Clawhmmer: a Streaming Hmmer-Search Implementation
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

RDA COVID-19 Recommendations and Guidelines on Data Sharing
RDA COVID-19 Recommendations and Guidelines on Data Sharing DOI: 10.15497/RDA00052 Authors: RDA COVID-19 Working Group Published: 30th June 2020 Abstract: This is the final version of the Recommendations and Guidelines from the RDA COVID19 Working Group, and has been endorsed through the official RDA process. Keywords: RDA; Recommendations; COVID-19. Language: English License: CC0 1.0 Universal (CC0 1.0) Public Domain Dedication RDA webpage: https://www.rd-alliance.org/group/rda-covid19-rda-covid19-omics-rda-covid19- epidemiology-rda-covid19-clinical-rda-covid19-1 Related resources: - RDA COVID-19 Guidelines and Recommendations – preliminary version, https://doi.org/10.15497/RDA00046 - Data Sharing in Epidemiology, https://doi.org/10.15497/RDA00049 - RDA COVID-19 Zotero Library, https://doi.org/10.15497/RDA00051 Citation and Download: RDA COVID-19 Working Group. Recommendations and Guidelines on data sharing. Research Data Alliance. 2020. DOI: https://doi.org/10.15497/RDA00052 RDA COVID-19 Recommendations and Guidelines on Data Sharing RDA Recommendation (FINAL Release) Produced by: RDA COVID-19 Working Group, 2020 Document Metadata Identifier DOI: https://doi.org/10.15497/rda00052 Citation To cite this document please use: RDA COVID-19 Working Group. Recommendations and Guidelines on data sharing. Research Data Alliance. 2020. DOI: https://doi.org/10.15497/rda00052 Title RDA COVID-19; Recommendations and Guidelines on Data Sharing, Final release 30 June 2020 Description This is the final version of the Recommendations and Guidelines -

HMMER User's Guide
HMMER User's Guide Biological sequence analysis using pro®le hidden Markov models http://hmmer.wustl.edu/ Version 2.1.1; December 1998 Sean Eddy Dept. of Genetics, Washington University School of Medicine 4566 Scott Ave., St. Louis, MO 63110, USA [email protected] With contributions by Ewan Birney ([email protected]) Copyright (C) 1992-1998, Washington University in St. Louis. Permission is granted to make and distribute verbatim copies of this manual provided the copyright notice and this permission notice are retained on all copies. The HMMER software package is a copyrighted work that may be freely distributed and modi®ed under the terms of the GNU General Public License as published by the Free Software Foundation; either version 2 of the License, or (at your option) any later version. Some versions of HMMER may have been obtained under specialized commercial licenses from Washington University; for details, see the ®les COPYING and LICENSE that came with your copy of the HMMER software. This program is distributed in the hope that it will be useful, but WITHOUT ANY WARRANTY; without even the implied warranty of MERCHANTABILITY or FITNESS FOR A PARTICULAR PURPOSE. See the Appendix for a copy of the full text of the GNU General Public License. 1 Contents 1 Tutorial 5 1.1 The programs in HMMER . 5 1.2 Files used in the tutorial . 6 1.3 Searching a sequence database with a single pro®le HMM . 6 HMM construction with hmmbuild . 7 HMM calibration with hmmcalibrate . 7 Sequence database search with hmmsearch . 8 Searching major databases like NR or SWISSPROT . -

Apply Parallel Bioinformatics Applications on Linux PC Clusters
Tunghai Science Vol. : 125−141 125 July, 2003 Apply Parallel Bioinformatics Applications on Linux PC Clusters Yu-Lun Kuo and Chao-Tung Yang* Abstract In addition to the traditional massively parallel computers, distributed workstation clusters now play an important role in scientific computing perhaps due to the advent of commodity high performance processors, low-latency/high-band width networks and powerful development tools. As we know, bioinformatics tools can speed up the analysis of large-scale sequence data, especially about sequence alignment. To fully utilize the relatively inexpensive CPU cycles available to today’s scientists, a PC cluster consists of one master node and seven slave nodes (16 processors totally), is proposed and built for bioinformatics applications. We use the mpiBLAST and HMMer on parallel computer to speed up the process for sequence alignment. The mpiBLAST software uses a message-passing library called MPI (Message Passing Interface) and the HMMer software uses a software package called PVM (Parallel Virtual Machine), respectively. The system architecture and performances of the cluster are also presented in this paper. Keywords: Parallel computing, Bioinformatics, BLAST, HMMer, PC Clusters, Speedup. 1. Introduction Extraordinary technological improvements over the past few years in areas such as microprocessors, memory, buses, networks, and software have made it possible to assemble groups of inexpensive personal computers and/or workstations into a cost effective system that functions in concert and posses tremendous processing power. Cluster computing is not new, but in company with other technical capabilities, particularly in the area of networking, this class of machines is becoming a high-performance platform for parallel and distributed applications [1, 2, 11, 12, 13, 14, 15, 16, 17]. -

HMMER User's Guide
HMMER User’s Guide Biological sequence analysis using profile hidden Markov models http://hmmer.org/ Version 3.0rc1; February 2010 Sean R. Eddy for the HMMER Development Team Janelia Farm Research Campus 19700 Helix Drive Ashburn VA 20147 USA http://eddylab.org/ Copyright (C) 2010 Howard Hughes Medical Institute. Permission is granted to make and distribute verbatim copies of this manual provided the copyright notice and this permission notice are retained on all copies. HMMER is licensed and freely distributed under the GNU General Public License version 3 (GPLv3). For a copy of the License, see http://www.gnu.org/licenses/. HMMER is a trademark of the Howard Hughes Medical Institute. 1 Contents 1 Introduction 5 How to avoid reading this manual . 5 How to avoid using this software (links to similar software) . 5 What profile HMMs are . 5 Applications of profile HMMs . 6 Design goals of HMMER3 . 7 What’s still missing in HMMER3 . 8 How to learn more about profile HMMs . 9 2 Installation 10 Quick installation instructions . 10 System requirements . 10 Multithreaded parallelization for multicores is the default . 11 MPI parallelization for clusters is optional . 11 Using build directories . 12 Makefile targets . 12 3 Tutorial 13 The programs in HMMER . 13 Files used in the tutorial . 13 Searching a sequence database with a single profile HMM . 14 Step 1: build a profile HMM with hmmbuild . 14 Step 2: search the sequence database with hmmsearch . 16 Searching a profile HMM database with a query sequence . 22 Step 1: create an HMM database flatfile . 22 Step 2: compress and index the flatfile with hmmpress . -
![Downloaded Were Considered to Be True Positive While Those from the from UCSC Databases on 14Th September 2011 [70,71]](https://docslib.b-cdn.net/cover/6028/downloaded-were-considered-to-be-true-positive-while-those-from-the-from-ucsc-databases-on-14th-september-2011-70-71-876028.webp)
Downloaded Were Considered to Be True Positive While Those from the from UCSC Databases on 14Th September 2011 [70,71]
Basu et al. BMC Bioinformatics 2013, 14(Suppl 7):S14 http://www.biomedcentral.com/1471-2105/14/S7/S14 RESEARCH Open Access Examples of sequence conservation analyses capture a subset of mouse long non-coding RNAs sharing homology with fish conserved genomic elements Swaraj Basu1, Ferenc Müller2, Remo Sanges1* From Ninth Annual Meeting of the Italian Society of Bioinformatics (BITS) Catania, Sicily. 2-4 May 2012 Abstract Background: Long non-coding RNAs (lncRNA) are a major class of non-coding RNAs. They are involved in diverse intra-cellular mechanisms like molecular scaffolding, splicing and DNA methylation. Through these mechanisms they are reported to play a role in cellular differentiation and development. They show an enriched expression in the brain where they are implicated in maintaining cellular identity, homeostasis, stress responses and plasticity. Low sequence conservation and lack of functional annotations make it difficult to identify homologs of mammalian lncRNAs in other vertebrates. A computational evaluation of the lncRNAs through systematic conservation analyses of both sequences as well as their genomic architecture is required. Results: Our results show that a subset of mouse candidate lncRNAs could be distinguished from random sequences based on their alignment with zebrafish phastCons elements. Using ROC analyses we were able to define a measure to select significantly conserved lncRNAs. Indeed, starting from ~2,800 mouse lncRNAs we could predict that between 4 and 11% present conserved sequence fragments in fish genomes. Gene ontology (GO) enrichment analyses of protein coding genes, proximal to the region of conservation, in both organisms highlighted similar GO classes like regulation of transcription and central nervous system development. -

Biological Sequence Analysis Probabilistic Models of Proteins and Nucleic Acids
This page intentionally left blank Biological sequence analysis Probabilistic models of proteins and nucleic acids The face of biology has been changed by the emergence of modern molecular genetics. Among the most exciting advances are large-scale DNA sequencing efforts such as the Human Genome Project which are producing an immense amount of data. The need to understand the data is becoming ever more pressing. Demands for sophisticated analyses of biological sequences are driving forward the newly-created and explosively expanding research area of computational molecular biology, or bioinformatics. Many of the most powerful sequence analysis methods are now based on principles of probabilistic modelling. Examples of such methods include the use of probabilistically derived score matrices to determine the significance of sequence alignments, the use of hidden Markov models as the basis for profile searches to identify distant members of sequence families, and the inference of phylogenetic trees using maximum likelihood approaches. This book provides the first unified, up-to-date, and tutorial-level overview of sequence analysis methods, with particular emphasis on probabilistic modelling. Pairwise alignment, hidden Markov models, multiple alignment, profile searches, RNA secondary structure analysis, and phylogenetic inference are treated at length. Written by an interdisciplinary team of authors, the book is accessible to molecular biologists, computer scientists and mathematicians with no formal knowledge of each others’ fields. It presents the state-of-the-art in this important, new and rapidly developing discipline. Richard Durbin is Head of the Informatics Division at the Sanger Centre in Cambridge, England. Sean Eddy is Assistant Professor at Washington University’s School of Medicine and also one of the Principle Investigators at the Washington University Genome Sequencing Center. -

Genomic and Transcriptomic Surveys for the Study of Ncrnas with a Focus on Tropical Parasites
PhD Thesis PROGRAMA DE PÓS-GRADUAÇÃO EM BIOINFORMÁTICA UNIVERSIDADE FEDERAL DE MINAS GERAIS Genomic and transcriptomic surveys for the study of ncRNAs with a focus on tropical parasites Mainá Bitar Belo Horizonte February 2015 Universidade Federal de Minas Gerais PhD Thesis PROGRAMA DE PÓS-GRADUAÇÃO EM BIOINFORMÁTICA Genomic and transcriptomic surveys for the study of ncRNAs with a focus on tropical parasites PhD candidate: Mainá Bitar Advisor: Glória Regina Franco Co-advisor: Martin Alexander Smith Mainá Bitar Lourenço Genomic and transcriptomic surveys for the study of ncRNAs with a focus on tropical parasites Versão final Tese apresentada ao Programa Interunidades de Pós-Graduação em Bioinformática do Instituto de Ciências Biológicas da Universidade Federal de Minas Gerais como requisito parcial para a obtenção do título de Doutor em Bioinformática. Orientador: Profa. Dra. Glória Regina Franco BELO HORIZONTE 2015 043 Bitar, Mainá. Genomic and transcriptomic surveys for the study of ncRNAs with a focus on tropical parasites [manuscrito] / Mainá Bitar. – 2015. 134 f. : il. ; 29,5 cm. Orientador: Glória Regina Franco. Coorientador: Martin Alexander Smith. Tese (doutorado) – Universidade Federal de Minas Gerais, Instituto de Ciências Biológicas. Programa de Pós-Graduação em Bioinformática. 1. Bioinformática - Teses. 2. Trypanosoma cruzi. 3. Schistosoma mansoni. 4. Genômica. 5. Transcriptoma. 6. Trans-Splicing. I. Franco, Glória Regina. II. Smith, Martin Alexander. III. Universidade Federal de Minas Gerais. Instituto de Ciências Biológicas. IV. Título. CDU: 573:004 Ficha catalográfica elaborada por Fabiane C. M. Reis – CRB 6/2680 Esta tese é dedicada à minha mãe, que me deu a liberdade para sonhar e a força para viver a realidade. -

Software List for Biology, Bioinformatics and Biostatistics CCT
Software List for biology, bioinformatics and biostatistics v CCT - Delta Software Version Application short read assembler and it works on both small and large (mammalian size) ALLPATHS-LG 52488 genomes provides a fast, flexible C++ API & toolkit for reading, writing, and manipulating BAMtools 2.4.0 BAM files a high level of alignment fidelity and is comparable to other mainstream Barracuda 0.7.107b alignment programs allows one to intersect, merge, count, complement, and shuffle genomic bedtools 2.25.0 intervals from multiple files Bfast 0.7.0a universal DNA sequence aligner tool analysis and comprehension of high-throughput genomic data using the R Bioconductor 3.2 statistical programming BioPython 1.66 tools for biological computation written in Python a fast approach to detecting gene-gene interactions in genome-wide case- Boost 1.54.0 control studies short read aligner geared toward quickly aligning large sets of short DNA Bowtie 1.1.2 sequences to large genomes Bowtie2 2.2.6 Bowtie + fully supports gapped alignment with affine gap penalties BWA 0.7.12 mapping low-divergent sequences against a large reference genome ClustalW 2.1 multiple sequence alignment program to align DNA and protein sequences assembles transcripts, estimates their abundances for differential expression Cufflinks 2.2.1 and regulation in RNA-Seq samples EBSEQ (R) 1.10.0 identifying genes and isoforms differentially expressed EMBOSS 6.5.7 a comprehensive set of sequence analysis programs FASTA 36.3.8b a DNA and protein sequence alignment software package FastQC -

On the Necessity of Dissecting Sequence Similarity Scores Into
Wong et al. BMC Bioinformatics 2014, 15:166 http://www.biomedcentral.com/1471-2105/15/166 METHODOLOGY ARTICLE Open Access On the necessity of dissecting sequence similarity scores into segment-specific contributions for inferring protein homology, function prediction and annotation Wing-Cheong Wong1*, Sebastian Maurer-Stroh1,2, Birgit Eisenhaber1 and Frank Eisenhaber1,3,4* Abstract Background: Protein sequence similarities to any types of non-globular segments (coiled coils, low complexity regions, transmembrane regions, long loops, etc. where either positional sequence conservation is the result of a very simple, physically induced pattern or rather integral sequence properties are critical) are pertinent sources for mistaken homologies. Regretfully, these considerations regularly escape attention in large-scale annotation studies since, often, there is no substitute to manual handling of these cases. Quantitative criteria are required to suppress events of function annotation transfer as a result of false homology assignments. Results: The sequence homology concept is based on the similarity comparison between the structural elements, the basic building blocks for conferring the overall fold of a protein. We propose to dissect the total similarity score into fold-critical and other, remaining contributions and suggest that, for a valid homology statement, the fold-relevant score contribution should at least be significant on its own. As part of the article, we provide the DissectHMMER software program for dissecting HMMER2/3 scores into segment-specific contributions. We show that DissectHMMER reproduces HMMER2/3 scores with sufficient accuracy and that it is useful in automated decisions about homology for instructive sequence examples. To generalize the dissection concept for cases without 3D structural information, we find that a dissection based on alignment quality is an appropriate surrogate. -
PTIR: Predicted Tomato Interactome Resource
www.nature.com/scientificreports OPEN PTIR: Predicted Tomato Interactome Resource Junyang Yue1,*, Wei Xu1,*, Rongjun Ban2,*, Shengxiong Huang1, Min Miao1, Xiaofeng Tang1, Guoqing Liu1 & Yongsheng Liu1,3 Received: 15 October 2015 Protein-protein interactions (PPIs) are involved in almost all biological processes and form the basis Accepted: 08 April 2016 of the entire interactomics systems of living organisms. Identification and characterization of these Published: 28 April 2016 interactions are fundamental to elucidating the molecular mechanisms of signal transduction and metabolic pathways at both the cellular and systemic levels. Although a number of experimental and computational studies have been performed on model organisms, the studies exploring and investigating PPIs in tomatoes remain lacking. Here, we developed a Predicted Tomato Interactome Resource (PTIR), based on experimentally determined orthologous interactions in six model organisms. The reliability of individual PPIs was also evaluated by shared gene ontology (GO) terms, co-evolution, co-expression, co-localization and available domain-domain interactions (DDIs). Currently, the PTIR covers 357,946 non-redundant PPIs among 10,626 proteins, including 12,291 high-confidence, 226,553 medium-confidence, and 119,102 low-confidence interactions. These interactions are expected to cover 30.6% of the entire tomato proteome and possess a reasonable distribution. In addition, ten randomly selected PPIs were verified using yeast two-hybrid (Y2H) screening or a bimolecular fluorescence complementation (BiFC) assay. The PTIR was constructed and implemented as a dedicated database and is available at http://bdg.hfut.edu.cn/ptir/index.html without registration. The increasing number of complete genome sequences has revealed the entire structure and composition of proteins, based mainly on theoretical predictions utilizing their corresponding DNA sequences. -

Download PDF of This Story
B NY RA DY BARRETT ILLUSTRATION BY MIKE PERRY TE H NEW JANELIA COMPUTING CLUSTER PUTS A PREMIUM ON EXPANDABILITY AND SPEED. ple—it’s pretty obvious to anyone which words are basically the same. That would be like two genes from humans and apes.” But in organisms that are more diver- gent, Eddy needs to understand how DNA sequences tend to change over time. “And it becomes a difficult specialty, with seri- ous statistical analysis,” he says. From a computational standpoint, that means churning through a lot of opera- tions. Comparing two typical-sized protein sequences, to take a simple example, would require a whopping 10200 opera- Computational biologists have a need for to help investigators conduct genome tions. Classic algorithms, available since speed. The computing cluster at HHMI’s searches and catalog the inner workings the 1960s, can trim that search to 160,000 Janelia Farm Research Campus delivers and structures of the brain. computations—a task that would take the performance they require—at a mind- only a millisecond or so on any modern boggling 36 trillion operations per second. F ASTER Answers processor. But in the genome business, In the course of their work, Janelia A group leader at Janelia Farm, Eddy deals people routinely do enormous numbers researchers generate millions of digitized in the realm of millions of computations of these sequence comparisons—trillions images and gigabytes of data files, and they daily as he compares sequences of DNA. and trillions of them. These “routine” cal- run algorithms daily that demand robust He is a rare breed, both biologist and code culations could take years if they had to be computational horsepower. -
Scaling HMMER Performance on Multicore Architectures
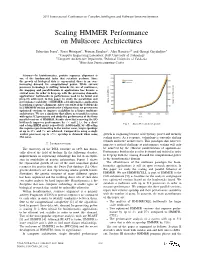
2011 International Conference on Complex, Intelligent, and Software Intensive Systems Scaling HMMER Performance on Multicore Architectures Sebastian Isaza∗, Ernst Houtgast∗, Friman Sanchezy, Alex Ramirezyz and Georgi Gaydadjiev∗ ∗Computer Engineering Laboratory, Delft University of Technology yComputer Architecture Department, Technical University of Catalonia zBarcelona Supercomputing Center Abstract—In bioinformatics, protein sequence alignment is one of the fundamental tasks that scientists perform. Since the growth of biological data is exponential, there is an ever- increasing demand for computational power. While current processor technology is shifting towards the use of multicores, the mapping and parallelization of applications has become a critical issue. In order to keep up with the processing demands, applications’ bottlenecks to performance need to be found and properly addressed. In this paper we study the parallelism and performance scalability of HMMER, a bioinformatics application to perform sequence alignment. After our study of the bottlenecks in a HMMER version ported to the Cell processor, we present two optimized versions to improve scalability in a larger multicore architecture. We use a simulator that allows us to model a system with up to 512 processors and study the performance of the three parallel versions of HMMER. Results show that removing the I/O bottleneck improves performance by 3× and 2:4× for a short Fig. 1. Swiss-Prot database growth. and a long HMM query respectively. Additionally, by offloading the sequence pre-formatting to the worker cores, larger speedups of up to 27× and 7× are achieved. Compared to using a single worker processor, up to 156× speedup is obtained when using growth is stagnating because of frequency, power and memory 256 cores.