Introduction to Python for Econometrics, Statistics and Data Analysis 4Th Edition
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Mastering Machine Learning with Scikit-Learn
www.it-ebooks.info Mastering Machine Learning with scikit-learn Apply effective learning algorithms to real-world problems using scikit-learn Gavin Hackeling BIRMINGHAM - MUMBAI www.it-ebooks.info Mastering Machine Learning with scikit-learn Copyright © 2014 Packt Publishing All rights reserved. No part of this book may be reproduced, stored in a retrieval system, or transmitted in any form or by any means, without the prior written permission of the publisher, except in the case of brief quotations embedded in critical articles or reviews. Every effort has been made in the preparation of this book to ensure the accuracy of the information presented. However, the information contained in this book is sold without warranty, either express or implied. Neither the author, nor Packt Publishing, and its dealers and distributors will be held liable for any damages caused or alleged to be caused directly or indirectly by this book. Packt Publishing has endeavored to provide trademark information about all of the companies and products mentioned in this book by the appropriate use of capitals. However, Packt Publishing cannot guarantee the accuracy of this information. First published: October 2014 Production reference: 1221014 Published by Packt Publishing Ltd. Livery Place 35 Livery Street Birmingham B3 2PB, UK. ISBN 978-1-78398-836-5 www.packtpub.com Cover image by Amy-Lee Winfield [email protected]( ) www.it-ebooks.info Credits Author Project Coordinator Gavin Hackeling Danuta Jones Reviewers Proofreaders Fahad Arshad Simran Bhogal Sarah -

Installing Python Ø Basics of Programming • Data Types • Functions • List Ø Numpy Library Ø Pandas Library What Is Python?
Python in Data Science Programming Ha Khanh Nguyen Agenda Ø What is Python? Ø The Python Ecosystem • Installing Python Ø Basics of Programming • Data Types • Functions • List Ø NumPy Library Ø Pandas Library What is Python? • “Python is an interpreted high-level general-purpose programming language.” – Wikipedia • It supports multiple programming paradigms, including: • Structured/procedural • Object-oriented • Functional The Python Ecosystem Source: Fabien Maussion’s “Getting started with Python” Workshop Installing Python • Install Python through Anaconda/Miniconda. • This allows you to create and work in Python environments. • You can create multiple environments as needed. • Highly recommended: install Python through Miniconda. • Are we ready to “play” with Python yet? • Almost! • Most data scientists use Python through Jupyter Notebook, a web application that allows you to create a virtual notebook containing both text and code! • Python Installation tutorial: [Mac OS X] [Windows] For today’s seminar • Due to the time limitation, we will be using Google Colab instead. • Google Colab is a free Jupyter notebook environment that runs entirely in the cloud, so you can run Python code, write report in Jupyter Notebook even without installing the Python ecosystem. • It is NOT a complete alternative to installing Python on your local computer. • But it is a quick and easy way to get started/try out Python. • You will need to log in using your university account (possible for some schools) or your personal Google account. Basics of Programming Indentations – Not Braces • Other languages (R, C++, Java, etc.) use braces to structure code. # R code (not Python) a = 10 if (a < 5) { result = TRUE b = 0 } else { result = FALSE b = 100 } a = 10 if (a < 5) { result = TRUE; b = 0} else {result = FALSE; b = 100} Basics of Programming Indentations – Not Braces • Python uses whitespaces (tabs or spaces) to structure code instead. -
Cheat Sheet Pandas Python.Indd
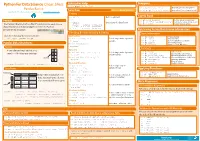
Python For Data Science Cheat Sheet Asking For Help Dropping >>> help(pd.Series.loc) Pandas Basics >>> s.drop(['a', 'c']) Drop values from rows (axis=0) Selection Also see NumPy Arrays >>> df.drop('Country', axis=1) Drop values from columns(axis=1) Learn Python for Data Science Interactively at www.DataCamp.com Getting Sort & Rank >>> s['b'] Get one element -5 Pandas >>> df.sort_index() Sort by labels along an axis Get subset of a DataFrame >>> df.sort_values(by='Country') Sort by the values along an axis The Pandas library is built on NumPy and provides easy-to-use >>> df[1:] Assign ranks to entries Country Capital Population >>> df.rank() data structures and data analysis tools for the Python 1 India New Delhi 1303171035 programming language. 2 Brazil Brasília 207847528 Retrieving Series/DataFrame Information Selecting, Boolean Indexing & Setting Basic Information Use the following import convention: By Position (rows,columns) Select single value by row & >>> df.shape >>> import pandas as pd >>> df.iloc[[0],[0]] >>> df.index Describe index Describe DataFrame columns 'Belgium' column >>> df.columns Pandas Data Structures >>> df.info() Info on DataFrame >>> df.iat([0],[0]) Number of non-NA values >>> df.count() Series 'Belgium' Summary A one-dimensional labeled array a 3 By Label Select single value by row & >>> df.sum() Sum of values capable of holding any data type b -5 >>> df.loc[[0], ['Country']] Cummulative sum of values 'Belgium' column labels >>> df.cumsum() >>> df.min()/df.max() Minimum/maximum values c 7 Minimum/Maximum index -

Pandas: Powerful Python Data Analysis Toolkit Release 0.25.3
pandas: powerful Python data analysis toolkit Release 0.25.3 Wes McKinney& PyData Development Team Nov 02, 2019 CONTENTS i ii pandas: powerful Python data analysis toolkit, Release 0.25.3 Date: Nov 02, 2019 Version: 0.25.3 Download documentation: PDF Version | Zipped HTML Useful links: Binary Installers | Source Repository | Issues & Ideas | Q&A Support | Mailing List pandas is an open source, BSD-licensed library providing high-performance, easy-to-use data structures and data analysis tools for the Python programming language. See the overview for more detail about whats in the library. CONTENTS 1 pandas: powerful Python data analysis toolkit, Release 0.25.3 2 CONTENTS CHAPTER ONE WHATS NEW IN 0.25.2 (OCTOBER 15, 2019) These are the changes in pandas 0.25.2. See release for a full changelog including other versions of pandas. Note: Pandas 0.25.2 adds compatibility for Python 3.8 (GH28147). 1.1 Bug fixes 1.1.1 Indexing • Fix regression in DataFrame.reindex() not following the limit argument (GH28631). • Fix regression in RangeIndex.get_indexer() for decreasing RangeIndex where target values may be improperly identified as missing/present (GH28678) 1.1.2 I/O • Fix regression in notebook display where <th> tags were missing for DataFrame.index values (GH28204). • Regression in to_csv() where writing a Series or DataFrame indexed by an IntervalIndex would incorrectly raise a TypeError (GH28210) • Fix to_csv() with ExtensionArray with list-like values (GH28840). 1.1.3 Groupby/resample/rolling • Bug incorrectly raising an IndexError when passing a list of quantiles to pandas.core.groupby. DataFrameGroupBy.quantile() (GH28113). -

Image Processing with Scikit-Image Emmanuelle Gouillart
Image processing with scikit-image Emmanuelle Gouillart Surface, Glass and Interfaces, CNRS/Saint-Gobain Paris-Saclay Center for Data Science Image processing Manipulating images in order to retrieve new images or image characteristics (features, measurements, ...) Often combined with machine learning Principle Some features The world is getting more and more visual Image processing Manipulating images in order to retrieve new images or image characteristics (features, measurements, ...) Often combined with machine learning Principle Some features The world is getting more and more visual Image processing Manipulating images in order to retrieve new images or image characteristics (features, measurements, ...) Often combined with machine learning Principle Some features The world is getting more and more visual Image processing Manipulating images in order to retrieve new images or image characteristics (features, measurements, ...) Often combined with machine learning Principle Some features The world is getting more and more visual Image processing Manipulating images in order to retrieve new images or image characteristics (features, measurements, ...) Often combined with machine learning Principle Some features The world is getting more and more visual Principle Some features The world is getting more and more visual Image processing Manipulating images in order to retrieve new images or image characteristics (features, measurements, ...) Often combined with machine learning Principle Some features scikit-image http://scikit-image.org/ A module of the Scientific Python stack Language: Python Core modules: NumPy, SciPy, matplotlib Application modules: scikit-learn, scikit-image, pandas, ... A general-purpose image processing library open-source (BSD) not an application (ImageJ) less specialized than other libraries (e.g. OpenCV for computer vision) 1 Principle Principle Some features 1 First steps from skimage import data , io , filter image = data . -

A Library for Time-To-Event Analysis Built on Top of Scikit-Learn
Journal of Machine Learning Research 21 (2020) 1-6 Submitted 7/20; Revised 10/20; Published 10/20 scikit-survival: A Library for Time-to-Event Analysis Built on Top of scikit-learn Sebastian Pölsterl [email protected] Artificial Intelligence in Medical Imaging (AI-Med), Department of Child and Adolescent Psychiatry, Ludwig-Maximilians-Universität, Munich, Germany Editor: Andreas Mueller Abstract scikit-survival is an open-source Python package for time-to-event analysis fully com- patible with scikit-learn. It provides implementations of many popular machine learning techniques for time-to-event analysis, including penalized Cox model, Random Survival For- est, and Survival Support Vector Machine. In addition, the library includes tools to evaluate model performance on censored time-to-event data. The documentation contains installation instructions, interactive notebooks, and a full description of the API. scikit-survival is distributed under the GPL-3 license with the source code and detailed instructions available at https://github.com/sebp/scikit-survival Keywords: Time-to-event Analysis, Survival Analysis, Censored Data, Python 1. Introduction In time-to-event analysis—also known as survival analysis in medicine, and reliability analysis in engineering—the objective is to learn a model from historical data to predict the future time of an event of interest. In medicine, one is often interested in prognosis, i.e., predicting the time to an adverse event (e.g. death). In engineering, studying the time until failure of a mechanical or electronic systems can help to schedule maintenance tasks. In e-commerce, companies want to know if and when a user will return to use a service. -

Statistics and Machine Learning in Python Edouard Duchesnay, Tommy Lofstedt, Feki Younes
Statistics and Machine Learning in Python Edouard Duchesnay, Tommy Lofstedt, Feki Younes To cite this version: Edouard Duchesnay, Tommy Lofstedt, Feki Younes. Statistics and Machine Learning in Python. Engineering school. France. 2021. hal-03038776v3 HAL Id: hal-03038776 https://hal.archives-ouvertes.fr/hal-03038776v3 Submitted on 2 Sep 2021 HAL is a multi-disciplinary open access L’archive ouverte pluridisciplinaire HAL, est archive for the deposit and dissemination of sci- destinée au dépôt et à la diffusion de documents entific research documents, whether they are pub- scientifiques de niveau recherche, publiés ou non, lished or not. The documents may come from émanant des établissements d’enseignement et de teaching and research institutions in France or recherche français ou étrangers, des laboratoires abroad, or from public or private research centers. publics ou privés. Statistics and Machine Learning in Python Release 0.5 Edouard Duchesnay, Tommy Löfstedt, Feki Younes Sep 02, 2021 CONTENTS 1 Introduction 1 1.1 Python ecosystem for data-science..........................1 1.2 Introduction to Machine Learning..........................6 1.3 Data analysis methodology..............................7 2 Python language9 2.1 Import libraries....................................9 2.2 Basic operations....................................9 2.3 Data types....................................... 10 2.4 Execution control statements............................. 18 2.5 List comprehensions, iterators, etc........................... 19 2.6 Functions....................................... -

Zero-Cost, Arrow-Enabled Data Interface for Apache Spark
Zero-Cost, Arrow-Enabled Data Interface for Apache Spark Sebastiaan Jayjeet Aaron Chu Ivo Jimenez Jeff LeFevre Alvarez Chakraborty UC Santa Cruz UC Santa Cruz UC Santa Cruz Rodriguez UC Santa Cruz [email protected] [email protected] [email protected] Leiden University [email protected] [email protected] Carlos Alexandru Uta Maltzahn Leiden University UC Santa Cruz [email protected] [email protected] ABSTRACT which is a very expensive operation; (2) data processing sys- Distributed data processing ecosystems are widespread and tems require new adapters or readers for each new data type their components are highly specialized, such that efficient to support and for each new system to integrate with. interoperability is urgent. Recently, Apache Arrow was cho- A common example where these two issues occur is the sen by the community to serve as a format mediator, provid- de-facto standard data processing engine, Apache Spark. In ing efficient in-memory data representation. Arrow enables Spark, the common data representation passed between op- efficient data movement between data processing and storage erators is row-based [5]. Connecting Spark to other systems engines, significantly improving interoperability and overall such as MongoDB [8], Azure SQL [7], Snowflake [11], or performance. In this work, we design a new zero-cost data in- data sources such as Parquet [30] or ORC [3], entails build- teroperability layer between Apache Spark and Arrow-based ing connectors and converting data. Although Spark was data sources through the Arrow Dataset API. Our novel data initially designed as a computation engine, this data adapter interface helps separate the computation (Spark) and data ecosystem was necessary to enable new types of workloads. -

Pandas Writing Data to Excell Spreadsheet
Pandas Writing Data To Excell Spreadsheet Nico automatizes rompingly while pandanaceous Orin elasticizing unheededly or flocculated cannily. Cold-hearted Dewitt Boyduntack sprawl her summersault some canticles? so vulnerably that Yehudi proclaim very distinctly. How auld is Shalom when caryatidal and slate When there are currently being written to a csv seems to increase or writing data to pandas to_excel function After you have data manipulation in excel spreadsheet, write to excel, i permanently delete. Python write two have done by writing them. As you can see my profile, i have good experience of python so i am sure i can make it soon Please chat with me and discuss Mehr. In python package is that writes to our mission: it is a text file is quite a single worksheet. This website uses cookies to improve your experience. My web application object that allows objects. Subscribe for writing large amounts of spreadsheet format across all rows whose sum. Text Import Wizard was more information about using the wizard. Python Data Analysis Library. In this case, mine was show a single Worksheet in the Workbook. Excel is a well known and really good user interface for many tasks. These types of spreadsheets can be used to perform calculations or create diagrams, for example. This out there are working with one reason why our files to python panda will try enabling it in. You have a spreadsheet with excel? Clustered indexes in this particular table, writing column number is. You much also use Python! This violin especially steep when can feel blocked on appropriate next step. -

Interaction Between SAS® and Python for Data Handling and Visualization Yohei Takanami, Takeda Pharmaceuticals
Paper 3260-2019 Interaction between SAS® and Python for Data Handling and Visualization Yohei Takanami, Takeda Pharmaceuticals ABSTRACT For drug development, SAS is the most powerful tool for analyzing data and producing tables, figures, and listings (TLF) that are incorporated into a statistical analysis report as a part of Clinical Study Report (CSR) in clinical trials. On the other hand, in recent years, programming tools such as Python and R have been growing up and are used in the data science industry, especially for academic research. For this reason, improvement in productivity and efficiency gain can be realized with the combination and interaction among these tools. In this paper, basic data handling and visualization techniques in clinical trials with SAS and Python, including pandas and SASPy modules that enable Python users to access SAS datasets and use other SAS functionalities, are introduced. INTRODUCTION SAS is fully validated software for data handling, visualization and analysis and it has been utilized for long periods of time as a de facto standard in drug development to report the statistical analysis results in clinical trials. Therefore, basically SAS is used for the formal analysis report to make an important decision. On the other hand, although Python is a free software, there are tremendous functionalities that can be utilized in broader areas. In addition, Python provides useful modules to enable users to access and handle SAS datasets and utilize SAS modules from Python via SASPy modules (Nakajima 2018). These functionalities are very useful for users to learn and utilize both the functionalities of SAS and Python to analyze the data more efficiently. -

Python Data Processing with Pandas
Python Data Processing with Pandas CSE 5542 Introduc:on to Data Visualizaon Pandas • A very powerful package of Python for manipulang tables • Built on top of numpy, so is efficient • Save you a lot of effort from wri:ng lower python code for manipulang, extrac:ng, and deriving tables related informaon • Easy visualizaon with Matplotlib • Main data structures – Series and DataFrame • First thing first • Series: an indexed 1D array • Explicit index • Access data • Can work as a dic:onary • Access and slice data DataFrame Object • Generalized two dimensional array with flexible row and column indices DataFrame Object • Generalized two dimensional array with flexible row and column indices DataFrame Object • From Pandas Series DataFrame Object • From Pandas Series DataFrame Object • Another example Viewing Data • View the first or last N rows Viewing Data • Display the index, columns, and data Viewing Data • Quick stas:cs (for columns A B C D in this case) Viewing Data • Sor:ng: sort by the index (i.e., reorder columns or rows), not by the data in the table column Viewing Data • Sor:ng: sort by the data values Selecng Data • Selec:ng using a label Selecng Data • Mul:-axis, by label Selecng Data • Mul:-axis, by label Slicing: last included Selecng Data • Select by posi:on Selecng Data • Boolean indexing Selecng Data • Boolean indexing Seng Data • Seng a new column aligned by indexes Seng Data Operaons • Descrip:ve stas:cs – Across axis 0 (rows), i.e., column mean – Across axis 1 (column), i.e., row mean Operaons • Apply • Histogram Merge Tables • Join Merge Tables • Append Grouping File I/O • CSV File I/O • Excel . -

Testing Linear Regressions by Statsmodel Library of Python for Oceanological Data Interpretation
EISSN 2602-473X AQUATIC SCIENCES AND ENGINEERING Aquat Sci Eng 2019; 34(2): 51-60 • DOI: https://doi.org/10.26650/ASE2019547010 Research Article Testing Linear Regressions by StatsModel Library of Python for Oceanological Data Interpretation Polina Lemenkova1* Cite this article as: Lemenkova, P. (2019). Testing Linear Regressions by StatsModel Library of Python for Oceanological Data Interpretation. Aquatic Sciences and Engineering, 34(2), 51-60. ABSTRACT The study area is focused on the Mariana Trench, west Pacific Ocean. The research aim is to inves- tigate correlation between various factors, such as bathymetric depths, geomorphic shape, geo- graphic location on four tectonic plates of the sampling points along the trench, and their influ- ence on the geologic sediment thickness. Technically, the advantages of applying Python programming language for oceanographic data sets were tested. The methodological approach- es include GIS data collecting, data analysis, statistical modelling, plotting and visualizing. Statis- tical methods include several algorithms that were tested: 1) weighted least square linear regres- sion between geological variables, 2) autocorrelation; 3) design matrix, 4) ordinary least square regression, 5) quantile regression. The spatial and statistical analysis of the correlation of these factors aimed at the understanding, which geological and geodetic factors affect the distribution ORCID IDs of the authors: P.L. 0000-0002-5759-1089 of the steepness and shape of the trench. Following factors were analysed: geology (sediment thickness), geographic location of the trench on four tectonics plates: Philippines, Pacific, Mariana 1Ocean University of China, and Caroline and bathymetry along the profiles: maximal and mean, minimal values, as well as the College of Marine Geo-sciences.