A Computational Framework to Study Sub-Cellular RNA Localization
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Management of Large Sets of Image Data Capture, Databases, Image Processing, Storage, Visualization Karol Kozak
Management of large sets of image data Capture, Databases, Image Processing, Storage, Visualization Karol Kozak Download free books at Karol Kozak Management of large sets of image data Capture, Databases, Image Processing, Storage, Visualization Download free eBooks at bookboon.com 2 Management of large sets of image data: Capture, Databases, Image Processing, Storage, Visualization 1st edition © 2014 Karol Kozak & bookboon.com ISBN 978-87-403-0726-9 Download free eBooks at bookboon.com 3 Management of large sets of image data Contents Contents 1 Digital image 6 2 History of digital imaging 10 3 Amount of produced images – is it danger? 18 4 Digital image and privacy 20 5 Digital cameras 27 5.1 Methods of image capture 31 6 Image formats 33 7 Image Metadata – data about data 39 8 Interactive visualization (IV) 44 9 Basic of image processing 49 Download free eBooks at bookboon.com 4 Click on the ad to read more Management of large sets of image data Contents 10 Image Processing software 62 11 Image management and image databases 79 12 Operating system (os) and images 97 13 Graphics processing unit (GPU) 100 14 Storage and archive 101 15 Images in different disciplines 109 15.1 Microscopy 109 360° 15.2 Medical imaging 114 15.3 Astronomical images 117 15.4 Industrial imaging 360° 118 thinking. 16 Selection of best digital images 120 References: thinking. 124 360° thinking . 360° thinking. Discover the truth at www.deloitte.ca/careers Discover the truth at www.deloitte.ca/careers © Deloitte & Touche LLP and affiliated entities. Discover the truth at www.deloitte.ca/careers © Deloitte & Touche LLP and affiliated entities. -

Bioimage Analysis Tools
Bioimage Analysis Tools Kota Miura, Sébastien Tosi, Christoph Möhl, Chong Zhang, Perrine Paul-Gilloteaux, Ulrike Schulze, Simon Norrelykke, Christian Tischer, Thomas Pengo To cite this version: Kota Miura, Sébastien Tosi, Christoph Möhl, Chong Zhang, Perrine Paul-Gilloteaux, et al.. Bioimage Analysis Tools. Kota Miura. Bioimage Data Analysis, Wiley-VCH, 2016, 978-3-527-80092-6. hal- 02910986 HAL Id: hal-02910986 https://hal.archives-ouvertes.fr/hal-02910986 Submitted on 3 Aug 2020 HAL is a multi-disciplinary open access L’archive ouverte pluridisciplinaire HAL, est archive for the deposit and dissemination of sci- destinée au dépôt et à la diffusion de documents entific research documents, whether they are pub- scientifiques de niveau recherche, publiés ou non, lished or not. The documents may come from émanant des établissements d’enseignement et de teaching and research institutions in France or recherche français ou étrangers, des laboratoires abroad, or from public or private research centers. publics ou privés. 2 Bioimage Analysis Tools 1 2 3 4 5 6 Kota Miura, Sébastien Tosi, Christoph Möhl, Chong Zhang, Perrine Pau/-Gilloteaux, - Ulrike Schulze,7 Simon F. Nerrelykke,8 Christian Tischer,9 and Thomas Penqo'" 1 European Molecular Biology Laboratory, Meyerhofstraße 1, 69117 Heidelberg, Germany National Institute of Basic Biology, Okazaki, 444-8585, Japan 2/nstitute for Research in Biomedicine ORB Barcelona), Advanced Digital Microscopy, Parc Científic de Barcelona, dBaldiri Reixac 1 O, 08028 Barcelona, Spain 3German Center of Neurodegenerative -

Cellanimation: an Open Source MATLAB Framework for Microscopy Assays Walter Georgescu1,2,3,∗, John P
Vol. 28 no. 1 2012, pages 138–139 BIOINFORMATICS APPLICATIONS NOTE doi:10.1093/bioinformatics/btr633 Systems biology Advance Access publication November 24, 2011 CellAnimation: an open source MATLAB framework for microscopy assays Walter Georgescu1,2,3,∗, John P. Wikswo1,2,3,4,5 and Vito Quaranta1,3,6 1Vanderbilt Institute for Integrative Biosystems Research and Education, 2Department of Biomedical Engineering, Vanderbilt University, 3Center for Cancer Systems Biology at Vanderbilt, Vanderbilt University Medical Center, Nashville, TN, USA, 4Department of Molecular Physiology and Biophysics, 5Department of Physics and Astronomy, Vanderbilt University and 6Department of Cancer Biology, Vanderbilt University Medical Center, Nashville, TN, USA Associate Editor: Jonathan Wren ABSTRACT 1 INTRODUCTION Motivation: Advances in microscopy technology have led to At present there are a number of microscopy applications on the the creation of high-throughput microscopes that are capable of market, both open-source and commercial, and addressing both generating several hundred gigabytes of images in a few days. very specific and broader microscopy needs. In general, commercial Analyzing such wealth of data manually is nearly impossible and software packages such as Imaris® and MetaMorph® tend to requires an automated approach. There are at present a number include more complete sets of algorithms, covering different kinds of open-source and commercial software packages that allow the of experimental setups, at the cost of ease of customization and user -

Titel Untertitel
KNIME Image Processing Nycomed Chair for Bioinformatics and Information Mining Department of Computer and Information Science Konstanz University, Germany Why Image Processing with KNIME? KNIME UGM 2013 2 The “Zoo” of Image Processing Tools Development Processing UI Handling ImgLib2 ImageJ OMERO OpenCV ImageJ2 BioFormats MatLab Fiji … NumPy CellProfiler VTK Ilastik VIGRA CellCognition … Icy Photoshop … = Single, individual, case specific, incompatible solutions KNIME UGM 2013 3 The “Zoo” of Image Processing Tools Development Processing UI Handling ImgLib2 ImageJ OMERO OpenCV ImageJ2 BioFormats MatLab Fiji … NumPy CellProfiler VTK Ilastik VIGRA CellCognition … Icy Photoshop … → Integration! KNIME UGM 2013 4 KNIME as integration platform KNIME UGM 2013 5 Integration: What and How? KNIME UGM 2013 6 Integration ImgLib2 • Developed at MPI-CBG Dresden • Generic framework for data (image) processing algoritms and data-structures • Generic design of algorithms for n-dimensional images and labelings • http://fiji.sc/wiki/index.php/ImgLib2 → KNIME: used as image representation (within the data cells); basis for algorithms KNIME UGM 2013 7 Integration ImageJ/Fiji • Popular, highly interactive image processing tool • Huge base of available plugins • Fiji: Extension of ImageJ1 with plugin-update mechanism and plugins • http://rsb.info.nih.gov/ij/ & http://fiji.sc/ → KNIME: ImageJ Macro Node KNIME UGM 2013 8 Integration ImageJ2 • Next-generation version of ImageJ • Complete re-design of ImageJ while maintaining backwards compatibility • Based on ImgLib2 -

A Bioimage Informatics Platform for High-Throughput Embryo Phenotyping James M
Briefings in Bioinformatics, 19(1), 2018, 41–51 doi: 10.1093/bib/bbw101 Advance Access Publication Date: 14 October 2016 Paper A bioimage informatics platform for high-throughput embryo phenotyping James M. Brown, Neil R. Horner, Thomas N. Lawson, Tanja Fiegel, Simon Greenaway, Hugh Morgan, Natalie Ring, Luis Santos, Duncan Sneddon, Lydia Teboul, Jennifer Vibert, Gagarine Yaikhom, Henrik Westerberg and Ann-Marie Mallon Corresponding author: James Brown, MRC Harwell Institute, Harwell Campus, Oxfordshire, OX11 0RD. Tel. þ44-0-1235-841237; Fax: +44-0-1235-841172; E-mail: [email protected] Abstract High-throughput phenotyping is a cornerstone of numerous functional genomics projects. In recent years, imaging screens have become increasingly important in understanding gene–phenotype relationships in studies of cells, tissues and whole organisms. Three-dimensional (3D) imaging has risen to prominence in the field of developmental biology for its ability to capture whole embryo morphology and gene expression, as exemplified by the International Mouse Phenotyping Consortium (IMPC). Large volumes of image data are being acquired by multiple institutions around the world that encom- pass a range of modalities, proprietary software and metadata. To facilitate robust downstream analysis, images and metadata must be standardized to account for these differences. As an open scientific enterprise, making the data readily accessible is essential so that members of biomedical and clinical research communities can study the images for them- selves without the need for highly specialized software or technical expertise. In this article, we present a platform of soft- ware tools that facilitate the upload, analysis and dissemination of 3D images for the IMPC. -

Machine Learning for Blob Detection in High-Resolution 3D Microscopy Images
DEGREE PROJECT IN COMPUTER SCIENCE AND ENGINEERING, SECOND CYCLE, 30 CREDITS STOCKHOLM, SWEDEN 2018 Machine learning for blob detection in high-resolution 3D microscopy images MARTIN TER HAAK KTH ROYAL INSTITUTE OF TECHNOLOGY SCHOOL OF ELECTRICAL ENGINEERING AND COMPUTER SCIENCE Machine learning for blob detection in high-resolution 3D microscopy images MARTIN TER HAAK EIT Digital Data Science Date: June 6, 2018 Supervisor: Vladimir Vlassov Examiner: Anne Håkansson Electrical Engineering and Computer Science (EECS) iii Abstract The aim of blob detection is to find regions in a digital image that dif- fer from their surroundings with respect to properties like intensity or shape. Bio-image analysis is a common application where blobs can denote regions of interest that have been stained with a fluorescent dye. In image-based in situ sequencing for ribonucleic acid (RNA) for exam- ple, the blobs are local intensity maxima (i.e. bright spots) correspond- ing to the locations of specific RNA nucleobases in cells. Traditional methods of blob detection rely on simple image processing steps that must be guided by the user. The problem is that the user must seek the optimal parameters for each step which are often specific to that image and cannot be generalised to other images. Moreover, some of the existing tools are not suitable for the scale of the microscopy images that are often in very high resolution and 3D. Machine learning (ML) is a collection of techniques that give computers the ability to ”learn” from data. To eliminate the dependence on user parameters, the idea is applying ML to learn the definition of a blob from labelled images. -

Outline of Machine Learning
Outline of machine learning The following outline is provided as an overview of and topical guide to machine learning: Machine learning – subfield of computer science[1] (more particularly soft computing) that evolved from the study of pattern recognition and computational learning theory in artificial intelligence.[1] In 1959, Arthur Samuel defined machine learning as a "Field of study that gives computers the ability to learn without being explicitly programmed".[2] Machine learning explores the study and construction of algorithms that can learn from and make predictions on data.[3] Such algorithms operate by building a model from an example training set of input observations in order to make data-driven predictions or decisions expressed as outputs, rather than following strictly static program instructions. Contents What type of thing is machine learning? Branches of machine learning Subfields of machine learning Cross-disciplinary fields involving machine learning Applications of machine learning Machine learning hardware Machine learning tools Machine learning frameworks Machine learning libraries Machine learning algorithms Machine learning methods Dimensionality reduction Ensemble learning Meta learning Reinforcement learning Supervised learning Unsupervised learning Semi-supervised learning Deep learning Other machine learning methods and problems Machine learning research History of machine learning Machine learning projects Machine learning organizations Machine learning conferences and workshops Machine learning publications -

A Study on the Core Area Image Processing: Current Research
The International journal of analytical and experimental modal analysis ISSN NO: 0886-9367 A Study on the Core Area Image Processing: Current Research, Softwares and Applications K.Kanimozhi1, Dr.K.Thangadurai2 1Research Scholar, PG and Research Department of Computer Science, Government Arts College, Karur-639005 2Assistant professor and Head, PG and Research Department of Computer Science, Government Arts College, Karur-639005 [email protected] [email protected] Abstract —Among various core subjects such as, networking, data mining, data warehousing, cloud computing, artificial intelligence, image processing, place a vital role on new emerging ideas. Image processing is a technique to perform some operations with an image to get its in enhanced version or to retrieve some information from it. The image can be of signal dispension, video for frame or a photograph while the system treats that as a 2D- signal. This image processing area has its applications in various wide range such as business, engineering, transport systems, remote sensing, tracking applications, surveillance systems, bio medical fields, visual inspection systems etc.., Its purpose can be of: to create best version of images(image sharpening and restoring), retrieving of images, pattern measuring and recognizing images. Keywords — Types, Flow, Projects, MATLAB, Python. I. INTRODUCTION In computer concepts, digital image processing technically means that of using computer algorithms to process digital images. Initially in1960’s the image processing had been applied for those highly technical applications such as satellite imagery, medical imaging, character recognition, wire-photo conversion standards, video phone and photograph enhancement. Since the cost of processing those images was very high however with the equipment used too. -

NEUBIAS, the Network of Bioimage Analysts and Its Activities
20160712 GerBI NEUBIAS, the network of bioimage analysts and its activities Kota Miura Freelance Image Analyst Nikon Imaging Center, Uni-Heidelberg National Institute of Basic Biology, Okazaki Arnold Dodel Stem Tillia sp. 20160712 GerBI Definition of Image Analysis Gonzalez & Woods, “Digital Image Processing” “ Image analysis is a process of discovering, identifying, and understanding patterns that are relevant to the performance of an image-based task. One of the principal goals of image analysis by computer is to endow a machine with the capability to approximate, in some sense, a similar capability in human beings.” e.g. “Computer Vision” 20160712 GerBI Definition of Bioimage Analysis In biology, image analysis is a process of identifying spatial and temporal distribution of biological components in images, and measure their characteristics to study their underlying mechanisms in an unbiased way. We do not have to be bothered with similarity to the human recognition - we have more emphasis on the objectivity of quantitative measurement, rather than how that computer-based recognition becomes in agreement with human recognition. 20160712 GerBI Survey Mar. 2015 1800 people answered in three days 20160712 GerBI Imaging Bottleneck Imaging -> Experiments -> Microscopy (50 million Euros in Germany) -> Image Processing & Analysis -> Results Bottleneck! 20160712 GerBI A Failure of Optimism There will be An Ultimate Software Package that solves all image analysis problems in biology, in a matter of several clicks. ... most likely not. 20160712 GerBI NEUBIAS: Web Platform components: The Image Data implementation of image processing and analysis algorithms. workflow component Workflow: a sequence of components for component biological research. component Software/Library: A package of various component components. -

Phenotypic Image Analysis Software Tools for Exploring and Understanding Big Image Data from Cell-Based Assays
Cell Systems Review Phenotypic Image Analysis Software Tools for Exploring and Understanding Big Image Data from Cell-Based Assays Kevin Smith,1,2 Filippo Piccinini,3 Tamas Balassa,4 Krisztian Koos,4 Tivadar Danka,4 Hossein Azizpour,1,2 and Peter Horvath4,5,* 1KTH Royal Institute of Technology, School of Electrical Engineering and Computer Science, Lindstedtsvagen€ 3, 10044 Stockholm, Sweden 2Science for Life Laboratory, Tomtebodavagen€ 23A, 17165 Solna, Sweden 3Istituto Scientifico Romagnolo per lo Studio e la Cura dei Tumori (IRST) IRCCS, Via P. Maroncelli 40, Meldola, FC 47014, Italy 4Synthetic and Systems Biology Unit, Hungarian Academy of Sciences, Biological Research Center (BRC), Temesva´ ri krt. 62, 6726 Szeged, Hungary 5Institute for Molecular Medicine Finland, University of Helsinki, Tukholmankatu 8, 00014 Helsinki, Finland *Correspondence: [email protected] https://doi.org/10.1016/j.cels.2018.06.001 Phenotypic image analysis is the task of recognizing variations in cell properties using microscopic image data. These variations, produced through a complex web of interactions between genes and the environ- ment, may hold the key to uncover important biological phenomena or to understand the response to a drug candidate. Today, phenotypic analysis is rarely performed completely by hand. The abundance of high-dimensional image data produced by modern high-throughput microscopes necessitates computa- tional solutions. Over the past decade, a number of software tools have been developed to address this need. They use statistical learning methods to infer relationships between a cell’s phenotype and data from the image. In this review, we examine the strengths and weaknesses of non-commercial phenotypic image analysis software, cover recent developments in the field, identify challenges, and give a perspective on future possibilities. -

Neural Network Fast-Classifies Biological Images Using
Neural network fast-classifies biological images using features selected after their random-forests-importance to power smart microscopy. Mael¨ Ballueta,b, Florian Sizairea,g, Youssef El Habouza, Thomas Walterc,d,e,Jer´ emy´ Pontb, Baptiste Girouxb, Otmane Boucharebb, Marc Tramiera,f, Jacques Pecreauxa,∗ aCNRS, Univ Rennes, IGDR - UMR 6290, F-35043 Rennes, France bInscoper SAS, F-35510 Cesson-S´evign´e,France cCentre for Computational Biology (CBIO), MINES ParisTech, PSL University, F-75272 Paris, France dInstitut Curie, F-75248 Paris, France eINSERM, U900, F-75248 Paris, France fUniv Rennes, BIOSIT, UMS CNRS 3480, US INSERM 018, F-35000 Rennes, France gPresent address: Biologics Research, Sanofi R&D, F94400 Vitry sur Seine, France Abstract Artificial intelligence is nowadays used for cell detection and classification in optical microscopy, during post-acquisition analysis. The microscopes are now fully automated and next expected to be smart, to make acquisition decisions based on the images. It calls for analysing them on the fly. Biology further imposes training on a reduced dataset due to cost and time to prepare the samples and have the datasets annotated by experts. We propose here a real-time image processing, compliant with these specifications by balancing accurate detection and execution performance. We characterised the images using a generic, high-dimensional feature extractor. We then classified the images using machine learning for the sake of understanding the contribution of each feature in decision and execution time. We found that the non-linear-classifier random forests outperformed Fisher’s linear discriminant. More importantly, the most discriminant and time-consuming features could be excluded without any significant loss in accuracy, offering a substantial gain in execution time. -
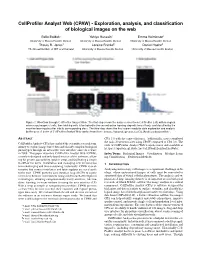
Cellprofiler Analyst Web (CPAW)
CellProfiler Analyst Web (CPAW) - Exploration, analysis, and classification of biological images on the web Bella Baidak* Yahiya Hussain† Emma Kelminson‡ University of Massachusetts Boston University of Massachusetts Boston University of Massachusetts Boston Thouis R. Jones§ Loraine Franke¶ Daniel Haehn|| The Broad Institute of MIT and Harvard University of Massachusetts Boston University of Massachusetts Boston Figure 1: Workflow through CellProfiler Analyst Web: The first step shows the data received from CellProfiler (left) with biological microscopy images of cells, then fetching cells to be trained in the second active learning step with tensorflow.js and classified by the machine learning classifier into its corresponding class. The third step shows the final scores ready for data exploration and analysis by the user. A demo of CellProfiler Analyst Web can be found here: https://mpsych.github.io/CellProfilerAnalystWeb/. ABSTRACT CPA 3.0 with the same efficiency. Additionally, users completed the tasks 20 percent faster using CPAW compared to CPA 3.0. The CellProfiler Analyst (CPA) has enabled the scientific research com- code of CellProfiler Analyst Web is open-source and available at munity to explore image-based data and classify complex biological https://mpsych.github.io/CellProfilerAnalystWeb/. phenotypes through an interactive user interface since its release in 2008. This paper describes CellProfiler Analyst Web (CPAW), Index Terms: Biological Images—Visualization—Machine Learn- a newly redesigned and web-based version of the software, allow- ing Classification—Evaluation Methods; ing for greater accessibility, quicker setup, and facilitating a simple workflow for users. Installation and managing new versions has 1 INTRODUCTION been challenging and time-consuming, historically.