Expansive and Diverse Phenotypic Landscape of Field Aedes Aegypti Larvae with Differential Susceptibility
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Historical Perspectives on Apple Production: Fruit Tree Pest Management, Regulation and New Insecticidal Chemistries
Historical Perspectives on Apple Production: Fruit Tree Pest Management, Regulation and New Insecticidal Chemistries. Peter Jentsch Extension Associate Department of Entomology Cornell University's Hudson Valley Lab 3357 Rt. 9W; PO box 727 Highland, NY 12528 email: [email protected] Phone 845-691-7151 Mobile: 845-417-7465 http://www.nysaes.cornell.edu/ent/faculty/jentsch/ 2 Historical Perspectives on Fruit Production: Fruit Tree Pest Management, Regulation and New Chemistries. by Peter Jentsch I. Historical Use of Pesticides in Apple Production Overview of Apple Production and Pest Management Prior to 1940 Synthetic Pesticide Development and Use II. Influences Changing the Pest Management Profile in Apple Production Chemical Residues in Early Insect Management Historical Chemical Regulation Recent Regulation Developments Changing Pest Management Food Quality Protection Act of 1996 The Science Behind The Methodology Pesticide Revisions – Requirements For New Registrations III. Resistance of Insect Pests to Insecticides Resistance Pest Management Strategies IV. Reduced Risk Chemistries: New Modes of Action and the Insecticide Treadmill Fermentation Microbial Products Bt’s, Abamectins, Spinosads Juvenile Hormone Analogs Formamidines, Juvenile Hormone Analogs And Mimics Insect Growth Regulators Azadirachtin, Thiadiazine Neonicotinyls Major Reduced Risk Materials: Carboxamides, Carboxylic Acid Esters, Granulosis Viruses, Diphenyloxazolines, Insecticidal Soaps, Benzoyl Urea Growth Regulators, Tetronic Acids, Oxadiazenes , Particle Films, Phenoxypyrazoles, Pyridazinones, Spinosads, Tetrazines , Organotins, Quinolines. 3 I Historical Use of Pesticides in Apple Production Overview of Apple Production and Pest Management Prior to 1940 The apple has a rather ominous origin. Its inception is framed in the biblical text regarding the genesis of mankind. The backdrop appears to be the turbulent setting of what many scholars believe to be present day Iraq. -

Transfluthrin (Insecticides, Acaricides and Products to Control Other Arthropods)
Regulation (EU) n°528/2012 concerning the making available on the market and use of biocidal products Evaluation of active substances Assessment Report Transfluthrin (insecticides, acaricides and products to control other arthropods) 13 March 2014 RMS: the Netherlands Transfluthrin (PT18) Assessment report Finalised in the Standing Committee on Biocidal Products at its meeting on 13 March 2014 CONTENTS 1. STATEMENT OF SUBJECT MATTER AND PURPOSE .................................. 4 1.1. Principle of evaluation .................................................................................... 4 1.2. Purpose of the assessment report ................................................................... 4 1.3. Procedure followed .......................................................................................... 4 2. OVERALL SUMMARY AND CONCLUSIONS ................................................... 6 2.1. Presentation of the Active Substance ............................................................. 6 2.1.1. Identity, Physico-Chemical Properties & Methods of Analysis ....... 6 2.1.2. Intended Uses and Efficacy ................................................................ 8 2.1.3. Classification and Labelling .............................................................. 8 2.2. Summary of the Risk Assessment ................................................................ 11 2.2.1. Human Health Risk Assessment ...................................................... 11 2.2.1.1. Hazard identification ........................................................................ -

Tenth Meeting of the WHO Vector Control Advisory Group
MEETING REPORT 13–15 May 2019 Tenth meeting of the WHO Vector Control Advisory Group MEETING REPORT 13–15 May 2019 Tenth meeting of the WHO Vector Control Advisory Group WHO/CDS/VCAG/2019.02 © World Health Organization 2019 Some rights reserved. This work is available under the Creative Commons Attribution-NonCommercial- ShareAlike 3.0 IGO licence (CC BY-NC-SA 3.0 IGO; https://creativecommons.org/licenses/by-nc-sa/3.0/igo). Under the terms of this licence, you may copy, redistribute and adapt the work for non-commercial purposes, provided the work is appropriately cited, as indicated below. In any use of this work, there should be no suggestion that WHO endorses any specific organization, products or services. The use of the WHO logo is not permitted. If you adapt the work, then you must license your work under the same or equivalent Creative Commons licence. If you create a translation of this work, you should add the following disclaimer along with the suggested citation: “This translation was not created by the World Health Organization (WHO). WHO is not responsible for the content or accuracy of this translation. The original English edition shall be the binding and authentic edition”. Any mediation relating to disputes arising under the licence shall be conducted in accordance with the mediation rules of the World Intellectual Property Organization. Suggested citation. Tenth meeting of the WHO Vector Control Advisory Group. Geneva: World Health Organization; 2018 (WHO/CDS/VCAG/2019.02). Licence: CC BY-NC-SA 3.0 IGO. Cataloguing-in-Publication (CIP) data. -

ORIGINAL ARTICLES AAEM Ann Agric Environ Med 2011, 18, 85–88
ORIGINAL ARTICLES AAEM Ann Agric Environ Med 2011, 18, 85–88 CONTENT OF TRANSFLUTHRIN IN INDOOR AIR DURING THE USE OF ELECTRO-VAPORIZERS Teresa Nazimek1, Magdalena Wasak1, Wojciech Zgrajka1, Waldemar Andrzej Turski1, 2 1Department of Toxicology, Institute of Rural Health, Lublin, Poland 2Department of Experimental and Clinical Pharmacology, Medical University of Lublin, Lublin, Poland Nazimek T, Wasak M, Zgrajka W, Turski WA: Content of transfl uthrin in indoor air dur- ing the use of electro-vaporizers. Ann Agric Environ Med 2011, 18, 85–88. Abstract: The quality of indoor air evokes increasing interest; however, no standards have been developed which determine the content of pesticides in the air of living space. At present, insecticides are increasingly more frequently applied to control household pests, fl ies, mosquitoes, termites and other harmful insects. In this study, the content of transfl uthrin was measured indoors after the application of two consumer products containing this active substance, using commercially available electro-vaporizers. It was found that during the application of insecticides in the form of gel and liquid the mean concentration of transfl uthrin in the air was 1.295–2.422 μg/m3 and 3.817–5.227 μg/m3, respectively. The concentration of an active agent in the air did not depend on the day of application. The concentration of transfl uthrin was higher when used in the form of a liquid than a gel preparation. 18–24 hours after the discontinuation of the use of the preparation no active agent was found in the air. As long as the standards are developed regulating the content of insecticides in the air of living spaces and utility rooms, the most important method of preventing their potential hazardous effect is informing the users of these preparations about the occurrence of active substances in indoor air, and eventual risk of exposure to the effect of pesticides during their application at home. -

Recent Advances on Detection of Insecticides Using Optical Sensors
sensors Review Recent Advances on Detection of Insecticides Using Optical Sensors Nurul Illya Muhamad Fauzi 1, Yap Wing Fen 1,2,*, Nur Alia Sheh Omar 1,2 and Hazwani Suhaila Hashim 2 1 Functional Devices Laboratory, Institute of Advanced Technology, Universiti Putra Malaysia, Serdang 43400, Selangor, Malaysia; [email protected] (N.I.M.F.); [email protected] (N.A.S.O.) 2 Department of Physics, Faculty of Science, Universiti Putra Malaysia, Serdang 43400, Selangor, Malaysia; [email protected] * Correspondence: [email protected] Abstract: Insecticides are enormously important to industry requirements and market demands in agriculture. Despite their usefulness, these insecticides can pose a dangerous risk to the safety of food, environment and all living things through various mechanisms of action. Concern about the environmental impact of repeated use of insecticides has prompted many researchers to develop rapid, economical, uncomplicated and user-friendly analytical method for the detection of insecticides. In this regards, optical sensors are considered as favorable methods for insecticides analysis because of their special features including rapid detection time, low cost, easy to use and high selectivity and sensitivity. In this review, current progresses of incorporation between recognition elements and optical sensors for insecticide detection are discussed and evaluated well, by categorizing it based on insecticide chemical classes, including the range of detection and limit of detection. Additionally, this review aims to provide powerful insights to researchers for the future development of optical sensors in the detection of insecticides. Citation: Fauzi, N.I.M.; Fen, Y.W.; Omar, N.A.S.; Hashim, H.S. Recent Keywords: insecticides; optical sensor; recognition element Advances on Detection of Insecticides Using Optical Sensors. -

Recommended Classification of Pesticides by Hazard and Guidelines to Classification 2019 Theinternational Programme on Chemical Safety (IPCS) Was Established in 1980
The WHO Recommended Classi cation of Pesticides by Hazard and Guidelines to Classi cation 2019 cation Hazard of Pesticides by and Guidelines to Classi The WHO Recommended Classi The WHO Recommended Classi cation of Pesticides by Hazard and Guidelines to Classi cation 2019 The WHO Recommended Classification of Pesticides by Hazard and Guidelines to Classification 2019 TheInternational Programme on Chemical Safety (IPCS) was established in 1980. The overall objectives of the IPCS are to establish the scientific basis for assessment of the risk to human health and the environment from exposure to chemicals, through international peer review processes, as a prerequisite for the promotion of chemical safety, and to provide technical assistance in strengthening national capacities for the sound management of chemicals. This publication was developed in the IOMC context. The contents do not necessarily reflect the views or stated policies of individual IOMC Participating Organizations. The Inter-Organization Programme for the Sound Management of Chemicals (IOMC) was established in 1995 following recommendations made by the 1992 UN Conference on Environment and Development to strengthen cooperation and increase international coordination in the field of chemical safety. The Participating Organizations are: FAO, ILO, UNDP, UNEP, UNIDO, UNITAR, WHO, World Bank and OECD. The purpose of the IOMC is to promote coordination of the policies and activities pursued by the Participating Organizations, jointly or separately, to achieve the sound management of chemicals in relation to human health and the environment. WHO recommended classification of pesticides by hazard and guidelines to classification, 2019 edition ISBN 978-92-4-000566-2 (electronic version) ISBN 978-92-4-000567-9 (print version) ISSN 1684-1042 © World Health Organization 2020 Some rights reserved. -

LC-MS/MS and GC-MS/MS Multi Residue Pesticide Analysis in Fruit and Vegetable Extracts on a Single Tandem Quadrupole Mass Spectrometer
36 May / June 2017 LC-MS/MS and GC-MS/MS Multi Residue Pesticide Analysis in Fruit and Vegetable Extracts on a Single Tandem Quadrupole Mass Spectrometer by Kari Organtini1, Gareth Cleland1, Eimear McCall2, and Simon Hird2 1Waters Corporation, Milford, MA, USA 2Waters Corporation, Wilmslow, UK Hundreds of pesticides are commercially available and are approved for use on various fruit and vegetable plants to prevent pest infestation and improve shelf-life of fresh produce. Maximum Residue Levels (MRLs) are set at the highest level of pesticide that the relevant regulatory body would expect to find in that crop when it has been treated in accordance with good agricultural practice. In the EU, if a pesticide is not explicitly mentioned in the MRL legislation, a default MRL is used for enforcement. This default value is set to be equal to the limit of quantification (LOQ) achievable with the analytical methods used for analysis. National authorities control and enforce MRLs by testing samples for pesticide residue levels using analytical surveillance programs. These programs check for compliance with MRLs, assess dietary exposure, and check for use of unauthorised pesticides. The food industry also carries out its own due diligence analyses. Mass spectrometry (MS) coupled with fragile or easily fragmented compounds. in this study (listed in appendix tables) were both gas chromatography (GC) and liquid APGC ionisation can occur using two chosen to cover a wide range of different chromatography (LC) is needed to provide mechanisms; proton transfer (wet source) pesticide classes and chemistries. The multi comprehensive analysis of a wide range of or charge transfer (dry source). -

Pesticide Pollution in Nepal
Status of Pesticide pollution in Nepal Kanti Shrestha, Ph.D Chief Scientific officer Nepal Academy of Science and Technology, Khumaltar, Lalitpur 20th June 2014 National Symposium on Pesticide Pollution GLOBAL CONSUMPTION PATTERN OF PESTICIDES S.N. Country Pesticide Use-a.i.(kg/ha) 1 India 0.38 2 China 2.0-2.5 3 Pakistan 1.3 4 Bangladesh 0.4 5 Bhutan 0.1 6 Republic of Korea 6.6 7 Japan 12 8 Indonesia 0.575 9 Thailand 1.1 10 Netherlands 9.4 11 Europe 1.9 12 USA 1.5 13 Germany 2.5 14 Norway 0.4 15 Africa 1.23 16 Nepal 0.142 Source: Pesticide use by country", World Resources Institute, livemint.com, 2007, Less pesticide on your food: Good news, 2012 and Trends in pesticide use and drivers for safer pest management, 2008 Pesticide trader • Private pesticide dealership started from 1995 when AIC stopped bulk purchasing of pesticides. • Network of private wholesalers & retailers in most of the districts Direct purchase • Some institution have been directly purchasing pesticides independently. • National Seed Company, Cotton Development Board, Nepal Food Corporation as well as the Epidemiology and Disease Control Division. List of Company involved in Pesticide formulation S. No. Name of Company Address /Farm 1 Kissan Agro Chemicals Birgunj, Parsa 2 Amit Pesticide Udhyog Birgunj, Parsa 3 Nepal Agro Industries Parsauni-2 Bara 4 Khoteja Agro Pvt. Ltd. Jagatpur, Chitwan 5 Nepal Krishi Rasayan Birgunj, Parsa Total registered pesticides 2068/6/31 S N Type of Pesticides Trade Name Common Name 1 Insecticides 500 40 2 Fungicides 229 33 3 Rodenticides 10 2 4 Weedicides 88 15 5 Bio-pesticides 19 7 6 Bactericides 6 2 7 Acaricides 7 3 Total 859 102 Classification Organochlorines (eg.DDT, BHC etc.) Organophosphates (eg.Malathion, Phorate, Quinalphos, Fenitrothion etc.) Carbamates (eg. -

The Insecticides Act, 1968 (Act No.46 of 1968)
The Insecticides Act, 1968 (Act No.46 of 1968) An Act to regulate the import, manufactures, sale, transport, distribution and use of insecticides with a view to prevent risk to human beings or animals and for matters connected therewith. [2 nd September 1968] Be it enacted by Parliament in the Nineteenth Year of the Republic of India as follows: 1. Short title, extent and commencement. * a. This Act may be called the Insecticides Act, 1968. b. It extends to the whole of India. c. It shall come into force on such date as the Central Government may, by notification in the official Gazette, appoint and different dates may be appointed for different States and for different provisions of Act. 2. Application of other laws not barred * The provisions of this Act shall be in addition to, and not in derogation of, any other law for the time being in force. 3. Definitions- In this Act, unless the context otherwise requires- a. "animals" means animals useful to human beings and includes fish and fowl, and such kinds of wild life as the Central Government may, by notification in the official Gazette, specify, being kinds which in its opinion, it is desirable to protect or preserve; b. "Board" means the Central Insecticides Board constituted under Sec.4; c. "Central Insecticides Laboratory" means the Central Insecticides Laboratory established, or as the case may be, the institution specified under Sec.16; d. "Import" means bringing into any place within the territories to which this Act extends from a place outside those territories; e. "Insecticide" means- i. -
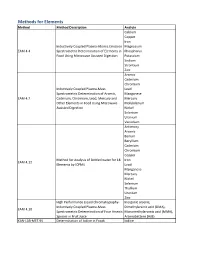
Method Description
Methods for Elements Method Method Description Analyte Calcium Copper Iron Inductively Coupled Plasma-Atomic Emission Magnesium EAM 4.4 Spectrometric Determination of Elements in Phosphorus Food Using Microwave Assisted Digestion Potassium Sodium Strontium Zinc Arsenic Cadmium Chromium Inductively Coupled Plasma-Mass Lead Spectrometric Determination of Arsenic, Manganese EAM 4.7 Cadmium, Chromium, Lead, Mercury and Mercury Other Elements in Food Using Microwave Molybdenum Assisted Digestion Nickel Selenium Uranium Vanadium Antimony Arsenic Barium Beryllium Cadmium Chromium Copper Method for Analysis of Bottled water for 18 Iron EAM 4.12 Elements by ICPMS Lead Manganese Mercury Nickel Selenium Thallium Uranium Zinc High Performance Liquid Chromatography- Inorganic arsenic, Inductively Coupled Plasma-Mass Dimethylarsinic acid (DMA), EAM 4.10 Spectrometric Determination of Four Arsenic Monomethylarsonic acid (MMA), Species in Fruit Juice Arsenobetaine (AsB) KAN-LAB-MET.95 Determination of Iodine in Foods Iodine Methods for Radionuclides Method Method Description Analyte Determination of Strontium-90 in Foods by WEAC.RN.METHOD.2.0 Strontium-90 Internal Gas-Flow Proportional Counting Americium-241 Cesium-134 Cesium-137 Determination of Gamma-Ray Emitting Cobalt-60 WEAC.RN.METHOD.3.0 Radionuclides in Foods by High-Purity Potassium-40 Germanium Spectrometry Radium-226 Ruthenium-103 Ruthenium-106 Thorium-232 Methods for Pesticides/Industrial Chemicals Method Method Description Analyte Extraction Method: Analysis of Pesticides KAN-LAB-PES.53 and -

The 2018 European Union Report on Pesticide Residues in Food
SCIENTIFIC REPORT ADOPTED: 24 February 2020 doi: 10.2903/j.efsa.2020.6057 The 2018 European Union report on pesticide residues in food European Food Safety Authority (EFSA), Paula Medina-Pastor and Giuseppe Triacchini Abstract Under EU legislation (Article 32, Regulation (EC) No 396/2005), EFSA provides an annual report which analyses pesticide residue levels in foods on the European market. The analysis is based on data from the official national control activities carried out by EU Member States, Iceland and Norway and includes a subset of data from the EU-coordinated control programme which uses a randomised sampling strategy. For 2018, 95.5% of the overall 91,015 samples analysed fell below the maximum residue level (MRL), 4.5% exceeded this level, of which 2.7% were non-compliant, i.e. samples exceeding the MRL after taking into account the measurement uncertainty. For the subset of 11,679 samples analysed as part of the EU-coordinated control programme, 1.4% exceeded the MRL and 0.9% were non-compliant. Table grapes and sweet peppers/bell peppers were among the food products that most frequently exceeded the MRLs. To assess acute and chronic risk to consumer health, dietary exposure to pesticide residues was estimated and compared with health-based guidance values. The findings suggest that the assessed levels for the food commodities analysed are unlikely to pose concern for consumer health. However, a number of recommendations are proposed to increase the efficiency of European control systems (e.g. optimising traceability), thereby continuing to ensure a high level of consumer protection. -

Registration Division Conventional Pesticides -Branch and Product
Registration Division Conventional Pesticides - Branch and Product Manager (PM) Assignments For a list of Branch contacts, please click the following link: http://www2.epa.gov/pesticide-contacts/contacts-office-pesticide-programs-registration-division Branch FB=Fungicide Branch. FHB=Fungicide Herbicide Branch. HB=Herbicide Branch. Abbreviations: IVB*= Invertebrate-Vertebrate Branch 1, 2 or 3. MUERB=Minor Use and Emergency Response Branch. Chemical Branch PM 1-Decanol FHB RM 20 1-Naphthaleneacetamide FHB RM 20 2, 4-D, Choline salt HB RM 23 2,4-D HB RM 23 2,4-D, 2-ethylhexyl ester HB RM 23 2,4-D, butoxyethyl ester HB RM 23 2,4-D, diethanolamine salt HB RM 23 2,4-D, dimethylamine salt HB RM 23 2,4-D, isopropyl ester HB RM 23 2,4-D, isopropylamine salt HB RM 23 2,4-D, sodium salt HB RM 23 2,4-D, triisopropanolamine salt HB RM 23 2,4-DB HB RM 23 2,4-DP HB RM 23 2,4-DP, diethanolamine salt HB RM 23 2,4-DP-p HB RM 23 2,4-DP-p, 2-ethylhexyl ester FB RM 21 2,4-DP-p, DMA salt HB RM 23 2-EEEBC FB RM 21 2-Phenylethyl propionate FHB RM 20 4-Aminopyridine IVB3 RM 07 4-Chlorophenoxyacetic acid FB RM 22 4-vinylcyclohexene diepoxide IVB3 RM 07 Abamectin IVB3 RM 07 Acephate IVB2 RM 10 Acequinocyl IVB3 RM 01 Acetaminophen IVB3 RM 07 Acetamiprid IVB3 RM 01 Acetic acid, (2,4-dichlorophenoxy)-, compd. with methanamine (1:1) HB RM 23 Acetic acid, trifluoro- FHB RM 20 Acetochlor HB RM 25 Acibenzolar-s-methyl FHB RM 24 Acid Blue 9 HB RM 23 Acid Yellow 23 HB RM 23 Sunday, June 06, 2021 Page 1 of 17 Chemical Branch PM Acifluorfen HB RM 23 Acrinathrin IVB1 RM 03