Difference Between Sigma Factors and Transcription Factors
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Characterization of the Osmoregulated Escherichia Coli Prou Promoter and Identification of Prov As a Membrane-Associated Protein
Molecular Microbiology (1989) 3(11), 1521-1531 Characterization of the osmoregulated Escherichia coli proU promoter and identification of ProV as a membrane-associated protein G. May, E. Faatz, J. M. Lucht, M. Haardt, The prop-encoded transport system has a iow affinity for M. Bolliger^ and E. Bremer* giycine betaine and is present in the cytopiasmic mem- Department of Biotogy. University of Konstanz, PO Box brane (Milner et ai. 1988). The protZ-encoded transport 5560, D'775O Konstanz. FRG. system has a high affinity for glycine betaine and is binding-protein-dependent (May ef ai. 1986; Higgins et ai, 1987a; Barron efa/., 1987;foranoverview, see Ames, Summary 1986). Analyses of the cloned proU region from £ coli The Escherichia coli proU operon encodes a high- have demonstrated that this iocus consists of at ieast affinity, binding-protein-dependent transport system three genes (proV. proW, and proX) that are organized in for the osmoprotectant gtycine betaine. Expression of an operon (Gowrishankar et ai, 1986; Faatz et ai, 1988; proU is osmoregulated, and transcription of this Dattananda and Gowrishankar, 1989). From the recently operon is greatly increased In cells grown at high determined proU DNA sequence {Gowrishankar, 1989), osmoiarity. Characterization of the proU operon and the products of these genes have been deduced. The proV its promoter provrded results similar to those pub- gene encodes a hydrophiiic protein {M, ^ 44162) with lished elsewhere (Gowrishankar, 1989; Stirling et a/., homology to the energy-coupling component of binding- 1989). The previously identified proU601 mutation, protein-dependent transport systems. A hydrophobic which leads to increased prot>expression both at low- polypeptide (M, = 37619) is encoded by proW and is and high osmolarity, is a G to A transition in the thought to be located in the cytoplasmic membrane. -

DNA-Binding and Transcriptional Properties of Human Transcription Factor TFIID After Mild Proteolysis MICHAEL W
MOLECULAR AND CELLULAR BIOLOGY, JUlY 1990, p. 3415-3420 Vol. 10, No. 7 0270-7306/90/073415-06$02.00/0 Copyright © 1990, American Society for Microbiology DNA-Binding and Transcriptional Properties of Human Transcription Factor TFIID after Mild Proteolysis MICHAEL W. VAN DYKE'* AND MICHELE SAWADOGO2 Department of Tumor Biology' and Department of Molecular Genetics,2 The University of Texas M. D. Anderson Cancer Center, Houston, Texas 77030 Received 4 January 1990/Accepted 11 April 1990 The existence of separable functions within the human class H general transcription factor TFIID was probed for differential sensitivity to mild proteolytic treatment. Independent of whether TFIID was bound to DNA or free in solution, partial digestion with either one of a variety of nonspecific endoproteases generated a protease- resistant protein product that retained specific DNA recognition, as revealed by DNase I footprinting. However, in contrast to native TFIID, which interacts with the adenovirus major late (ML) promoter over a very broad DNA region, partially proteolyzed TFIID interacted with only a small region of the ML promoter immediately surrounding the TATA sequence. This novel footprint was very similar to that observed with the TATA factor purified from yeast cells. Partially proteolyzed human TFIID could form stable complexes that were resistant to challenge by exogenous templates. It could also nucleate the assembly of transcription complexes on the ML promoter with an efficiency comparable to that of native TFIID, yielding similar levels of transcription initiation. These results suggest a model in which the human TFHID protein is composed of at least two different regions or polypeptides: a protease-resistant "core," which by itself is sufficient for promoter recognition and basal transcriptional levels, and a protease-sensitive "tail," which interacts with downstream promoter regions and may be involved in regulatory processes. -

Genome-Wide Mutational Biases Fuel Transcriptional Diversity in the Mycobacterium Tuberculosis Complex
ARTICLE https://doi.org/10.1038/s41467-019-11948-6 OPEN Genome-wide mutational biases fuel transcriptional diversity in the Mycobacterium tuberculosis complex Álvaro Chiner-Oms 1,2,12, Michael Berney 3,12, Christine Boinett4,5, Fernando González-Candelas 1,6, Douglas B. Young7, Sebastien Gagneux8,9, William R. Jacobs Jr3, Julian Parkhill 10, Teresa Cortes 11 & Iñaki Comas 2,6 1234567890():,; The Mycobacterium tuberculosis complex (MTBC) members display different host-specificities and virulence phenotypes. Here, we have performed a comprehensive RNAseq and methy- lome analysis of the main clades of the MTBC and discovered unique transcriptional profiles. The majority of genes differentially expressed between the clades encode proteins involved in host interaction and metabolic functions. A significant fraction of changes in gene expression can be explained by positive selection on single mutations that either create or disrupt transcriptional start sites (TSS). Furthermore, we show that clinical strains have different methyltransferases inactivated and thus different methylation patterns. Under the tested conditions, differential methylation has a minor direct role on transcriptomic differences between strains. However, disruption of a methyltransferase in one clinical strain revealed important expression differences suggesting indirect mechanisms of expression regulation. Our study demonstrates that variation in transcriptional profiles are mainly due to TSS mutations and have likely evolved due to differences in host characteristics. 1 Unidad Mixta “Infección y Salud Pública” FISABIO-CSISP/Universidad de Valencia, Instituto de Biología Integrativa de Sistemas-I2SysBio, Valencia, Spain. 2 Instituto de Biomedicina de Valencia, IBV-CSIC, Valencia, Spain. 3 Department of Microbiology and Immunology and Department of Molecular Genetics, Albert Einstein College of Medicine, New York, USA. -

Diarrheagenic Escherichia Coli Signaling and Interactions with Host Innate Immunity And
Diarrheagenic Escherichia coli signaling and interactions with host innate immunity and intestinal microbiota by Gaochan Wang B.S., China Agricultural University, 2007 M.S., The Ohio State University, 2012 AN ABSTRACT OF A DISSERTATION submitted in partial fulfillment of the requirements for the degree DOCTOR OF PHILOSOPHY Department of Diagnostic Medicine/Pathobiology College of Veterinary Medicine KANSAS STATE UNIVERSITY Manhattan, Kansas 2017 Abstract Diarrheagenic Escherichia coli (E. coli) strains are common etiological agents of diarrhea. Diarrheagenic E. coli are classified into enterotoxigenic E. coli (ETEC), Shiga toxin-producing E. coli (STEC or enterohemorrhagic E. coli [EHEC]), enteropathogenic E. coli (EPEC), enteroinvasive E. coli (EIEC), enteroaggregative E. coli (EAEC), diffuse-adherent E. coli (DAEC), and adherent invasive E. coli (AIEC). In addition to encoding toxins that cause diarrhea, diarrheagenic E. coli have evolved numerous strategies to interfere with host defenses. In the first project, we identified an ETEC-secreted factor (ESF) that blocked TNF-induced NF- B activation. One of the consequences of TNF-induced NF-B activation is the production of pro-inflammatory cytokines that help to eliminate pathogens. Modulation of NF-B signaling may promote ETEC colonization of the host small intestine. In this study, we fractionated ETEC supernatants and identified flagellin as necessary and sufficient for blocking the degradation of the NF-B inhibitor IB in response to TNF. In the second project, we attempted to identify an ETEC cAMP importer. ETEC diarrhea leads to cAMP release into the lumen of the small intestine. cAMP is a key secondary messenger that regulates ETEC adhesin expression. We hypothesized that a cAMP importer is present in ETEC, accounting for its hypersensitivity to extracellular cAMP. -

Spatial Protein Interaction Networks of the Intrinsically Disordered Transcription Factor C(%3$
Spatial protein interaction networks of the intrinsically disordered transcription factor C(%3$ Dissertation zur Erlangung des akademischen Grades Doctor rerum naturalium (Dr. rer. nat.) im Fach Biologie/Molekularbiologie eingereicht an der Lebenswissenschaftlichen Fakultät der Humboldt-Universität zu Berlin Von Evelyn Ramberger, M.Sc. Präsidentin der Humboldt-Universität zu Berlin Prof. Dr.-Ing.Dr. Sabine Kunst Dekan der Lebenswissenschaftlichen Fakultät der Humboldt-Universität zu Berlin Prof. Dr. Bernhard Grimm Gutachter: 1. Prof. Dr. Achim Leutz 2. Prof. Dr. Matthias Selbach 3. Prof. Dr. Gunnar Dittmar Tag der mündlichen Prüfung: 12.8.2020 For T. Table of Contents Selbstständigkeitserklärung ....................................................................................1 List of Figures ............................................................................................................2 List of Tables ..............................................................................................................3 Abbreviations .............................................................................................................4 Zusammenfassung ....................................................................................................6 Summary ....................................................................................................................7 1. Introduction ............................................................................................................8 1.1. Disordered proteins -

NF-Kappab Activation in Infections with Helicobacter Pylori Or Legionella Pneumophila
Dissertation NF-kappaB activation in infections with Helicobacter pylori or Legionella pneumophila zur Erlangung des akademischen Grades doctor rerum naturalium (Dr. rer. nat) im Fach Biologie eingereicht an der Mathematisch-Naturwissenschaftlichen Fakultät I der Humboldt Universität zu Berlin von Dipl.-Biol. Sina Bartfeld (geb. 06.01.1978 in Berlin) Präsident der Humboldt Universität zu Berlin Prof. Dr. Dr. h.c. Christoph Markschies Dekan der Mathematisch-Naturwissenschaftlichen Fakultät I Prof. Dr. Lutz-Helmut Schön Gutachter: 1. Prof. Thomas F. Meyer 2. Prof. Wolfgang Uckert 3. Prof. Claus Scheidereit Tag der mündlichen Prüfung : 30.6.2009 Table of content Table of content Abbreviations ................................................................................................ 4 Abstract ........................................................................................................ 6 Zusammenfassung ......................................................................................... 7 1 Introduction .............................................................................................. 8 1.1 The transcription factor family NF-κB ....................................................... 9 1.2 The bacterium Helicobacter pylori .......................................................... 16 1.3 The intracellular bacterium Legionella pneumophila .............................. 21 1.4 RNAi-based screens ................................................................................. 23 1.5 Aims of this thesis ................................................................................... -

Chapter 3. the Beginnings of Genomic Biology – Molecular
Chapter 3. The Beginnings of Genomic Biology – Molecular Genetics Contents 3. The beginnings of Genomic Biology – molecular genetics 3.1. DNA is the Genetic Material 3.6.5. Translation initiation, elongation, and termnation 3.2. Watson & Crick – The structure of DNA 3.6.6. Protein Sorting in Eukaryotes 3.3. Chromosome structure 3.7. Regulation of Eukaryotic Gene Expression 3.3.1. Prokaryotic chromosome structure 3.7.1. Transcriptional Control 3.3.2. Eukaryotic chromosome structure 3.7.2. Pre-mRNA Processing Control 3.3.3. Heterochromatin & Euchromatin 3.4. DNA Replication 3.7.3. mRNA Transport from the Nucleus 3.4.1. DNA replication is semiconservative 3.7.4. Translational Control 3.4.2. DNA polymerases 3.7.5. Protein Processing Control 3.4.3. Initiation of replication 3.7.6. Degradation of mRNA Control 3.4.4. DNA replication is semidiscontinuous 3.7.7. Protein Degradation Control 3.4.5. DNA replication in Eukaryotes. 3.8. Signaling and Signal Transduction 3.4.6. Replicating ends of chromosomes 3.8.1. Types of Cellular Signals 3.5. Transcription 3.8.2. Signal Recognition – Sensing the Environment 3.5.1. Cellular RNAs are transcribed from DNA 3.8.3. Signal transduction – Responding to the Environment 3.5.2. RNA polymerases catalyze transcription 3.5.3. Transcription in Prokaryotes 3.5.4. Transcription in Prokaryotes - Polycistronic mRNAs are produced from operons 3.5.5. Beyond Operons – Modification of expression in Prokaryotes 3.5.6. Transcriptions in Eukaryotes 3.5.7. Processing primary transcripts into mature mRNA 3.6. Translation 3.6.1. -

IGA 8/E Chapter 8
8 RNA: Transcription and Processing WORKING WITH THE FIGURES 1. In Figure 8-3, why are the arrows for genes 1 and 2 pointing in opposite directions? Answer: The arrows for genes 1 and 2 indicate the direction of transcription, which is always 5 to 3. The two genes are transcribed from opposite DNA strands, which are antiparallel, so the genes must be transcribed in opposite directions to maintain the 5 to 3 direction of transcription. 2. In Figure 8-5, draw the “one gene” at much higher resolution with the following components: DNA, RNA polymerase(s), RNA(s). Answer: At the higher resolution, the feathery structures become RNA transcripts, with the longer transcripts occurring nearer the termination of the gene. The RNA in this drawing has been straightened out to illustrate the progressively longer transcripts. 3. In Figure 8-6, describe where the gene promoter is located. Chapter Eight 271 Answer: The promoter is located to the left (upstream) of the 3 end of the template strand. From this sequence it cannot be determined how far the promoter would be from the 5 end of the mRNA. 4. In Figure 8-9b, write a sequence that could form the hairpin loop structure. Answer: Any sequence that contains inverted complementary regions separated by a noncomplementary one would form a hairpin. One sequence would be: ACGCAAGCUUACCGAUUAUUGUAAGCUUGAAG The two bold-faced sequences would pair and form a hairpin. The intervening non-bold sequence would be the loop. 5. How do you know that the events in Figure 8-13 are occurring in the nucleus? Answer: The figure shows a double-stranded DNA molecule from which RNA is being transcribed. -

A Sigma Factor and Anti-Sigma Factor That Control Swarming Motility and Biofilm Formation in Pseudomonas Aeruginosa
A sigma factor and anti-sigma factor that control swarming motility and biofilm formation in Pseudomonas aeruginosa The Harvard community has made this article openly available. Please share how this access benefits you. Your story matters Citation McGuffie, Bryan A. 2015. A sigma factor and anti-sigma factor that control swarming motility and biofilm formation in Pseudomonas aeruginosa. Doctoral dissertation, Harvard University, Graduate School of Arts & Sciences. Citable link http://nrs.harvard.edu/urn-3:HUL.InstRepos:17467530 Terms of Use This article was downloaded from Harvard University’s DASH repository, and is made available under the terms and conditions applicable to Other Posted Material, as set forth at http:// nrs.harvard.edu/urn-3:HUL.InstRepos:dash.current.terms-of- use#LAA A σ factor and anti-σ factor that control swarming motility and biofilm formation in Pseudomonas aeruginosa A dissertation presented by Bryan Alexander McGuffie to The Division of Medical Sciences in partial fulfillment of the requirements for the degree of Doctor of Philosophy in the subject of Microbiology and Molecular Genetics Harvard University Cambridge, Massachusetts May 2015 © 2015 Bryan Alexander McGuffie All rights reserved. Dissertation Advisor: Dr. Simon Dove Bryan Alexander McGuffie A σ factor and anti-σ factor that control swarming motility and biofilm formation in P. aeruginosa Abstract Pseudomonas aeruginosa is an environmental bacterium and opportunistic human pathogen of major clinical significance. It is the principal cause of morbidity and mortality in patients with cystic fibrosis (CF) and a leading cause of nosocomial infections. Although the organism is unicellular, P. aeruginosa exhibits two forms of multicellular behaviors when associated with a surface under the right conditions: swarming motility and biofilm formation. -
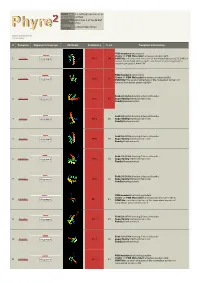
Phyre 2 Results for Q15583
Email [email protected] Description Q15583 Wed May 9 17:58:48 BST Date 2012 Unique Job 376ce548d294102d ID Detailed template information # Template Alignment Coverage 3D Model Confidence % i.d. Template Information PDB header:transcription Chain: A: PDB Molecule:homeobox protein tgif1; 1 c2lk2A_ Alignment 99.9 94 PDBTitle: solution nmr structure of homeobox domain (171-248) of human homeobox2 protein tgif1, northeast structural genomics consortium target3 hr4411b PDB header:transcription Chain: A: PDB Molecule:homeobox protein tgif2lx; 2 c2dmnA_ 99.8 62 Alignment PDBTitle: the solution structure of the homeobox domain of human2 homeobox protein tgif2lx Fold:DNA/RNA-binding 3-helical bundle 3 d1x2na1 Alignment 99.8 49 Superfamily:Homeodomain-like Family:Homeodomain Fold:DNA/RNA-binding 3-helical bundle 4 d1lfup_ Alignment 99.8 36 Superfamily:Homeodomain-like Family:Homeodomain Fold:DNA/RNA-binding 3-helical bundle 5 d1pufb_ Alignment 99.8 36 Superfamily:Homeodomain-like Family:Homeodomain Fold:DNA/RNA-binding 3-helical bundle 6 d1mnmc_ Alignment 99.8 33 Superfamily:Homeodomain-like Family:Homeodomain Fold:DNA/RNA-binding 3-helical bundle 7 d1du6a_ Alignment 99.8 33 Superfamily:Homeodomain-like Family:Homeodomain PDB header:dna binding protein Chain: A: PDB Molecule:homeobox protein barh-like 1; 8 c2dmtA_ 99.7 21 Alignment PDBTitle: solution structure of the homeobox domain of homeobox2 protein barh-like 1 Fold:DNA/RNA-binding 3-helical bundle 9 d1le8b_ Alignment 99.7 25 Superfamily:Homeodomain-like Family:Homeodomain Fold:DNA/RNA-binding -

Transcription
Transcription (CHAPTER 12- Brooker Text) Feb 19, 2007 BIO 184 Dr. Tom Peavy Sequence Complexity in the Genome 60-70% of human DNA fragments are unique DNA sequences 1 • Bacterial mRNA may be polycistronic, which means it encodes two or more polypeptides The Stages of Transcription • Transcription occurs in three stages – Initiation – Elongation – Termination • These steps involve protein-DNA interactions – Proteins such as RNA polymerase interact with DNA sequences Initiation • promoter = recognition site for transcription factors • transcription factors + RNA polymerase to bind to the promoter (initial phase) = closed promoter complex • then DNA is denatured (bubble) = open promoter complex 2 Prokaryotic Transcription • E. coli RNA polymerase = holoenzyme – Core enzyme (Four subunits = α2ββ’) – Sigma factor (One subunit = σ) Initiation stages involve RNA pol holoenzyme • Binding loosely to the DNA • Scaning for promoter region • Forming Open promoter complex • Synthesizing Short stretch of RNA • Releasing Sigma factor Bacterial Promoter Sequence elements that play a key role in transcription (pribnow box) • The RNA transcript is synthesized during ELONGATION step • The DNA strand used as a template for RNA synthesis is termed the template or noncoding strand • The opposite DNA strand is called the coding strand – It has the same base sequence as the RNA transcript • Except that T in DNA corresponds to U in RNA 3 Termination of Bacterial Transcription • short RNA-DNA hybrid is forced to separate = release of newly made RNA • E. coli has -

Mechanism to Control the Cell Lysis and the Cell Survival Strategy in Stationary Phase Under Heat Stress Rashed Noor*
Noor SpringerPlus (2015) 4:599 DOI 10.1186/s40064-015-1415-7 REVIEW Open Access Mechanism to control the cell lysis and the cell survival strategy in stationary phase under heat stress Rashed Noor* Abstract An array of stress signals triggering the bacterial cellular stress response is well known in Escherichia coli and other bacteria. Heat stress is usually sensed through the misfolded outer membrane porin (OMP) precursors in the peri- plasm, resulting in the activation of σE (encoded by rpoE), which binds to RNA polymerase to start the transcription of genes required for responding against the heat stress signal. At the elevated temperatures, σE also serves as the transcription factor for σH (the main heat shock sigma factor, encoded by rpoH), which is involved in the expression of several genes whose products deal with the cytoplasmic unfolded proteins. Besides, oxidative stress in form of the reactive oxygen species (ROS) that accumulate due to heat stress, has been found to give rise to viable but non- culturable (VBNC) cells at the early stationary phase, which is in turn lysed by the σE-dependent process. Such lysis of the defective cells may generate nutrients for the remaining population to survive with the capacity of formation of colony forming units (CFUs). σH is also known to regulate the transcription of the major heat shock proteins (HSPs) required for heat shock response (HSR) resulting in cellular survival. Present review concentrated on the cellular sur- vival against heat stress employing the harmonized impact of σE and σH regulons and the HSPs as well as their inter connectivity towards the maintenance of cellular survival.