Codon Usage Vis-A-Vis Start and Stop Codon Context Analysis of Three Dicot Species
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Bio 102 Practice Problems Genetic Code and Mutation
Bio 102 Practice Problems Genetic Code and Mutation Multiple choice: Unless otherwise directed, circle the one best answer: 1. Choose the one best answer: Beadle and Tatum mutagenized Neurospora to find strains that required arginine to live. Based on the classification of their mutants, they concluded that: A. one gene corresponds to one protein. B. DNA is the genetic material. C. "inborn errors of metabolism" were responsible for many diseases. D. DNA replication is semi-conservative. E. protein cannot be the genetic material. 2. Choose the one best answer. Which one of the following is NOT part of the definition of a gene? A. A physical unit of heredity B. Encodes a protein C. Segement of a chromosome D. Responsible for an inherited characteristic E. May be linked to other genes 3. A mutation converts an AGA codon to a TGA codon (in DNA). This mutation is a: A. Termination mutation B. Missense mutation C. Frameshift mutation D. Nonsense mutation E. Non-coding mutation 4. Beadle and Tatum performed a series of complex experiments that led to the idea that one gene encodes one enzyme. Which one of the following statements does not describe their experiments? A. They deduced the metabolic pathway for the synthesis of an amino acid. B. Many different auxotrophic mutants of Neurospora were isolated. C. Cells unable to make arginine cannot survive on minimal media. D. Some mutant cells could survive on minimal media if they were provided with citrulline or ornithine. E. Homogentisic acid accumulates and is excreted in the urine of diseased individuals. 5. -

Expanding the Genetic Code Lei Wang and Peter G
Reviews P. G. Schultz and L. Wang Protein Science Expanding the Genetic Code Lei Wang and Peter G. Schultz* Keywords: amino acids · genetic code · protein chemistry Angewandte Chemie 34 2005 Wiley-VCH Verlag GmbH & Co. KGaA, Weinheim DOI: 10.1002/anie.200460627 Angew. Chem. Int. Ed. 2005, 44,34–66 Angewandte Protein Science Chemie Although chemists can synthesize virtually any small organic molecule, our From the Contents ability to rationally manipulate the structures of proteins is quite limited, despite their involvement in virtually every life process. For most proteins, 1. Introduction 35 modifications are largely restricted to substitutions among the common 20 2. Chemical Approaches 35 amino acids. Herein we describe recent advances that make it possible to add new building blocks to the genetic codes of both prokaryotic and 3. In Vitro Biosynthetic eukaryotic organisms. Over 30 novel amino acids have been genetically Approaches to Protein encoded in response to unique triplet and quadruplet codons including Mutagenesis 39 fluorescent, photoreactive, and redox-active amino acids, glycosylated 4. In Vivo Protein amino acids, and amino acids with keto, azido, acetylenic, and heavy-atom- Mutagenesis 43 containing side chains. By removing the limitations imposed by the existing 20 amino acid code, it should be possible to generate proteins and perhaps 5. An Expanded Code 46 entire organisms with new or enhanced properties. 6. Outlook 61 1. Introduction The genetic codes of all known organisms specify the same functional roles to amino acid residues in proteins. Selectivity 20 amino acid building blocks. These building blocks contain a depends on the number and reactivity (dependent on both limited number of functional groups including carboxylic steric and electronic factors) of a particular amino acid side acids and amides, a thiol and thiol ether, alcohols, basic chain. -

Translational Control Using an Expanded Genetic Code
Review Translational Control using an Expanded Genetic Code Yusuke Kato Division of Biotechnology, Institute of Agrobiological Sciences, National Agriculture and Food Research Organization (NARO), Oowashi 1-2, Tsukuba, Ibaraki 305-8634, Japan; [email protected]; Tel.: +81-29-838-6059 Received: 19 December 2018; Accepted: 15 February 2019; Published: 18 February 2019 Abstract: A bio-orthogonal and unnatural substance, such as an unnatural amino acid (Uaa), is an ideal regulator to control target gene expression in a synthetic gene circuit. Genetic code expansion technology has achieved Uaa incorporation into ribosomal synthesized proteins in vivo at specific sites designated by UAG stop codons. This site-specific Uaa incorporation can be used as a controller of target gene expression at the translational level by conditional read-through of internal UAG stop codons. Recent advances in optimization of site-specific Uaa incorporation for translational regulation have enabled more precise control over a wide range of novel important applications, such as Uaa-auxotrophy-based biological containment, live-attenuated vaccine, and high-yield zero-leakage expression systems, in which Uaa translational control is exclusively used as an essential genetic element. This review summarizes the history and recent advance of the translational control by conditional stop codon read-through, especially focusing on the methods using the site-specific Uaa incorporation. Keywords: genetic switch; synthetic biology; unnatural amino acids; translational regulation; biological containment; stop codon read-through 1. Introduction Conditional induction of gene expression is one of the most important technologies for constructing synthetic gene circuits, and such regulation has applications from basic research to industrial use. -

Reducing the Genetic Code Induces Massive Rearrangement of the Proteome
Reducing the genetic code induces massive rearrangement of the proteome Patrick O’Donoghuea,b, Laure Pratc, Martin Kucklickd, Johannes G. Schäferc, Katharina Riedele, Jesse Rinehartf,g, Dieter Söllc,h,1, and Ilka U. Heinemanna,1 Departments of aBiochemistry and bChemistry, The University of Western Ontario, London, ON N6A 5C1, Canada; Departments of cMolecular Biophysics and Biochemistry, fCellular and Molecular Physiology, and hChemistry, and gSystems Biology Institute, Yale University, New Haven, CT 06520; dDepartment of Microbiology, Technical University of Braunschweig, Braunschweig 38106, Germany; and eDivision of Microbial Physiology and Molecular Biology, University of Greifswald, Greifswald 17487, Germany Contributed by Dieter Söll, October 22, 2014 (sent for review September 29, 2014; reviewed by John A. Leigh) Expanding the genetic code is an important aim of synthetic Opening codons by reducing the genetic code is highly biology, but some organisms developed naturally expanded ge- promising, but it is unknown how removing 1 amino acid from netic codes long ago over the course of evolution. Less than 1% of the genetic code might impact the proteome or cellular viability. all sequenced genomes encode an operon that reassigns the stop Many genetic code variations are found in nature (15), including codon UAG to pyrrolysine (Pyl), a genetic code variant that results stop or sense codon reassignments, codon recoding, and natural from the biosynthesis of Pyl-tRNAPyl. To understand the selective code expansion (16). Pyrrolysine (Pyl) is a rare example of nat- advantage of genetically encoding more than 20 amino acids, we ural genetic code expansion. Evidence for genetically encoded constructed a markerless tRNAPyl deletion strain of Methanosarcina Pyl is found in <1% of all sequenced genomes (17). -

How Genes Work
Help Me Understand Genetics How Genes Work Reprinted from MedlinePlus Genetics U.S. National Library of Medicine National Institutes of Health Department of Health & Human Services Table of Contents 1 What are proteins and what do they do? 1 2 How do genes direct the production of proteins? 5 3 Can genes be turned on and off in cells? 7 4 What is epigenetics? 8 5 How do cells divide? 10 6 How do genes control the growth and division of cells? 12 7 How do geneticists indicate the location of a gene? 16 Reprinted from MedlinePlus Genetics (https://medlineplus.gov/genetics/) i How Genes Work 1 What are proteins and what do they do? Proteins are large, complex molecules that play many critical roles in the body. They do most of the work in cells and are required for the structure, function, and regulation of thebody’s tissues and organs. Proteins are made up of hundreds or thousands of smaller units called amino acids, which are attached to one another in long chains. There are 20 different types of amino acids that can be combined to make a protein. The sequence of amino acids determineseach protein’s unique 3-dimensional structure and its specific function. Aminoacids are coded by combinations of three DNA building blocks (nucleotides), determined by the sequence of genes. Proteins can be described according to their large range of functions in the body, listed inalphabetical order: Antibody. Antibodies bind to specific foreign particles, such as viruses and bacteria, to help protect the body. Example: Immunoglobulin G (IgG) (Figure 1) Enzyme. -

Site-Specific Incorporation of Unnatural Amino Acids Into Escherichia Coli Recombinant Protein: Methodology Development and Recent Achievement
Review Site-Specific Incorporation of Unnatural Amino Acids into Escherichia coli Recombinant Protein: Methodology Development and Recent Achievement Sviatlana Smolskaya 1,* and Yaroslav A. Andreev 1,2 1 Sechenov First Moscow State Medical University, Institute of Molecular Medicine, Trubetskaya str. 8, bld. 2, 119991 Moscow, Russia 2 Shemyakin-Ovchinnikov Institute of Bioorganic Chemistry, Russian Academy of Sciences, ul. Miklukho- Maklaya 16/10, 117997 Moscow, Russia; [email protected] * Correspondence: [email protected]; Tel.: +7-903-215-44-89 Received: 30 May 2019; Accepted: 25 June 2019; Published: 28 June 2019 Abstract: More than two decades ago a general method to genetically encode noncanonical or unnatural amino acids (NAAs) with diverse physical, chemical, or biological properties in bacteria, yeast, animals and mammalian cells was developed. More than 200 NAAs have been incorporated into recombinant proteins by means of non-endogenous aminoacyl-tRNA synthetase (aa-RS)/tRNA pair, an orthogonal pair, that directs site-specific incorporation of NAA encoded by a unique codon. The most established method to genetically encode NAAs in Escherichia coli is based on the usage of the desired mutant of Methanocaldococcus janaschii tyrosyl-tRNA synthetase (MjTyrRS) and cognate suppressor tRNA. The amber codon, the least-used stop codon in E. coli, assigns NAA. Until very recently the genetic code expansion technology suffered from a low yield of targeted proteins due to both incompatibilities of orthogonal pair with host cell translational machinery and the competition of suppressor tRNA with release factor (RF) for binding to nonsense codons. Here we describe the latest progress made to enhance nonsense suppression in E. -

Basic Molecular Genetics for Epidemiologists F Calafell, N Malats
398 GLOSSARY Basic molecular genetics for epidemiologists F Calafell, N Malats ............................................................................................................................. J Epidemiol Community Health 2003;57:398–400 This is the first of a series of three glossaries on CHROMOSOME molecular genetics. This article focuses on basic Linear or (in bacteria and organelles) circular DNA molecule that constitutes the basic physical molecular terms. block of heredity. Chromosomes in diploid organ- .......................................................................... isms such as humans come in pairs; each member of a pair is inherited from one of the parents. general increase in the number of epide- Humans carry 23 pairs of chromosomes (22 pairs miological research articles that apply basic of autosomes and two sex chromosomes); chromo- science methods in their studies, resulting somes are distinguished by their length (from 48 A to 257 million base pairs) and by their banding in what is known as both molecular and genetic epidemiology, is evident. Actually, genetics has pattern when stained with appropriate methods. come into the epidemiological scene with plenty Homologous chromosome of new sophisticated concepts and methodologi- cal issues. Each of the chromosomes in a pair with respect to This fact led the editors of the journal to offer the other. Homologous chromosomes carry the you a glossary of terms commonly used in papers same set of genes, and recombine with each other applying genetic methods to health problems to during meiosis. facilitate your “walking” around the journal Sex chromosome issues and enjoying the articles while learning. Sex determining chromosome. In humans, as in Obviously, the topics are so extensive and inno- all other mammals, embryos carrying XX sex vative that a single short glossary would not be chromosomes develop as females, whereas XY sufficient to provide you with the minimum embryos develop as males. -
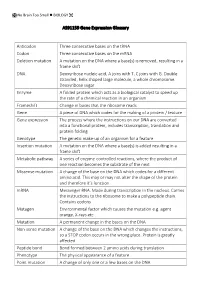
AS91159 Gene Expression Glossary Anticodon Three Consecutive Bases
No Brain Too Small BIOLOGY AS91159 Gene Expression Glossary Anticodon Three consecutive bases on the tRNA Codon Three consecutive bases on the mRNA Deletion mutation A mutation on the DNA where a base(s) is removed, resulting in a frame shift DNA Deoxyribose nucleic acid, A joins with T, C joins with G. Double stranded, helix shaped large molecule, a whole chromosome. Deoxyribose sugar Enzyme A folded protein which acts as a biological catalyst to speed up the rate of a chemical reaction in an organism Frameshift Change in bases that the ribosome reads Gene A piece of DNA which codes for the making of a protein / feature Gene expression The process where the instructions on our DNA are converted into a functional protein, includes transcription, translation and protein folding Genotype The genetic make-up of an organism for a feature Insertion mutation A mutation on the DNA where a base(s) is added resulting in a frame shift Metabolic pathway A series of enzyme-controlled reactions, where the product of one reaction becomes the substrate of the next Missense mutation A change of the base on the DNA which codes for a different amino acid. This may or may not alter the shape of the protein and therefore it’s function mRNA Messenger RNA. Made during transcription in the nucleus. Carries the instructions to the ribosome to make a polypeptide chain. Contains codons Mutagen Environmental factor which causes the mutation e.g. agent orange, X-rays etc Mutation A permanent change in the bases on the DNA Non-sense mutation A change of the base on the DNA which changes the instructions, so a STOP codon occurs in the wrong place. -

Analysis of Stop Codons Within Prokaryotic Protein-Coding Genes Suggests Frequent Readthrough Events
International Journal of Molecular Sciences Article Analysis of Stop Codons within Prokaryotic Protein-Coding Genes Suggests Frequent Readthrough Events Frida Belinky 1 , Ishan Ganguly 1, Eugenia Poliakov 2, Vyacheslav Yurchenko 3,4,* and Igor B. Rogozin 1,* 1 National Center for Biotechnology Information, National Library of Medicine, National Institutes of Health, Bethesda, MD 20894, USA; [email protected] (F.B.); [email protected] (I.G.) 2 National Eye Institute, National Institutes of Health, Bethesda, MD 20892, USA; [email protected] 3 Life Science Research Centre, Faculty of Science, University of Ostrava, 710 00 Ostrava, Czech Republic 4 Martsinovsky Institute of Medical Parasitology, Tropical and Vector Borne Diseases, Sechenov University, 119435 Moscow, Russia * Correspondence: [email protected] (V.Y.); [email protected] (I.B.R.) Abstract: Nonsense mutations turn a coding (sense) codon into an in-frame stop codon that is assumed to result in a truncated protein product. Thus, nonsense substitutions are the hallmark of pseudogenes and are used to identify them. Here we show that in-frame stop codons within bacterial protein-coding genes are widespread. Their evolutionary conservation suggests that many of them are not pseudogenes, since they maintain dN/dS values (ratios of substitution rates at non-synonymous and synonymous sites) significantly lower than 1 (this is a signature of purifying selection in protein- coding regions). We also found that double substitutions in codons—where an intermediate step is a nonsense substitution—show a higher rate of evolution compared to null models, indicating that a stop codon was introduced and then changed back to sense via positive selection. -

LETTER Doi:10.1038/Nature14896
LETTER doi:10.1038/nature14896 Structural basis for stop codon recognition in eukaryotes Alan Brown1*, Sichen Shao1*, Jason Murray1, Ramanujan S. Hegde1 & V. Ramakrishnan1 Termination of protein synthesis occurs when a translating that leads to its dissociation, permitting eRF1 to accommodate fully in ribosome encounters one of three universally conserved stop the A-site. This change is thought to bring a universally conserved codons: UAA, UAG or UGA. Release factors recognize stop codons GGQ motif close to the ester bond between the nascent polypeptide in the ribosomal A-site to mediate release of the nascent chain and and the tRNA, stimulating its hydrolysis. Concomitant with these recycling of the ribosome. Bacteria decode stop codons using two events, the ATPase ABCE1 is recruited to the ribosome after eRF3 separate release factors with differing specificities for the second dissociation and, together with eRF1, catalyses splitting of the ribo- and third bases1. By contrast, eukaryotes rely on an evolutionarily somal subunits to recycle post-termination complexes3,4,7,8. unrelated omnipotent release factor (eRF1) to recognize all three We reasoned that a catalytically inactive eRF1 mutant may trap a stop codons2. The molecular basis of eRF1 discrimination for stop pre-hydrolysis termination complex with two key features. First, eRF1 codons over sense codons is not known. Here we present cryo- would be in complex with the stop codon it had recognized. Second, electron microscopy (cryo-EM) structures at 3.5–3.8 A˚ resolution the unreleased nascent polypeptide would provide a unique affinity of mammalian ribosomal complexes containing eRF1 interacting handle to enrich this species for structural analysis. -

CHAPTER 9 the Molecular Genetics of Gene Expression Assigned: 9.1
CHAPTER 9 The Molecular Genetics of Gene Expression Assigned: 9.1, 9.2, 9.3, 9.4, 9.9, 9.13,9.14, 9.16 9-1. Only statement (d) is true. 9-2. The 5’ end of the RNA transcript is transcribed first, so (a) through (c) are all wrong. The 5’ end of the RNA transcript is transcribed from the 3’ end of the DNA template, so (e) is wrong. The additional statement in (e) that the amino end of the polypeptide is synthesized first serves as confirmation of the correctness of statement (d). 9-3.The initiation codon is AUG and the stop codons are UAA, UAG, and AGA. In a random sequence, the probability of an initiation codon is (1/4)3 = 1/64, and the probability of a stop codon is 3(1/4)3 = 3/64. The average distance between stop codons is 64/3 = 21.3 codons. 9-4. The mRNA sequence is 5’-AAAAUGCCCUUAAUCUCAGCGUCCUAC- 3’ and the first AUG is the initiation codon; so the resulting gamino acid sequence is Met—Pro—Leu—Ile—Ser—Ala—Ser—Tyr or in the single-letter abbreviations M—P—L—I—S—A—S—Y. 9-5. Because the pairing of codon and anticodon are antiparallel, it is convenient to write the anticodon as 3’-UAI-5’. The first two codon positions are therefore 5’-AU-3’, and the third is A or U or C (because each can pair with I). The possible codons are 5’-AUA-3’, 5’-AUU-3’, and 5’-AUC-3’, all of which code for isoleucine. -

Suppression of Nonsense Mutations by New Emerging Technologies
International Journal of Molecular Sciences Review Suppression of Nonsense Mutations by New Emerging Technologies Pedro Morais 1, Hironori Adachi 2 and Yi-Tao Yu 2,* 1 ProQR Therapeutics, Zernikedreef 9, 2333 CK Leiden, The Netherlands; [email protected] 2 Department of Biochemistry and Biophysics, Center for RNA Biology, University of Rochester Medical Center, 601 Elmwood Avenue, Rochester, NY 14642, USA; [email protected] * Correspondence: [email protected]; Tel.: +1-(585)-275-1271; Fax: +1-(585)-275-6007 Received: 28 May 2020; Accepted: 18 June 2020; Published: 20 June 2020 Abstract: Nonsense mutations often result from single nucleotide substitutions that change a sense codon (coding for an amino acid) to a nonsense or premature termination codon (PTC) within the coding region of a gene. The impact of nonsense mutations is two-fold: (1) the PTC-containing mRNA is degraded by a surveillance pathway called nonsense-mediated mRNA decay (NMD) and (2) protein translation stops prematurely at the PTC codon, and thus no functional full-length protein is produced. As such, nonsense mutations result in a large number of human diseases. Nonsense suppression is a strategy that aims to correct the defects of hundreds of genetic disorders and reverse disease phenotypes and conditions. While most clinical trials have been performed with small molecules, there is an increasing need for sequence-specific repair approaches that are safer and adaptable to personalized medicine. Here, we discuss recent advances in both conventional strategies as well as new technologies. Several of these will soon be tested in clinical trials as nonsense therapies, even if they still have some limitations and challenges to overcome.