based Scripting Languages Jython 2.1 on java1.4.1_02 (JIT: null) Type "copyright", "credits" or "license" for more information. »Jython : Python dialect >>> import javax.swing as swing >>> win = swing.JFrame("Welcome to Jython") »Clojure : LISP dialect >>> win.size = (200, 200) »Scala : OOP +Functional Hybrid >>> win.show() -

Amazon Codeguru Profiler
Amazon CodeGuru Profiler User Guide Amazon CodeGuru Profiler User Guide Amazon CodeGuru Profiler: User Guide Copyright © Amazon Web Services, Inc. and/or its affiliates. All rights reserved. Amazon's trademarks and trade dress may not be used in connection with any product or service that is not Amazon's, in any manner that is likely to cause confusion among customers, or in any manner that disparages or discredits Amazon. All other trademarks not owned by Amazon are the property of their respective owners, who may or may not be affiliated with, connected to, or sponsored by Amazon. Amazon CodeGuru Profiler User Guide Table of Contents What is Amazon CodeGuru Profiler? ..................................................................................................... 1 What can I do with CodeGuru Profiler? ......................................................................................... 1 What languages are supported by CodeGuru Profiler? ..................................................................... 1 How do I get started with CodeGuru Profiler? ................................................................................ 1 Setting up ......................................................................................................................................... 3 Set up in the Lambda console ..................................................................................................... 3 Step 1: Sign up for AWS .................................................................................................... -

Characteristics of Dynamic JVM Languages
Characteristics of Dynamic JVM Languages Aibek Sarimbekov Andrej Podzimek Lubomir Bulej University of Lugano Charles University in Prague University of Lugano fi[email protected] [email protected]ff.cuni.cz fi[email protected] Yudi Zheng Nathan Ricci Walter Binder University of Lugano Tufts University University of Lugano fi[email protected] [email protected] fi[email protected] Abstract However, since the JVM was originally conceived for The Java Virtual Machine (JVM) has become an execution a statically-typed language, the performance of the JVM platform targeted by many programming languages. How- and its JIT compiler with dynamically-typed languages is ever, unlike with Java, a statically-typed language, the per- often lacking, lagging behind purpose-built language-specific formance of the JVM and its Just-In-Time (JIT) compiler JIT compilers. Making the JVM perform well with various with dynamically-typed languages lags behind purpose-built statically- and dynamically-typed languages clearly requires language-specific JIT compilers. In this paper, we aim to significant effort, not only in optimizing the JVM itself, but contribute to the understanding of the workloads imposed on also, more importantly, in optimizing the bytecode-emitting the JVM by dynamic languages. We use various metrics to language compiler, instead of just relying on the original JIT characterize the dynamic behavior of a variety of programs to gain performance [8]. This in turn requires that developers written in three dynamic languages (Clojure, Python, and of both the language compilers and the JVM understand the Ruby) executing on the JVM. We identify the differences characteristics of the JVM workloads produced by various with respect to Java, and briefly discuss their implications. -

Debian Java Insights and Challenges
Debian Java Insights and challenges Markus Koschany FOSDEM 19 Brussels / Belgium February, 3rd 2019 Markus Koschany Debian Java: Insights and challenges FOSDEM 19 1/7 The importance of Java Source / binary packages maintained by the Java team: 1033 / 1644 (+10,84 % since Debian 9) Source lines of code (Rank 3) : 90,744,884 Popcon value OpenJDK-8 (installed): 78104 / 199604 Popular libraries: apache-commons-*, javamail, xerces2, bouncycastle Popular applications: libreoffice, netbeans, pdfsam, sweethome3d, freeplane, freecol Frequently used for scientific research, medical care and bioinformatics. Markus Koschany Debian Java: Insights and challenges FOSDEM 19 2/7 What is new in Buster? OpenJDK 11 transition completed. (required more than 400! package updates) Build tools: Ant and Maven are up-to-date. Gradle is stuck at the last pre-Kotlin version. SBT is still being worked on. JVM languages: Groovy 2.14, Scala 2.11.12 (2.12 requires SBT), Clojure 1.9, Jython 1.7.1, JRuby 9.1.13 (?), Kotlin is wanted but hard to bootstrap. IDE: Eclipse is gone (lack of maintainers) but there is Netbeans 10 now. Server: Jetty 9.4 and Tomcat 9 fully up-to-date with systemd integration. Reproducibility rate is at 85% (was 75%) https://reproducible-builds.org Markus Koschany Debian Java: Insights and challenges FOSDEM 19 3/7 Packaging challenges “None of the packages in the main archive area require software outside of that area to function” Internet downloads at build time are not allowed No prebuilt jar or class files! Java is version-centric. Every developer has to update every dependency themself in this model. -

ERDA User Guide
User Guide 22. July 2021 1 / 116 Table of Contents Introduction..........................................................................................................................................3 Requirements and Terms of Use...........................................................................................................3 How to Access UCPH ERDA...............................................................................................................3 Sign-up.............................................................................................................................................4 Login................................................................................................................................................7 Overview..........................................................................................................................................7 Home................................................................................................................................................8 Files..................................................................................................................................................9 File Sharing and Data Exchange....................................................................................................15 Share Links...............................................................................................................................15 Workgroup Shared Folders.......................................................................................................19 -

Reading the Runes for Java Runtimes the Latest IBM Java Sdks
Java Technology Centre Reading the runes for Java runtimes The latest IBM Java SDKs ... and beyond Tim Ellison [email protected] © 2009 IBM Corporation Java Technology Centre Goals . IBM and Java . Explore the changing landscape of hardware and software influences . Discuss the impact to Java runtime technology due to these changes . Show how IBM is leading the way with these changes 2 Mar 9, 2009 © 2009 IBM Corporation Java Technology Centre IBM and Java . Java is critically important to IBM – Provides fundamental infrastructure to IBM software portfolio – Delivers standard development environment – Enables cost effective multi platform support – Delivered to Independent Software Vendors supporting IBM server platforms . IBM is investing strategically in virtual machine technology – Since Java 5.0, a single Java platform technology supports ME, SE and EE – Technology base on which to delivery improved performance, reliability and serviceability • Some IBM owned code (Virtual machine, JIT compiler, ...) • Some open source code (Apache XML parser, Apache Core libraries, Zlib, ...) • Some Sun licensed code (class libraries, tools, ...) . Looking to engender accelerated and open innovation in runtime technologies – Support for Eclipse, Apache (Harmony, XML, Derby, Geronimo, Tuscany) – Broad participation of relevant standards bodies such as JCP and OSGi 3 Mar 9, 2009 © 2009 IBM Corporation Java Technology Centre IBM Java – 2009 key initiatives . Consumability – Deliver value without complexity. – Ensure that problems with our products can be addressed quickly, allowing customers to keep focus on their own business issues. – Deliver a consistent model for solving customer problems. “Scaling Up” - Emerging hardware and applications – Provide a Java implementation that can scale to the most demanding application needs. -

Glassfish Server 3.0.1 Scripting Framework Guide
Oracle® GlassFish Server 3.0.1 Scripting Framework Guide Part No: 821–1760–10 June 2010 Copyright © 2010, Oracle and/or its affiliates. All rights reserved. This software and related documentation are provided under a license agreement containing restrictions on use and disclosure and are protected by intellectual property laws. Except as expressly permitted in your license agreement or allowed by law, you may not use, copy, reproduce, translate, broadcast, modify, license, transmit, distribute, exhibit, perform, publish, or display any part, in any form, or by any means. Reverse engineering, disassembly, or decompilation of this software, unless required by law for interoperability, is prohibited. The information contained herein is subject to change without notice and is not warranted to be error-free. If you find any errors, please report them to us in writing. If this is software or related software documentation that is delivered to the U.S. Government or anyone licensing it on behalf of the U.S. Government, the following notice is applicable: U.S. GOVERNMENT RIGHTS Programs, software, databases, and related documentation and technical data delivered to U.S. Government customers are “commercial computer software” or “commercial technical data” pursuant to the applicable Federal Acquisition Regulation and agency-specific supplemental regulations. As such, the use, duplication, disclosure, modification, and adaptation shall be subject to the restrictions and license terms setforth in the applicable Government contract, and, to the extent applicable by the terms of the Government contract, the additional rights set forth in FAR 52.227-19, Commercial Computer Software License (December 2007). Oracle America, Inc., 500 Oracle Parkway, Redwood City, CA 94065. -
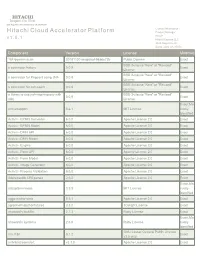
Hitachi Cloud Accelerator Platform Product Manager HCAP V 1
HITACHI Inspire the Next 2535 Augustine Drive Santa Clara, CA 95054 USA Contact Information : Hitachi Cloud Accelerator Platform Product Manager HCAP v 1 . 5 . 1 Hitachi Vantara LLC 2535 Augustine Dr. Santa Clara CA 95054 Component Version License Modified 18F/domain-scan 20181130-snapshot-988de72b Public Domain Exact BSD 3-clause "New" or "Revised" a connector factory 0.0.9 Exact License BSD 3-clause "New" or "Revised" a connector for Pageant using JNA 0.0.9 Exact License BSD 3-clause "New" or "Revised" a connector for ssh-agent 0.0.9 Exact License a library to use jsch-agent-proxy with BSD 3-clause "New" or "Revised" 0.0.9 Exact sshj License Exact,Ma activesupport 5.2.1 MIT License nually Identified Activiti - BPMN Converter 6.0.0 Apache License 2.0 Exact Activiti - BPMN Model 6.0.0 Apache License 2.0 Exact Activiti - DMN API 6.0.0 Apache License 2.0 Exact Activiti - DMN Model 6.0.0 Apache License 2.0 Exact Activiti - Engine 6.0.0 Apache License 2.0 Exact Activiti - Form API 6.0.0 Apache License 2.0 Exact Activiti - Form Model 6.0.0 Apache License 2.0 Exact Activiti - Image Generator 6.0.0 Apache License 2.0 Exact Activiti - Process Validation 6.0.0 Apache License 2.0 Exact Addressable URI parser 2.5.2 Apache License 2.0 Exact Exact,Ma adzap/timeliness 0.3.8 MIT License nually Identified aggs-matrix-stats 5.5.1 Apache License 2.0 Exact agronholm/pythonfutures 3.3.0 3Delight License Exact ahoward's lockfile 2.1.3 Ruby License Exact Exact,Ma ahoward's systemu 2.6.5 Ruby License nually Identified GNU Lesser General Public License ai's -

Magicdraw Macro Engine User Guide
MACRO ENGINE USER GUIDE version 17.0 No Magic, Inc. 2011 All material contained herein is considered proprietary information owned by No Magic, Inc. and is not to be shared, copied, or reproduced by any means. All information copyright 2009-2011 by No Magic, Inc. CONTENTS MACRO ENGINE 2 1. Introduction 2 2. Working with Macro Engine 2 2.1 Selecting a Default Macro Language 2 2.2 Creating a Macro 4 2.3 Adding a Macro, Entering and Editing Macro Information 5 2.3.1 Opening Macro Information Dialog 5 2.3.2 Adding a Macro and Its Information 7 2.3.3 Editing Macro Information 11 2.3.4 Macro Information Dialog Mnemonic Keys 12 2.4 Deleting and Executing Macros 12 2.4.1 Deleting a Macro 12 2.4.2 Executing a Macro 13 2.4.3 Organize Macros Dialog Mnemonic Keys 15 2.5 Macro Keyboard Shortcuts 17 2.5.1 Assigning a Keyboard Shortcut to a Macro 19 2.5.2 Removing a Keyboard Shortcut from a Macro 19 2.6 Opaque Objects 20 2.6.1 Getting an Opaque Object 20 2.6.2 Getting Element Property Values 21 2.6.3 Setting Element Property Values 22 2.6.4 Getting the Child of an Element 25 2.6.5 Getting the Owner of an Element 26 2.6.6 Creating a New Element 26 2.6.7 Creating a Relationship Between Elements 26 2.6.8 Removing an Element 26 2.6.9 Adding a Stereotype to an Element 27 2.6.10 Removing a Stereotype from an Element 27 2.6.11 Printing Element Details 27 2.7 Recording Macros 28 3. -

Using Other Tools with Jython
A P P E N D I X A Using Other Tools with Jython The primary focus of this appendix is to provide information on using some external Python packages with Jython, as well as providing information regarding the Jython registry. In some circumstances, the tools must be used or installed a bit differently on Jython than on CPython, and those differences will be noted. Because there is a good deal of documentation on the usage of these tools available on the web, this appendix will focus on using the tool specifically with Jython. However, relevant URLs will be cited for finding more documentation on each of the topics. The Jython Registry Because there is no good platform-independent equivalent of the Windows Registry or Unix environment variables, Java has its own environment variable namespace. Jython acquires its namespace from the following sources (later sources override defaults found in earlier places): • The Java system properties, typically passed in on the command line as options to the java interpreter. • The Jython “registry” file, containing prop=value pairs. Read on for the algorithm Jython uses to find the registry file. • The user’s personal registry file, containing similarly formatted prop/value pairs. The user’s registry file can be found at "user.home"+"/.jython" • Jython properties specified on the command line as options to the Jython class. See the -D option to the interpreter. Registry Properties The following properties are recognized by Jython. There may be others that aren’t documented here; consult the comments in registry file for details. python.cachedir The directory to use for caches—currently just package information. -

IT-Empfehlungen Für Den Nachhaltigen Umgang Mit Digitalen Daten in Den Altertumswissenschaften
IT-Empfehlungen für den nachhaltigen Umgang mit digitalen Daten in den Altertumswissenschaften Herausgeber: IANUS Version 1.0.0.0 29. März 2017 Koordination Förderung Herausgeber IANUS Autoren: siehe Autorenverzeichnis Titel: IT-Empfehlungen für den nachhaltigen Umgang mit digitalen Daten in den Altertumswissenschaften Sprache: Deutsch doi:10.13149/000.y47clt-t Zitierhinweis: IANUS (Hrsg.), IT-Empfehlungen für den nachhaltigen Umgang mit digi- talen Daten in den Altertumswissenschaften (2017) [Version 1.0.0.0] doi: http://dx.doi.org/10.13149/000.y47clt-t Kontakt: [email protected] www.ianus-fdz.de Lizenz: cba BYSA Dieses Werk bzw. Inhalt ist lizenziert unter einer Creative Commons Namensnennung – Weitergabe unter gleichen Bedingungen 4.0 International Lizenz. http://creativecommons.org/licenses/by-sa/4.0/ Versionshistorie Version Autor Datum Beschreibung 0.9 IANUS 18.02.2014 Erste veröffentlichte Fassung. 0.9 IANUS 11.03.2014 Änderung der Lizenzbedingungen. 0.95 IANUS 05.02.2015 Kapitel „3D und Virtual Reality“ und „Textdokumente“ neu. Anpassungen im Layout und kleinere Korrekturen. 0.96 IANUS 27.05.2015 Kapitel „Datenmanagement“ neu, Ergänzung Versionskontrolle und Korrekturen. Fortsetzung auf der nächsten Seite Fortsetzung der vorhergehenden Seite Version Autor Datum Beschreibung 0.97 IANUS 09.12.2015 Kapitel „Tabellen“ neu, Ergänzungen im Glossar und Korrekturen. 0.97.5 IANUS 21.07.2016 Kapitel „Video“, „Audio“ und „RTI“ neu, Ergänzungen im Glossar, Anpassungen im Kapitel „Archivierung bei IANUS“ und Korrekturen. 0.97.5 IANUS 13.09.2016 Einarbeitung Reviews für „Video“, „Audio“ und „RTI“, Ergänzungen im Glossar und Korrekturen. 0.98 IANUS 22.11.2016 Überarbeitung „Datenmanagement“ und „3D und Virtual Reality“, Ergänzung Bild in „RTI“.