VAPB/ALS8 Interacts with FFAT-Like Proteins Including the P97 Cofactor
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Kids First Pediatric Research Program (Kids First) Poster Session at ASHG Accelerating Pediatric Genomics Research Through Collaboration October 15Th, 2019
The Gabriella Miller Kids First Pediatric Research Program (Kids First) Poster Session at ASHG Accelerating Pediatric Genomics Research through Collaboration October 15th, 2019 Background The Gabriella Miller Kids First Pediatric Research Program (Kids First) is a trans- NIH Common Fund program initiated in response to the 2014 Gabriella Miller Kids First Research Act. The program’s vision is to alleviate suffering from childhood cancer and structural birth defects by fostering collaborative research to uncover the etiology of these diseases and support data sharing within the pediatric research community. This is implemented through developing the Gabriella Miller Kids First Data Resource (Kids First Data Resource) and populating this resource with whole genome sequence datasets and associated clinical and phenotypic information. Both childhood cancers and structural birth defects are critical and costly conditions associated with substantial morbidity and mortality. Elucidating the underlying genetic etiology of these diseases has the potential to profoundly improve preventative measures, diagnostics, and therapeutic interventions. Purpose During this evening poster session, attendees will gain a broad understanding of the utility of the genomic data generated by Kids First, learn about the progress of Kids First X01 cohort projects, and observe demonstrations of the tools and functionalities of the recently launched Kids First Data Resource Portal. The session is an opportunity for the scientific community and public to engage with Kids First investigators, collaborators, and a growing community of researchers, patient foundations, and families. Several other NIH and external data efforts will present posters and be available to discuss collaboration opportunities as we work together to accelerate pediatric research. -

Defining the Kv2.1–Syntaxin Molecular Interaction Identifies a First-In-Class Small Molecule Neuroprotectant
Defining the Kv2.1–syntaxin molecular interaction identifies a first-in-class small molecule neuroprotectant Chung-Yang Yeha,b,1, Zhaofeng Yec,d,1, Aubin Moutale, Shivani Gaura,b, Amanda M. Hentonf,g, Stylianos Kouvarosf,g, Jami L. Salomana, Karen A. Hartnett-Scotta,b, Thanos Tzounopoulosa,f,g, Rajesh Khannae, Elias Aizenmana,b,g,2, and Carlos J. Camachoc,2 aDepartment of Neurobiology, University of Pittsburgh School of Medicine, Pittsburgh, PA 15261; bPittsburgh Institute for Neurodegenerative Diseases, University of Pittsburgh School of Medicine, Pittsburgh, PA 15261; cDepartment of Computational and Systems Biology, University of Pittsburgh School of Medicine, Pittsburgh, PA 15261; dSchool of Medicine, Tsinghua University, Beijing 100871, China; eDepartment of Pharmacology, College of Medicine, University of Arizona, Tucson, AZ 85724; fDepartment of Otolaryngology, University of Pittsburgh School of Medicine, Pittsburgh, PA 15261; and gPittsburgh Hearing Research Center, University of Pittsburgh School of Medicine, Pittsburgh, PA 15261 Edited by Lily Yeh Jan, University of California, San Francisco, CA, and approved June 19, 2019 (received for review February 27, 2019) + The neuronal cell death-promoting loss of cytoplasmic K follow- (13). The Kv2.1-dependent cell death pathway is normally initiated ing injury is mediated by an increase in Kv2.1 potassium channels in by the oxidative liberation of zinc from intracellular metal-binding the plasma membrane. This phenomenon relies on Kv2.1 binding to proteins (14), leading to the sequential phosphorylation of syntaxin 1A via 9 amino acids within the channel intrinsically disor- Kv2.1 residues Y124 and S800 by Src and p38 kinases, respectively dered C terminus. Preventing this interaction with a cell and blood- (15–17). -

Chromatin-Associated Degradation Is Defined by UBXN-3/FAF1 To
ARTICLE Received 27 Jul 2015 | Accepted 5 Jan 2016 | Published 4 Feb 2016 DOI: 10.1038/ncomms10612 OPEN Chromatin-associated degradation is defined by UBXN-3/FAF1 to safeguard DNA replication fork progression Andre´ Franz1, Paul A. Pirson1, Domenic Pilger1,2, Swagata Halder2, Divya Achuthankutty2, Hamid Kashkar3, Kristijan Ramadan2 & Thorsten Hoppe1 The coordinated activity of DNA replication factors is a highly dynamic process that involves ubiquitin-dependent regulation. In this context, the ubiquitin-directed ATPase CDC-48/p97 recently emerged as a key regulator of chromatin-associated degradation in several of the DNA metabolic pathways that assure genome integrity. However, the spatiotemporal control of distinct CDC-48/p97 substrates in the chromatin environment remained unclear. Here, we report that progression of the DNA replication fork is coordinated by UBXN-3/FAF1. UBXN-3/FAF1 binds to the licensing factor CDT-1 and additional ubiquitylated proteins, thus promoting CDC-48/p97-dependent turnover and disassembly of DNA replication factor complexes. Consequently, inactivation of UBXN-3/FAF1 stabilizes CDT-1 and CDC-45/GINS on chromatin, causing severe defects in replication fork dynamics accompanied by pronounced replication stress and eventually resulting in genome instability. Our work identifies a critical substrate selection module of CDC-48/p97 required for chromatin- associated protein degradation in both Caenorhabditis elegans and humans, which is relevant to oncogenesis and aging. 1 Institute for Genetics and CECAD Research Center, University of Cologne, Joseph-Stelzmann-Str. 26, 50931 Cologne, Germany. 2 Department of Oncology, University of Oxford, Cancer Research UK/Medical Research Council Oxford, Institute for Radiation Oncology, Old Road Campus Research Building, OX3 7DQ Oxford, UK. -

NLRP2 and FAF1 Deficiency Blocks Early Embryogenesis in the Mouse
REPRODUCTIONRESEARCH NLRP2 and FAF1 deficiency blocks early embryogenesis in the mouse Hui Peng1,*, Haijun Liu2,*, Fang Liu1, Yuyun Gao1, Jing Chen1, Jianchao Huo1, Jinglin Han1, Tianfang Xiao1 and Wenchang Zhang1 1College of Animal Science, Fujian Agriculture and Forestry University, Fujian, Fuzhou, People’s Republic of China and 2Tianjin Institute of Animal Science and Veterinary Medicine, Tianjin, People’s Republic of China Correspondence should be addressed to W Zhang; Email: [email protected] *(H Peng and H Liu contributed equally to this work) Abstract Nlrp2 is a maternal effect gene specifically expressed by mouse ovaries; deletion of this gene from zygotes is known to result in early embryonic arrest. In the present study, we identified FAF1 protein as a specific binding partner of the NLRP2 protein in both mouse oocytes and preimplantation embryos. In addition to early embryos, both Faf1 mRNA and protein were detected in multiple tissues. NLRP2 and FAF1 proteins were co-localized to both the cytoplasm and nucleus during the development of oocytes and preimplantation embryos. Co-immunoprecipitation assays were used to confirm the specific interaction between NLRP2 and FAF1 proteins. Knockdown of the Nlrp2 or Faf1 gene in zygotes interfered with the formation of a NLRP2–FAF1 complex and led to developmental arrest during early embryogenesis. We therefore conclude that NLRP2 interacts with FAF1 under normal physiological conditions and that this interaction is probably essential for the successful development of cleavage-stage mouse embryos. Our data therefore indicated a potential role for NLRP2 in regulating early embryo development in the mouse. Reproduction (2017) 154 245–251 Introduction (Peng et al. -

Signal Peptide Peptidase‐Like 2C Impairs Vesicular Transport And
Article Signal peptide peptidase-like 2c impairs vesicular transport and cleaves SNARE proteins Alkmini A Papadopoulou1, Stephan A Müller2, Torben Mentrup3, Merav D Shmueli2,4,5, Johannes Niemeyer3, Martina Haug-Kröper1, Julia von Blume6, Artur Mayerhofer7, Regina Feederle2,8,9 , Bernd Schröder3,10 , Stefan F Lichtenthaler2,5,9 & Regina Fluhrer1,2,* Abstract Introduction Members of the GxGD-type intramembrane aspartyl proteases The high degree of compartmentalization in eukaryotic cells creates have emerged as key players not only in fundamental cellular a need for specific and precise protein trafficking. To meet this need, processes such as B-cell development or protein glycosylation, but cells have developed a complex system of vesicle transport that also in development of pathologies, such as Alzheimer’s disease or ensures safe sorting of cargo proteins, in particular between the dif- hepatitis virus infections. However, one member of this protease ferent compartments of the secretory pathway [1–3]. Vesicles origi- family, signal peptide peptidase-like 2c (SPPL2c), remains orphan nate from a donor membrane, translocate in a targeted manner, and and its capability of proteolysis as well as its physiological function get specifically tethered to the target membrane, before fusing with is still enigmatic. Here, we demonstrate that SPPL2c is catalytically it. Soluble N-ethylmaleimide-sensitive factor attachment protein active and identify a variety of SPPL2c candidate substrates using receptor (SNARE) proteins are known since three decades to medi- proteomics. The majority of the SPPL2c candidate substrates clus- ate specific membrane fusion [4,5]. So far, 38 SNARE proteins have ter to the biological process of vesicular trafficking. -

Genome Sequences of Tropheus Moorii and Petrochromis Trewavasae, Two Eco‑Morphologically Divergent Cichlid Fshes Endemic to Lake Tanganyika C
www.nature.com/scientificreports OPEN Genome sequences of Tropheus moorii and Petrochromis trewavasae, two eco‑morphologically divergent cichlid fshes endemic to Lake Tanganyika C. Fischer1,2, S. Koblmüller1, C. Börger1, G. Michelitsch3, S. Trajanoski3, C. Schlötterer4, C. Guelly3, G. G. Thallinger2,5* & C. Sturmbauer1,5* With more than 1000 species, East African cichlid fshes represent the fastest and most species‑rich vertebrate radiation known, providing an ideal model to tackle molecular mechanisms underlying recurrent adaptive diversifcation. We add high‑quality genome reconstructions for two phylogenetic key species of a lineage that diverged about ~ 3–9 million years ago (mya), representing the earliest split of the so‑called modern haplochromines that seeded additional radiations such as those in Lake Malawi and Victoria. Along with the annotated genomes we analysed discriminating genomic features of the study species, each representing an extreme trophic morphology, one being an algae browser and the other an algae grazer. The genomes of Tropheus moorii (TM) and Petrochromis trewavasae (PT) comprise 911 and 918 Mbp with 40,300 and 39,600 predicted genes, respectively. Our DNA sequence data are based on 5 and 6 individuals of TM and PT, and the transcriptomic sequences of one individual per species and sex, respectively. Concerning variation, on average we observed 1 variant per 220 bp (interspecifc), and 1 variant per 2540 bp (PT vs PT)/1561 bp (TM vs TM) (intraspecifc). GO enrichment analysis of gene regions afected by variants revealed several candidates which may infuence phenotype modifcations related to facial and jaw morphology, such as genes belonging to the Hedgehog pathway (SHH, SMO, WNT9A) and the BMP and GLI families. -

Incredibly Close—A Newly Identified Peroxisome–ER Contact Site in Humans
JCB: Spotlight Incredibly close—A newly identified peroxisome–ER contact site in humans Maya Schuldiner and Einat Zalckvar Department of Molecular Genetics, Weizmann Institute of Science, Rehovot 7610001, Israel Peroxisomes are tiny organelles that control important residents of contact sites and to mediate membrane association and diverse metabolic processes via their interplay with through their major sperm protein (MSP) domain (Wyles and Ridgway, 2004), they were exciting candidates for tethers. The other organelles, including the endoplasmic reticulum MSP domain of VAPs interacts with proteins that contain two (ER). In this issue, Costello et al. (2017. J. Cell Biol. phenylalanines (FF) in an acidic tract (FFAT) motif. When the https://doi.org/10.1083/jcb.201607055) and Hua et MSP and FFAT motifs are located in proteins that are anchored al. (2017. J. Cell Biol. https://doi.org/10.1083/jcb to opposing organelle membranes, the MSP–FFAT interac- tion zippers up the contact. Hua et al. (2017) looked among .201608128) identify a peroxisome–ER contact site in their PEX16-binding candidates for proteins with a FFAT do- human cells held together by a tethering complex of main and identified ACBD5. The observation that peroxisomal VAPA/B (vesicle-associated membrane protein–associated ACBD5 contained a FFAT domain and could be found in the same protein complex as the ER tethering proteins VAPA and proteins A/B) and ACBD5 (acyl Co-A binding protein 5). VAPB led both groups to investigate whether or not these pro- teins play a role in peroxisome–ER tethering. Peroxisomes are organelles enclosed by a single membrane We have recently put in place a suggestion to term a pro- that exist in most eukaryotic organisms and cells and are in- tein a tether only if it abides by three criteria (Eisenberg-Bord volved in various metabolic, as well as nonmetabolic, cellular et al., 2016). -
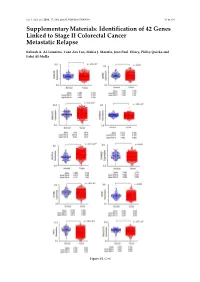
Identification of 42 Genes Linked to Stage II Colorectal Cancer Metastatic Relapse
Int. J. Mol. Sci. 2016, 17, 598; doi:10.3390/ijms17040598 S1 of S16 Supplementary Materials: Identification of 42 Genes Linked to Stage II Colorectal Cancer Metastatic Relapse Rabeah A. Al-Temaimi, Tuan Zea Tan, Makia J. Marafie, Jean Paul Thiery, Philip Quirke and Fahd Al-Mulla Figure S1. Cont. Int. J. Mol. Sci. 2016, 17, 598; doi:10.3390/ijms17040598 S2 of S16 Figure S1. Mean expression levels of fourteen genes of significant association with CRC DFS and OS that are differentially expressed in normal colon compared to CRC tissues. Each dot represents a sample. Table S1. Copy number aberrations associated with poor disease-free survival and metastasis in early stage II CRC as predicted by STAC and SPPS combined methodologies with resident gene symbols. CN stands for copy number, whereas CNV is copy number variation. Region Cytoband % of CNV Count of Region Event Gene Symbols Length Location Overlap Genes chr1:113,025,076–113,199,133 174,057 p13.2 CN Loss 0.0 2 AKR7A2P1, SLC16A1 chr1:141,465,960–141,822,265 356,305 q12–q21.1 CN Gain 95.9 1 SRGAP2B MIR5087, LOC10013000 0, FLJ39739, LOC10028679 3, PPIAL4G, PPIAL4A, NBPF14, chr1:144,911,564–146,242,907 1,331,343 q21.1 CN Gain 99.6 16 NBPF15, NBPF16, PPIAL4E, NBPF16, PPIAL4D, PPIAL4F, LOC645166, LOC388692, FCGR1C chr1:177,209,428–177,226,812 17,384 q25.3 CN Gain 0.0 0 chr1:197,652,888–197,676,831 23,943 q32.1 CN Gain 0.0 1 KIF21B chr1:201,015,278–201,033,308 18,030 q32.1 CN Gain 0.0 1 PLEKHA6 chr1:201,289,154–201,298,247 9093 q32.1 CN Gain 0.0 0 chr1:216,820,186–217,043,421 223,235 q41 CN -

Human Induced Pluripotent Stem Cell–Derived Podocytes Mature Into Vascularized Glomeruli Upon Experimental Transplantation
BASIC RESEARCH www.jasn.org Human Induced Pluripotent Stem Cell–Derived Podocytes Mature into Vascularized Glomeruli upon Experimental Transplantation † Sazia Sharmin,* Atsuhiro Taguchi,* Yusuke Kaku,* Yasuhiro Yoshimura,* Tomoko Ohmori,* ‡ † ‡ Tetsushi Sakuma, Masashi Mukoyama, Takashi Yamamoto, Hidetake Kurihara,§ and | Ryuichi Nishinakamura* *Department of Kidney Development, Institute of Molecular Embryology and Genetics, and †Department of Nephrology, Faculty of Life Sciences, Kumamoto University, Kumamoto, Japan; ‡Department of Mathematical and Life Sciences, Graduate School of Science, Hiroshima University, Hiroshima, Japan; §Division of Anatomy, Juntendo University School of Medicine, Tokyo, Japan; and |Japan Science and Technology Agency, CREST, Kumamoto, Japan ABSTRACT Glomerular podocytes express proteins, such as nephrin, that constitute the slit diaphragm, thereby contributing to the filtration process in the kidney. Glomerular development has been analyzed mainly in mice, whereas analysis of human kidney development has been minimal because of limited access to embryonic kidneys. We previously reported the induction of three-dimensional primordial glomeruli from human induced pluripotent stem (iPS) cells. Here, using transcription activator–like effector nuclease-mediated homologous recombination, we generated human iPS cell lines that express green fluorescent protein (GFP) in the NPHS1 locus, which encodes nephrin, and we show that GFP expression facilitated accurate visualization of nephrin-positive podocyte formation in -

Variability in the Degree of Expression of Phosphorylated I B in Chronic
6796 Vol. 10, 6796–6806, October 15, 2004 Clinical Cancer Research Featured Article Variability in the Degree of Expression of Phosphorylated IB␣ in Chronic Lymphocytic Leukemia Cases With Nodal Involvement Antonia Rodrı´guez,1 Nerea Martı´nez,1 changes in the expression profile (mRNA and protein ex- Francisca I. Camacho,1 Elena Ruı´z-Ballesteros,2 pression) and clinical outcome in a series of CLL cases with 2 1 lymph node involvement. Activation of NF-B, as deter- Patrocinio Algara, Juan-Fernando Garcı´a, ␣ 3 5 mined by the expression of p-I B , was associated with the Javier Mena´rguez, Toma´s Alvaro, expression of a set of genes comprising key genes involved in 6 4 Manuel F. Fresno, Fernando Solano, the control of B-cell receptor signaling, signal transduction, Manuela Mollejo,2 Carmen Martin,1 and and apoptosis, including SYK, LYN, BCL2, CCR7, BTK, Miguel A. Piris1 PIK3CD, and others. Cases with increased expression of ␣ 1Molecular Pathology Program, Centro Nacional de Investigaciones p-I B showed longer overall survival than cases with lower Oncolo´gicas, Madrid, Spain; 2Department of Genetics and Pathology, expression. A Cox regression model was derived to estimate 3 Hospital Virgen de la Salud, Toledo, Spain; Department of some parameters of prognostic interest: IgVH mutational Pathology, Hospital General Universitario Gregorio Maran˜o´n, ␣ 4 status, ZAP-70, and p-I B expression. The multivariate Madrid, Spain; Department of Hematology, Hospital Nuestra Sen˜ora analysis disclosed p-IB␣ and ZAP-70 expression as inde- del Prado, Talavera de la Reina, Toledo, Spain; 5Department of Pathology, Hospital Verge de la Cinta, Tortosa, Spain; pendent prognostic factors of survival. -

The Rhodococcus Equi Virulence Protein Vapa Disrupts Endolysosome Function and Stimulates Lysosome Biogenesis
This is a repository copy of The Rhodococcus equi virulence protein VapA disrupts endolysosome function and stimulates lysosome biogenesis. White Rose Research Online URL for this paper: https://eprints.whiterose.ac.uk/104956/ Version: Accepted Version Article: Rofe, Adam P, Davis, Luther J, Whittingham, Jean L et al. (3 more authors) (2016) The Rhodococcus equi virulence protein VapA disrupts endolysosome function and stimulates lysosome biogenesis. Microbiology Open. e00416. ISSN 2045-8827 https://doi.org/10.1002/mbo3.416 Reuse This article is distributed under the terms of the Creative Commons Attribution (CC BY) licence. This licence allows you to distribute, remix, tweak, and build upon the work, even commercially, as long as you credit the authors for the original work. More information and the full terms of the licence here: https://creativecommons.org/licenses/ Takedown If you consider content in White Rose Research Online to be in breach of UK law, please notify us by emailing [email protected] including the URL of the record and the reason for the withdrawal request. [email protected] https://eprints.whiterose.ac.uk/ The Rhodococcus equi virulence protein VapA disrupts endolysosome function and stimulates lysosome biogenesis. Adam P. Rofe1, Luther J. Davis2, Jean L. Whittingham3, Elizabeth C. Latimer- Bowman2, Anthony J. Wilkinson3 and Paul R. Pryor1,4* 1 Department of Biology, Wentworth Way, University of York, York. YO10 5DD. United Kingdom; 2 Cambridge Institute for Medical Research and Department of Clinical Biochemistry, University of Cambridge, Addenbrooke’s Hospital, Hills Rd, Cambridge, CB2 0XY; 3 Structural Biology Laboratory, Department of Chemistry, University of York York. -

A New Synuclein-Transgenic Mouse Model for Early Parkinson's Reveals Molecular Features of Preclinical Disease
bioRxiv preprint doi: https://doi.org/10.1101/2020.04.04.016642; this version posted April 5, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. A new synuclein-transgenic mouse model for early Parkinson's reveals molecular features of preclinical disease Diana M Hendrickx1,*,#, Pierre Garcia1,2,#, Amer Ashrafi1, Alessia Sciortino1, Kristopher J Schmit1, Heike Kollmus3, Nathalie Nicot4, Tony Kaoma5, Laurent Vallar6, Manuel Buttini1,*,$, Enrico Glaab1,$ 1 Luxembourg Centre for Systems Biomedicine (LCSB), University of Luxembourg, Belvaux, Luxembourg 2 Laboratoire National de Sant´e(LNS), Neuropathology Unit, Dudelange, Luxembourg 3 Department of Infection Genetics, Helmholtz Centre for Infection Research, Braunschweig, Germany 4 Quantitative Biology Unit, Luxembourg Institute of Health, Strassen, Luxembourg 5 Department of Oncology, Luxembourg Institute of Health, Strassen, Luxembourg 6 Genomics Research Unit, Luxembourg Institute of Health, Luxembourg, Luxembourg * [email protected]; [email protected] # equal contributor $ equal contributor Abstract Understanding Parkinson's disease (PD) in particular in its earliest phases is important for diagnosis and treatment. However, human brain samples are collected post-mortem, reflecting mainly end stage disease. Because brain samples of mouse models can be collected at any stage of the disease process, they are useful to investigate PD progression. Here, we compare ventral midbrain transcriptomics profiles from α-synuclein transgenic mice with a progressive, early PD-like striatum neurodegeneration across different ages using pathway, gene set and network analysis methods.