Full-Text.Pdf
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Review the Significance of Interleukin-6
Review The significance of interleukin-6 and C-reactive protein in systemic sclerosis: a systematic literature review C. Muangchan1,2,3, J.E. Pope2 1Research Fellow, Rheumatology; ABSTRACT Introduction 2Schulich School of Medicine & Dentistry, Objectives. Interleukin-6 (IL-6) may Systemic sclerosis (SSc) or scleroder- Western University of Canada (formerly play a role in the pathogenesis of SSc. ma is a systemic autoimmune rheumat- University of Western Ontario), St Joseph C-reactive protein (CRP), an acute ic disease characterised by autoimmun- Health Care, London, ON; 3Division of Rheumatology, Department of phase reactant induced by IL-6, may be ity; fibrosis and dysfunction in vascular Medicine, Faculty of Medicine, Mahidol a prognostic marker in SSc. The goal of regulatory mechanisms highlighted by University, Siriraj Hospital, Bangkok, this systematic review was to address vasculopathy of microcirculation (1). Thailand. the significance and clinical applica- SSc has increased extracellular matrix Chayawee Muangchan, Research Fellow tion of IL-6 and CRP in systemic scle- protein deposition due to increased fi- Janet Elizabeth Pope, MD rosis (SSc). broblast biosynthetic activity (2). SSc is Please address correspondence Methods. A literature search was con- rare and has a female predisposition (3, and reprint requests to: ducted to identify English-language 4). It is classified into diffuse cutane- Dr Janet Pope, original articles within PubMed, Sco- ous SSc (dcSSc) and limited cutaneous St. Joseph’s Health Care, London, SSc (lcSSc) subsets according to extent 268 Grosvenor St., pus, and Medline database from incep- London N6A 4V2, ON, Canada. tion to May 30, 2013 using keywords of cutaneous involvement (5). -

Chronic Mtor Activation Induces a Degradative Smooth Muscle Cell Phenotype
The Journal of Clinical Investigation RESEARCH ARTICLE Chronic mTOR activation induces a degradative smooth muscle cell phenotype Guangxin Li,1,2 Mo Wang,1 Alexander W. Caulk,3 Nicholas A. Cilfone,4 Sharvari Gujja,4 Lingfeng Qin,1 Pei-Yu Chen,5 Zehua Chen,4 Sameh Yousef,1 Yang Jiao,1 Changshun He,1 Bo Jiang,1 Arina Korneva,3 Matthew R. Bersi,3 Guilin Wang,6 Xinran Liu,7,8 Sameet Mehta,9 Arnar Geirsson,1,10 Jeffrey R. Gulcher,4 Thomas W. Chittenden,4 Michael Simons,5,10 Jay D. Humphrey,3,10 and George Tellides1,10,11 1Department of Surgery, Yale School of Medicine, New Haven, Connecticut, USA. 2Department of Breast and Thyroid Surgery, Peking University Shenzhen Hospital, Shenzhen, Guangdong Province, China. 3Department of Biomedical Engineering, Yale School of Engineering and Applied Science, New Haven, Connecticut, USA. 4Computational Statistics and Bioinformatics Group, Advanced Artificial Intelligence Research Laboratory, WuXi NextCODE, Cambridge, Massachusetts, USA. 5Internal Medicine, 6Molecular Biophysics and Biochemistry, and 7Cell Biology, Yale School of Medicine, New Haven, Connecticut, USA. 8Center for Cellular and Molecular Imaging, EM Core Facility, Yale School of Medicine, New Haven, Connecticut, USA. 9Genetics and 10Program in Vascular Biology and Therapeutics, Yale School of Medicine, New Haven, Connecticut, USA. 11Veterans Affairs Connecticut Healthcare System, West Haven, Connecticut, USA. Smooth muscle cell (SMC) proliferation has been thought to limit the progression of thoracic aortic aneurysm and dissection (TAAD) because loss of medial cells associates with advanced disease. We investigated effects of SMC proliferation in the aortic media by conditional disruption of Tsc1, which hyperactivates mTOR complex 1. -

Human and Mouse CD Marker Handbook Human and Mouse CD Marker Key Markers - Human Key Markers - Mouse
Welcome to More Choice CD Marker Handbook For more information, please visit: Human bdbiosciences.com/eu/go/humancdmarkers Mouse bdbiosciences.com/eu/go/mousecdmarkers Human and Mouse CD Marker Handbook Human and Mouse CD Marker Key Markers - Human Key Markers - Mouse CD3 CD3 CD (cluster of differentiation) molecules are cell surface markers T Cell CD4 CD4 useful for the identification and characterization of leukocytes. The CD CD8 CD8 nomenclature was developed and is maintained through the HLDA (Human Leukocyte Differentiation Antigens) workshop started in 1982. CD45R/B220 CD19 CD19 The goal is to provide standardization of monoclonal antibodies to B Cell CD20 CD22 (B cell activation marker) human antigens across laboratories. To characterize or “workshop” the antibodies, multiple laboratories carry out blind analyses of antibodies. These results independently validate antibody specificity. CD11c CD11c Dendritic Cell CD123 CD123 While the CD nomenclature has been developed for use with human antigens, it is applied to corresponding mouse antigens as well as antigens from other species. However, the mouse and other species NK Cell CD56 CD335 (NKp46) antibodies are not tested by HLDA. Human CD markers were reviewed by the HLDA. New CD markers Stem Cell/ CD34 CD34 were established at the HLDA9 meeting held in Barcelona in 2010. For Precursor hematopoetic stem cell only hematopoetic stem cell only additional information and CD markers please visit www.hcdm.org. Macrophage/ CD14 CD11b/ Mac-1 Monocyte CD33 Ly-71 (F4/80) CD66b Granulocyte CD66b Gr-1/Ly6G Ly6C CD41 CD41 CD61 (Integrin b3) CD61 Platelet CD9 CD62 CD62P (activated platelets) CD235a CD235a Erythrocyte Ter-119 CD146 MECA-32 CD106 CD146 Endothelial Cell CD31 CD62E (activated endothelial cells) Epithelial Cell CD236 CD326 (EPCAM1) For Research Use Only. -

Tools for Cell Therapy and Immunoregulation
RnDSy-lu-2945 Tools for Cell Therapy and Immunoregulation Target Cell TIM-4 SLAM/CD150 BTNL8 PD-L2/B7-DC B7-H1/PD-L1 (Human) Unknown PD-1 B7-1/CD80 TIM-1 SLAM/CD150 Receptor TIM Family SLAM Family Butyrophilins B7/CD28 Families T Cell Multiple Co-Signaling Molecules Co-stimulatory Co-inhibitory Ig Superfamily Regulate T Cell Activation Target Cell T Cell Target Cell T Cell B7-1/CD80 B7-H1/PD-L1 T cell activation requires two signals: 1) recognition of the antigenic peptide/ B7-1/CD80 B7-2/CD86 CTLA-4 major histocompatibility complex (MHC) by the T cell receptor (TCR) and 2) CD28 antigen-independent co-stimulation induced by interactions between B7-2/CD86 B7-H1/PD-L1 B7-1/CD80 co-signaling molecules expressed on target cells, such as antigen-presenting PD-L2/B7-DC PD-1 ICOS cells (APCs), and their T cell-expressed receptors. Engagement of the TCR in B7-H2/ICOS L 2Ig B7-H3 (Mouse) the absence of this second co-stimulatory signal typically results in T cell B7-H1/PD-L1 B7/CD28 Families 4Ig B7-H3 (Human) anergy or apoptosis. In addition, T cell activation can be negatively regulated Unknown Receptors by co-inhibitory molecules present on APCs. Therefore, integration of the 2Ig B7-H3 Unknown B7-H4 (Mouse) Receptors signals transduced by co-stimulatory and co-inhibitory molecules following TCR B7-H5 4Ig B7-H3 engagement directs the outcome and magnitude of a T cell response Unknown Ligand (Human) B7-H5 including the enhancement or suppression of T cell proliferation, B7-H7 Unknown Receptor differentiation, and/or cytokine secretion. -

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

A Single Gene Expressed in Fibroblastic Reticular Cells Predicts
bioRxiv preprint doi: https://doi.org/10.1101/2020.02.19.955666; this version posted February 23, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-ND 4.0 International license. A single gene expressed in fibroblastic reticular cells predicts response to cancer immunotherapy Daniele Biasci1,2, , James Thaventhiran1,2*, and Simon Tavaré2,3,* 1MRC Toxicology Unit, University of Cambridge, Robinson Way, Cambridge, CB2 0RE, United Kingdom 2Cancer Research UK Cambridge Institute, Robinson Way, Cambridge CB2 0RE, United Kingdom 3Herbert and Florence Irving Institute for Cancer Dynamics, Columbia University, Schermerhorn Hall, Suite 601, 1190 Amsterdam Ave, New York, NY 10027, United States 1 While the role of CD8+ T cells in mediating response to cancer 46 CD19, FCER2 (CD23), PAX5, BANK1, VPREB3, TCL1A, 2 immunotherapy is well established, the role of other cell types, 47 CLEC17A and FDCSP (Fig. 1A and Table S1). Literature 3 including B cells, remains more controversial. By conducting a 48 reports that these genes are predominantly expressed in B 4 large gene expression study of response to immune checkpoint 49 cells (12), with the exception of FDCSP, which was found to 5 blockade (ICB), here we show that pre-treatment expression of 50 be expressed follicular dendritic cells (FDCs) isolated from 6 B cell genes is associated with ICB response independently of 51 secondary lymphoid organs, but not in B cells (13, 14). In 7 CD8+ T cells. -

Single-Cell RNA Sequencing Demonstrates the Molecular and Cellular Reprogramming of Metastatic Lung Adenocarcinoma
ARTICLE https://doi.org/10.1038/s41467-020-16164-1 OPEN Single-cell RNA sequencing demonstrates the molecular and cellular reprogramming of metastatic lung adenocarcinoma Nayoung Kim 1,2,3,13, Hong Kwan Kim4,13, Kyungjong Lee 5,13, Yourae Hong 1,6, Jong Ho Cho4, Jung Won Choi7, Jung-Il Lee7, Yeon-Lim Suh8,BoMiKu9, Hye Hyeon Eum 1,2,3, Soyean Choi 1, Yoon-La Choi6,10,11, Je-Gun Joung1, Woong-Yang Park 1,2,6, Hyun Ae Jung12, Jong-Mu Sun12, Se-Hoon Lee12, ✉ ✉ Jin Seok Ahn12, Keunchil Park12, Myung-Ju Ahn 12 & Hae-Ock Lee 1,2,3,6 1234567890():,; Advanced metastatic cancer poses utmost clinical challenges and may present molecular and cellular features distinct from an early-stage cancer. Herein, we present single-cell tran- scriptome profiling of metastatic lung adenocarcinoma, the most prevalent histological lung cancer type diagnosed at stage IV in over 40% of all cases. From 208,506 cells populating the normal tissues or early to metastatic stage cancer in 44 patients, we identify a cancer cell subtype deviating from the normal differentiation trajectory and dominating the metastatic stage. In all stages, the stromal and immune cell dynamics reveal ontological and functional changes that create a pro-tumoral and immunosuppressive microenvironment. Normal resident myeloid cell populations are gradually replaced with monocyte-derived macrophages and dendritic cells, along with T-cell exhaustion. This extensive single-cell analysis enhances our understanding of molecular and cellular dynamics in metastatic lung cancer and reveals potential diagnostic and therapeutic targets in cancer-microenvironment interactions. 1 Samsung Genome Institute, Samsung Medical Center, Seoul 06351, Korea. -

Supplementary Table 1: Adhesion Genes Data Set
Supplementary Table 1: Adhesion genes data set PROBE Entrez Gene ID Celera Gene ID Gene_Symbol Gene_Name 160832 1 hCG201364.3 A1BG alpha-1-B glycoprotein 223658 1 hCG201364.3 A1BG alpha-1-B glycoprotein 212988 102 hCG40040.3 ADAM10 ADAM metallopeptidase domain 10 133411 4185 hCG28232.2 ADAM11 ADAM metallopeptidase domain 11 110695 8038 hCG40937.4 ADAM12 ADAM metallopeptidase domain 12 (meltrin alpha) 195222 8038 hCG40937.4 ADAM12 ADAM metallopeptidase domain 12 (meltrin alpha) 165344 8751 hCG20021.3 ADAM15 ADAM metallopeptidase domain 15 (metargidin) 189065 6868 null ADAM17 ADAM metallopeptidase domain 17 (tumor necrosis factor, alpha, converting enzyme) 108119 8728 hCG15398.4 ADAM19 ADAM metallopeptidase domain 19 (meltrin beta) 117763 8748 hCG20675.3 ADAM20 ADAM metallopeptidase domain 20 126448 8747 hCG1785634.2 ADAM21 ADAM metallopeptidase domain 21 208981 8747 hCG1785634.2|hCG2042897 ADAM21 ADAM metallopeptidase domain 21 180903 53616 hCG17212.4 ADAM22 ADAM metallopeptidase domain 22 177272 8745 hCG1811623.1 ADAM23 ADAM metallopeptidase domain 23 102384 10863 hCG1818505.1 ADAM28 ADAM metallopeptidase domain 28 119968 11086 hCG1786734.2 ADAM29 ADAM metallopeptidase domain 29 205542 11085 hCG1997196.1 ADAM30 ADAM metallopeptidase domain 30 148417 80332 hCG39255.4 ADAM33 ADAM metallopeptidase domain 33 140492 8756 hCG1789002.2 ADAM7 ADAM metallopeptidase domain 7 122603 101 hCG1816947.1 ADAM8 ADAM metallopeptidase domain 8 183965 8754 hCG1996391 ADAM9 ADAM metallopeptidase domain 9 (meltrin gamma) 129974 27299 hCG15447.3 ADAMDEC1 ADAM-like, -

The Regulation of Interleukin 7 Receptor Alpha Internalization, Recycling and Degradation by IL-7
Universidade de Lisboa Faculdade de Medicina Unidade de Biologia do Cancro, Instituto de Medicina Molecular The regulation of Interleukin 7 receptor alpha internalization, recycling and degradation by IL-7 - Possible implications in T-cell homeostasis, migration and leukaemogenesis - Catarina Martins de Oliveira Henriques Doutoramento em Ciências Biomédicas For the degree of Doctor of Philosophy 2009 Universidade de Lisboa Faculdade de Medicina Unidade de Biologia do Cancro, Instituto de Medicina Molecular The regulation of Interleukin 7 receptor alpha internalization, recycling and degradation by IL-7 - Possible implications in T-cell homeostasis, migration and leukaemogenesis - Catarina Martins de Oliveira Henriques (Recipient of a scholarship- SFRH7BD/21940/2005 from Fundação para a Ciência e Tecnologia) Tese orientada pelo Doutor João T. Barata, Prof Doutor.Gerard Graham e Prof. Doutora Leonor Parreira Doutoramento em Ciências Biomédicas, especialidade em Ciências Biopatológicas For the degree of Doctor of Philosophy 2009 As opiniões expressas são da exclusiva responsabilidade do seu autor A impressão desta dissertação foi aprovada pela Comissão Coordenadora do Conselho Científico da Faculdade de Medicina de Lisboa em reunião de 13 de Outubro de 2009. Para a Prof. Filomena Mota. Sem o seu apoio e inspiração há muitos anos atrás, eu não seria hoje uma bióloga nem esta tese teria alguma vez existido… Table of Contents Table of contents……………………………………………………….………………….................. i Aknowledgements……………………………………………………………………………………………. -

CD Markers Are Routinely Used for the Immunophenotyping of Cells
ptglab.com 1 CD MARKER ANTIBODIES www.ptglab.com Introduction The cluster of differentiation (abbreviated as CD) is a protocol used for the identification and investigation of cell surface molecules. So-called CD markers are routinely used for the immunophenotyping of cells. Despite this use, they are not limited to roles in the immune system and perform a variety of roles in cell differentiation, adhesion, migration, blood clotting, gamete fertilization, amino acid transport and apoptosis, among many others. As such, Proteintech’s mini catalog featuring its antibodies targeting CD markers is applicable to a wide range of research disciplines. PRODUCT FOCUS PECAM1 Platelet endothelial cell adhesion of blood vessels – making up a large portion molecule-1 (PECAM1), also known as cluster of its intracellular junctions. PECAM-1 is also CD Number of differentiation 31 (CD31), is a member of present on the surface of hematopoietic the immunoglobulin gene superfamily of cell cells and immune cells including platelets, CD31 adhesion molecules. It is highly expressed monocytes, neutrophils, natural killer cells, on the surface of the endothelium – the thin megakaryocytes and some types of T-cell. Catalog Number layer of endothelial cells lining the interior 11256-1-AP Type Rabbit Polyclonal Applications ELISA, FC, IF, IHC, IP, WB 16 Publications Immunohistochemical of paraffin-embedded Figure 1: Immunofluorescence staining human hepatocirrhosis using PECAM1, CD31 of PECAM1 (11256-1-AP), Alexa 488 goat antibody (11265-1-AP) at a dilution of 1:50 anti-rabbit (green), and smooth muscle KD/KO Validated (40x objective). alpha-actin (red), courtesy of Nicola Smart. PECAM1: Customer Testimonial Nicola Smart, a cardiovascular researcher “As you can see [the immunostaining] is and a group leader at the University of extremely clean and specific [and] displays Oxford, has said of the PECAM1 antibody strong intercellular junction expression, (11265-1-AP) that it “worked beautifully as expected for a cell adhesion molecule.” on every occasion I’ve tried it.” Proteintech thanks Dr. -
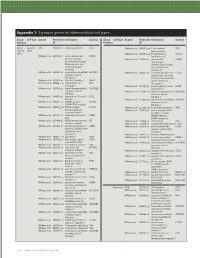
Technical Note, Appendix: an Analysis of Blood Processing Methods to Prepare Samples for Genechip® Expression Profiling (Pdf, 1
Appendix 1: Signature genes for different blood cell types. Blood Cell Type Source Probe Set Description Symbol Blood Cell Type Source Probe Set Description Symbol Fraction ID Fraction ID Mono- Lympho- GSK 203547_at CD4 antigen (p55) CD4 Whitney et al. 209813_x_at T cell receptor TRG nuclear cytes gamma locus cells Whitney et al. 209995_s_at T-cell leukemia/ TCL1A Whitney et al. 203104_at colony stimulating CSF1R lymphoma 1A factor 1 receptor, Whitney et al. 210164_at granzyme B GZMB formerly McDonough (granzyme 2, feline sarcoma viral cytotoxic T-lymphocyte- (v-fms) oncogene associated serine homolog esterase 1) Whitney et al. 203290_at major histocompatibility HLA-DQA1 Whitney et al. 210321_at similar to granzyme B CTLA1 complex, class II, (granzyme 2, cytotoxic DQ alpha 1 T-lymphocyte-associated Whitney et al. 203413_at NEL-like 2 (chicken) NELL2 serine esterase 1) Whitney et al. 203828_s_at natural killer cell NK4 (H. sapiens) transcript 4 Whitney et al. 212827_at immunoglobulin heavy IGHM Whitney et al. 203932_at major histocompatibility HLA-DMB constant mu complex, class II, Whitney et al. 212998_x_at major histocompatibility HLA-DQB1 DM beta complex, class II, Whitney et al. 204655_at chemokine (C-C motif) CCL5 DQ beta 1 ligand 5 Whitney et al. 212999_x_at major histocompatibility HLA-DQB Whitney et al. 204661_at CDW52 antigen CDW52 complex, class II, (CAMPATH-1 antigen) DQ beta 1 Whitney et al. 205049_s_at CD79A antigen CD79A Whitney et al. 213193_x_at T cell receptor beta locus TRB (immunoglobulin- Whitney et al. 213425_at Homo sapiens cDNA associated alpha) FLJ11441 fis, clone Whitney et al. 205291_at interleukin 2 receptor, IL2RB HEMBA1001323, beta mRNA sequence Whitney et al. -

Lysosomal Membrane Glycoproteins Bind Cholesterol and Contribute to Lysosomal Cholesterol Export
1 Lysosomal membrane glycoproteins bind cholesterol and 2 contribute to lysosomal cholesterol export 3 4 Jian Li and Suzanne R. Pfeffer* 5 Department of Biochemistry, Stanford University School of Medicine 6 Stanford, CA USA 94305-5307 7 8 *Correspondence to: [email protected]. 9 10 Abstract: LAMP1 and LAMP2 proteins are highly abundant, ubiquitous, mammalian proteins 11 that line the lysosome limiting membrane, and protect it from lysosomal hydrolase action. 12 LAMP2 deficiency causes Danon’s disease, an X-linked hypertrophic cardiomyopathy. LAMP2 13 is needed for chaperone-mediated autophagy, and its expression improves tissue function in 14 models of aging. We show here that LAMP1 and LAMP2 bind cholesterol in a manner that 15 buries the cholesterol 3β-hydroxyl group; they also bind tightly to NPC1 and NPC2 proteins that 16 export cholesterol from lysosomes. Quantitation of cellular LAMP2 and NPC1 protein levels 17 suggest that LAMP proteins represent a significant cholesterol binding site at the lysosome 18 limiting membrane, and may signal cholesterol availability. Functional rescue experiments show 19 that the ability of LAMP2 to facilitate cholesterol export from lysosomes relies on its ability to 20 bind cholesterol directly. 21 22 23 Introduction 24 Eukaryotic lysosomes are acidic, membrane-bound organelles that contain proteases, lipases and 25 nucleases and degrade cellular components to regenerate catabolic precursors for cellular use (1- 26 3). Lysosomes are crucial for the degradation of substrates from the cytoplasm, as well as 27 membrane bound compartments derived from the secretory, endocytic, autophagic and 28 phagocytic pathways. The limiting membrane of lysosomes is lined with so-called lysosomal 29 membrane glycoproteins (LAMPs) that are comprised of a short cytoplasmic domain, a single 30 transmembrane span, and a highly, N- and O-glycosylated lumenal domain (4-6).