Regulation of Monocyte/Macrophage Activation by Leukocyte Immunoglobulin-Like Receptor B4
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Imm Catalog.Pdf
$ Gene Symbol A B 3 C 4 D 9 E 10 F 11 G 12 H 13 I 14 J. K 17 L 18 M 19 N 20 O. P 22 R 26 S 27 T 30 U 32 V. W. X. Y. Z 33 A ® ® Gene Symbol Gene ID Antibody Monoclonal Antibody Polyclonal MaxPab Full-length Protein Partial-length Protein Antibody Pair KIt siRNA/Chimera Gene Symbol Gene ID Antibody Monoclonal Antibody Polyclonal MaxPab Full-length Protein Partial-length Protein Antibody Pair KIt siRNA/Chimera A1CF 29974 ● ● ADAMTS13 11093 ● ● ● ● ● A2M 2 ● ● ● ● ● ● ADAMTS20 80070 ● AACS 65985 ● ● ● ADAMTS5 11096 ● ● ● AANAT 15 ● ● ADAMTS8 11095 ● ● ● ● AATF 26574 ● ● ● ● ● ADAMTSL2 9719 ● AATK 9625 ● ● ● ● ADAMTSL4 54507 ● ● ABCA1 19 ● ● ● ● ● ADAR 103 ● ● ABCA5 23461 ● ● ADARB1 104 ● ● ● ● ABCA7 10347 ● ADARB2 105 ● ABCB9 23457 ● ● ● ● ● ADAT1 23536 ● ● ABCC4 10257 ● ● ● ● ADAT2 134637 ● ● ABCC5 10057 ● ● ● ● ● ADAT3 113179 ● ● ● ABCC8 6833 ● ● ● ● ADCY10 55811 ● ● ABCD2 225 ● ADD1 118 ● ● ● ● ● ● ABCD4 5826 ● ● ● ADD3 120 ● ● ● ABCG1 9619 ● ● ● ● ● ADH5 128 ● ● ● ● ● ● ABL1 25 ● ● ADIPOQ 9370 ● ● ● ● ● ABL2 27 ● ● ● ● ● ADK 132 ● ● ● ● ● ABO 28 ● ● ADM 133 ● ● ● ABP1 26 ● ● ● ● ● ADNP 23394 ● ● ● ● ABR 29 ● ● ● ● ● ADORA1 134 ● ● ACAA2 10449 ● ● ● ● ADORA2A 135 ● ● ● ● ● ● ● ACAN 176 ● ● ● ● ● ● ADORA2B 136 ● ● ACE 1636 ● ● ● ● ADRA1A 148 ● ● ● ● ACE2 59272 ● ● ADRA1B 147 ● ● ACER2 340485 ● ADRA2A 150 ● ● ACHE 43 ● ● ● ● ● ● ADRB1 153 ● ● ACIN1 22985 ● ● ● ADRB2 154 ● ● ● ● ● ACOX1 51 ● ● ● ● ● ADRB3 155 ● ● ● ● ACP5 54 ● ● ● ● ● ● ● ADRBK1 156 ● ● ● ● ACSF2 80221 ● ● ADRM1 11047 ● ● ● ● ACSF3 197322 ● ● AEBP1 165 ● ● ● ● ACSL4 2182 ● -

Human and Mouse CD Marker Handbook Human and Mouse CD Marker Key Markers - Human Key Markers - Mouse
Welcome to More Choice CD Marker Handbook For more information, please visit: Human bdbiosciences.com/eu/go/humancdmarkers Mouse bdbiosciences.com/eu/go/mousecdmarkers Human and Mouse CD Marker Handbook Human and Mouse CD Marker Key Markers - Human Key Markers - Mouse CD3 CD3 CD (cluster of differentiation) molecules are cell surface markers T Cell CD4 CD4 useful for the identification and characterization of leukocytes. The CD CD8 CD8 nomenclature was developed and is maintained through the HLDA (Human Leukocyte Differentiation Antigens) workshop started in 1982. CD45R/B220 CD19 CD19 The goal is to provide standardization of monoclonal antibodies to B Cell CD20 CD22 (B cell activation marker) human antigens across laboratories. To characterize or “workshop” the antibodies, multiple laboratories carry out blind analyses of antibodies. These results independently validate antibody specificity. CD11c CD11c Dendritic Cell CD123 CD123 While the CD nomenclature has been developed for use with human antigens, it is applied to corresponding mouse antigens as well as antigens from other species. However, the mouse and other species NK Cell CD56 CD335 (NKp46) antibodies are not tested by HLDA. Human CD markers were reviewed by the HLDA. New CD markers Stem Cell/ CD34 CD34 were established at the HLDA9 meeting held in Barcelona in 2010. For Precursor hematopoetic stem cell only hematopoetic stem cell only additional information and CD markers please visit www.hcdm.org. Macrophage/ CD14 CD11b/ Mac-1 Monocyte CD33 Ly-71 (F4/80) CD66b Granulocyte CD66b Gr-1/Ly6G Ly6C CD41 CD41 CD61 (Integrin b3) CD61 Platelet CD9 CD62 CD62P (activated platelets) CD235a CD235a Erythrocyte Ter-119 CD146 MECA-32 CD106 CD146 Endothelial Cell CD31 CD62E (activated endothelial cells) Epithelial Cell CD236 CD326 (EPCAM1) For Research Use Only. -

Tools for Cell Therapy and Immunoregulation
RnDSy-lu-2945 Tools for Cell Therapy and Immunoregulation Target Cell TIM-4 SLAM/CD150 BTNL8 PD-L2/B7-DC B7-H1/PD-L1 (Human) Unknown PD-1 B7-1/CD80 TIM-1 SLAM/CD150 Receptor TIM Family SLAM Family Butyrophilins B7/CD28 Families T Cell Multiple Co-Signaling Molecules Co-stimulatory Co-inhibitory Ig Superfamily Regulate T Cell Activation Target Cell T Cell Target Cell T Cell B7-1/CD80 B7-H1/PD-L1 T cell activation requires two signals: 1) recognition of the antigenic peptide/ B7-1/CD80 B7-2/CD86 CTLA-4 major histocompatibility complex (MHC) by the T cell receptor (TCR) and 2) CD28 antigen-independent co-stimulation induced by interactions between B7-2/CD86 B7-H1/PD-L1 B7-1/CD80 co-signaling molecules expressed on target cells, such as antigen-presenting PD-L2/B7-DC PD-1 ICOS cells (APCs), and their T cell-expressed receptors. Engagement of the TCR in B7-H2/ICOS L 2Ig B7-H3 (Mouse) the absence of this second co-stimulatory signal typically results in T cell B7-H1/PD-L1 B7/CD28 Families 4Ig B7-H3 (Human) anergy or apoptosis. In addition, T cell activation can be negatively regulated Unknown Receptors by co-inhibitory molecules present on APCs. Therefore, integration of the 2Ig B7-H3 Unknown B7-H4 (Mouse) Receptors signals transduced by co-stimulatory and co-inhibitory molecules following TCR B7-H5 4Ig B7-H3 engagement directs the outcome and magnitude of a T cell response Unknown Ligand (Human) B7-H5 including the enhancement or suppression of T cell proliferation, B7-H7 Unknown Receptor differentiation, and/or cytokine secretion. -
![Anti-NCR1 Antibody [29A1.4] (FITC) (ARG23568)](https://docslib.b-cdn.net/cover/3482/anti-ncr1-antibody-29a1-4-fitc-arg23568-113482.webp)
Anti-NCR1 Antibody [29A1.4] (FITC) (ARG23568)
Product datasheet [email protected] ARG23568 Package: 50 μg anti-NCR1 antibody [29A1.4] (FITC) Store at: 4°C Summary Product Description FITC-conjugated Rat Monoclonal antibody [29A1.4] recognizes NCR1. Rat anti Mouse CD335 antibody, clone 29A1.4 recognizes natural killer cell p46-related protein (NKp46), otherwise known as CD335, which is uniquely expressed by resting and activated natural killer (NK) cells, but no expression of CD335 has been detected on B cells, T cells, monocytes or granulocytes. It is a major NK cell lysis receptor for autologous pathogen-infected and tumor target cells during natural cytotoxicity responses. Ligands of CD335 include viral hemagglutinins (HAs) and heparan sulfate proteoglycans (HSPGs) on the surface of tumor cells. Staining with Rat anti mouse CD335 (29A1.4) is not strain specific and the antibody has been used to stain C57BL/6, SJL, CBA/CA and BALB/c strains. Clone 29A1.4 also activates NK cells in vitro. It does not deplete NK cells in vivo. Tested Reactivity Ms Tested Application FACS Host Rat Clonality Monoclonal Clone 29A1.4 Isotype IgG2a Target Name NCR1 Antigen Species Mouse Immunogen NKP46-IgG1 Fc fusion protein. Conjugation FITC Alternate Names CD antigen CD335; Natural killer cell p46-related protein; Lymphocyte antigen 94 homolog; Natural cytotoxicity triggering receptor 1; hNKp46; NKP46; NKp46; NK cell-activating receptor; LY94; CD335; NK- p46 Application Instructions Application table Application Dilution FACS 1:2 - 1:5 Application Note FACS: Use 10 µl of the suggested working dilution to label 10^6 cells in 100 µl. * The dilutions indicate recommended starting dilutions and the optimal dilutions or concentrations should be determined by the scientist. -

Molecular Profile of Tumor-Specific CD8+ T Cell Hypofunction in a Transplantable Murine Cancer Model
Downloaded from http://www.jimmunol.org/ by guest on September 25, 2021 T + is online at: average * The Journal of Immunology , 34 of which you can access for free at: 2016; 197:1477-1488; Prepublished online 1 July from submission to initial decision 4 weeks from acceptance to publication 2016; doi: 10.4049/jimmunol.1600589 http://www.jimmunol.org/content/197/4/1477 Molecular Profile of Tumor-Specific CD8 Cell Hypofunction in a Transplantable Murine Cancer Model Katherine A. Waugh, Sonia M. Leach, Brandon L. Moore, Tullia C. Bruno, Jonathan D. Buhrman and Jill E. Slansky J Immunol cites 95 articles Submit online. Every submission reviewed by practicing scientists ? is published twice each month by Receive free email-alerts when new articles cite this article. Sign up at: http://jimmunol.org/alerts http://jimmunol.org/subscription Submit copyright permission requests at: http://www.aai.org/About/Publications/JI/copyright.html http://www.jimmunol.org/content/suppl/2016/07/01/jimmunol.160058 9.DCSupplemental This article http://www.jimmunol.org/content/197/4/1477.full#ref-list-1 Information about subscribing to The JI No Triage! Fast Publication! Rapid Reviews! 30 days* Why • • • Material References Permissions Email Alerts Subscription Supplementary The Journal of Immunology The American Association of Immunologists, Inc., 1451 Rockville Pike, Suite 650, Rockville, MD 20852 Copyright © 2016 by The American Association of Immunologists, Inc. All rights reserved. Print ISSN: 0022-1767 Online ISSN: 1550-6606. This information is current as of September 25, 2021. The Journal of Immunology Molecular Profile of Tumor-Specific CD8+ T Cell Hypofunction in a Transplantable Murine Cancer Model Katherine A. -

Expression Tissue
The Journal of Immunology Killer Cell Ig-Like Receptor and Leukocyte Ig-Like Receptor Transgenic Mice Exhibit Tissue- and Cell-Specific Transgene Expression1 Danny Belkin,* Michaela Torkar,* Chiwen Chang,* Roland Barten,* Mauro Tolaini,† Anja Haude,* Rachel Allen,* Michael J. Wilson,* Dimitris Kioussis,† and John Trowsdale2* To generate an experimental model for exploring the function, expression pattern, and developmental regulation of human Ig-like activating and inhibitory receptors, we have generated transgenic mice using two human genomic clones: 52N12 (a 150-Kb clone encompassing the leukocyte Ig-like receptor (LILR)B1 (ILT2), LILRB4 (ILT3), and LILRA1 (LIR6) genes) and 1060P11 (a 160-Kb clone that contains ten killer cell Ig-like receptor (KIR) genes). Both the KIR and LILR families are encoded within the leukocyte receptor complex, and are involved in immune modulation. We have also produced a novel mAb to LILRA1 to facilitate expression studies. The LILR transgenes were expressed in a similar, but not identical, pattern to that observed in humans: LILRB1 was expressed in B cells, most NK cells, and a small number of T cells; LILRB4 was expressed in a B cell subset; and LILRA1 was found on a ring of cells surrounding B cell areas on spleen sections, consistent with other data showing monocyte/macrophage expression. KIR transgenic mice showed KIR2DL2 expression on a subset of NK cells and T cells, similar to the pattern seen in humans, and expression of KIR2DL4, KIR3DS1, and KIR2DL5 by splenic NK cells. These observations indicate that linked regulatory elements within the genomic clones are sufficient to allow appropriate expression of KIRs in mice, and illustrate that the presence of the natural ligands for these receptors, in the form of human MHC class I proteins, is not necessary for the expression of the KIRs observed in these mice. -

ORIGINAL ARTICLE Flow Cytometric Protein Expression Profiling As a Systematic Approach for Developing Disease-Specific Assays
Leukemia (2006) 20, 2102–2110 & 2006 Nature Publishing Group All rights reserved 0887-6924/06 $30.00 www.nature.com/leu ORIGINAL ARTICLE Flow cytometric protein expression profiling as a systematic approach for developing disease-specific assays: identification of a chronic lymphocytic leukaemia-specific assay for use in rituximab-containing regimens AC Rawstron, R de Tute, AS Jack and P Hillmen Haematological Malignancy Diagnostic Service (HMDS), Leeds Teaching Hospitals, Leeds, UK Depletion of disease below the levels detected by sensitive sustained remissions only occur in patients achieving an MRD- minimal residual disease (MRD) assays is associated with negative complete response.12 Therefore MRD is increasingly prolonged survival in chronic lymphocytic leukaemia (CLL). being used as an end point for therapeutic trials, and several Flow cytometric MRD assays are now sufficiently sensitive and rapid to guide the duration of therapy in CLL, but generally rely studies are now using the assessment of MRD to define the on assessment of CD20 expression, which cannot be accurately duration of therapy. measured during and after therapeutic approaches containing Approaches using allele-specific oligonucleotide polymerase rituximab. The aim of this study was to use analytical software chain reaction (ASO-PCR) to the immunoglobulin gene of the developed for microarray analysis to provide a systematic B-CLL cell are generally accepted to show the highest sensitivity approach for MRD flow assay development. Samples from CLL for MRD detection. However, more recent four-colour ap- patients (n ¼ 49), normal controls (n ¼ 21) and other B-lympho- proaches show sensitivities nearing that of ASO-PCR6,11,13 with proliferative disorders (n ¼ 12) were assessed with a panel of 66 antibodies. -

Receptor Nkp46 Cells by the NK Β Murine Pancreatic Recognition And
Recognition and Killing of Human and Murine Pancreatic β Cells by the NK Receptor NKp46 This information is current as Chamutal Gur, Jonatan Enk, Sameer A. Kassem, Yaron of September 27, 2021. Suissa, Judith Magenheim, Miri Stolovich-Rain, Tomer Nir, Hagit Achdout, Benjamin Glaser, James Shapiro, Yaakov Naparstek, Angel Porgador, Yuval Dor and Ofer Mandelboim J Immunol 2011; 187:3096-3103; Prepublished online 17 Downloaded from August 2011; doi: 10.4049/jimmunol.1101269 http://www.jimmunol.org/content/187/6/3096 http://www.jimmunol.org/ Supplementary http://www.jimmunol.org/content/suppl/2011/08/18/jimmunol.110126 Material 9.DC1 References This article cites 29 articles, 9 of which you can access for free at: http://www.jimmunol.org/content/187/6/3096.full#ref-list-1 Why The JI? Submit online. by guest on September 27, 2021 • Rapid Reviews! 30 days* from submission to initial decision • No Triage! Every submission reviewed by practicing scientists • Fast Publication! 4 weeks from acceptance to publication *average Subscription Information about subscribing to The Journal of Immunology is online at: http://jimmunol.org/subscription Permissions Submit copyright permission requests at: http://www.aai.org/About/Publications/JI/copyright.html Email Alerts Receive free email-alerts when new articles cite this article. Sign up at: http://jimmunol.org/alerts The Journal of Immunology is published twice each month by The American Association of Immunologists, Inc., 1451 Rockville Pike, Suite 650, Rockville, MD 20852 Copyright © 2011 by The American Association of Immunologists, Inc. All rights reserved. Print ISSN: 0022-1767 Online ISSN: 1550-6606. The Journal of Immunology Recognition and Killing of Human and Murine Pancreatic b Cells by the NK Receptor NKp46 Chamutal Gur,*,†,1 Jonatan Enk,*,1 Sameer A. -

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -
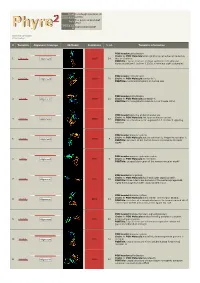
Phyre 2 Results for A1YIY0
Email [email protected] Description A1YIY0 Tue Jul 30 13:19:15 BST Date 2013 Unique Job 1035bc4b501530df ID Detailed template information # Template Alignment Coverage 3D Model Confidence % i.d. Template Information PDB header:cell adhesion Chain: A: PDB Molecule:down syndrome cell adhesion molecule 1 c3dmkA_ Alignment 100.0 14 (dscam) isoform PDBTitle: crystal structure of down syndrome cell adhesion molecule (dscam)2 isoform 1.30.30, n-terminal eight ig domains PDB header:cell adhesion 2 c2om5A_ Alignment 100.0 20 Chain: A: PDB Molecule:contactin 2; PDBTitle: n-terminal fragment of human tax1 PDB header:cell adhesion 3 c3jxaA_ Alignment 100.0 21 Chain: A: PDB Molecule:contactin 4; PDBTitle: immunoglobulin domains 1-4 of mouse cntn4 PDB header:signaling protein/transferase Chain: A: PDB Molecule:tek tyrosine kinase variant; 4 c4k0vA_ 100.0 12 Alignment PDBTitle: structural basis for angiopoietin-1 mediated signaling initiation PDB header:immune system Chain: A: PDB Molecule:natural cytotoxicity triggering receptor 1; 5 c1p6fA_ 100.0 9 Alignment PDBTitle: structure of the human natural cytotoxicity receptor nkp46 PDB header:immune system/receptor 6 c1ollA_ Alignment 99.9 9 Chain: A: PDB Molecule:nk receptor; PDBTitle: extracellular region of the human receptor nkp46 PDB header:viral protein Chain: A: PDB Molecule:hoc head outer capsid protein; 7 c3shsA_ 99.9 18 Alignment PDBTitle: three n-terminal domains of the bacteriophage rb49 highly immunogenic2 outer capsid protein (hoc) PDB header:immune system Chain: E: PDB Molecule:natural -

CD226 T Cells Expressing the Receptors TIGIT and Divergent Phenotypes of Human Regulatory
The Journal of Immunology Divergent Phenotypes of Human Regulatory T Cells Expressing the Receptors TIGIT and CD226 Christopher A. Fuhrman,*,1 Wen-I Yeh,*,1 Howard R. Seay,* Priya Saikumar Lakshmi,* Gaurav Chopra,† Lin Zhang,* Daniel J. Perry,* Stephanie A. McClymont,† Mahesh Yadav,† Maria-Cecilia Lopez,‡ Henry V. Baker,‡ Ying Zhang,x Yizheng Li,{ Maryann Whitley,{ David von Schack,x Mark A. Atkinson,* Jeffrey A. Bluestone,‡ and Todd M. Brusko* Regulatory T cells (Tregs) play a central role in counteracting inflammation and autoimmunity. A more complete understanding of cellular heterogeneity and the potential for lineage plasticity in human Treg subsets may identify markers of disease pathogenesis and facilitate the development of optimized cellular therapeutics. To better elucidate human Treg subsets, we conducted direct transcriptional profiling of CD4+FOXP3+Helios+ thymic-derived Tregs and CD4+FOXP3+Helios2 T cells, followed by comparison with CD4+FOXP32Helios2 T conventional cells. These analyses revealed that the coinhibitory receptor T cell Ig and ITIM domain (TIGIT) was highly expressed on thymic-derived Tregs. TIGIT and the costimulatory factor CD226 bind the common ligand CD155. Thus, we analyzed the cellular distribution and suppressive activity of isolated subsets of CD4+CD25+CD127lo/2 T cells expressing CD226 and/or TIGIT. We observed TIGIT is highly expressed and upregulated on Tregs after activation and in vitro expansion, and is associated with lineage stability and suppressive capacity. Conversely, the CD226+TIGIT2 population was associated with reduced Treg purity and suppressive capacity after expansion, along with a marked increase in IL-10 and effector cytokine production. These studies provide additional markers to delineate functionally distinct Treg subsets that may help direct cellular therapies and provide important phenotypic markers for assessing the role of Tregs in health and disease. -

ANGPTL2/LILRB2 Signaling Promotes the Propagation of Lung Cancer Cells
www.impactjournals.com/oncotarget/ Oncotarget, Vol. 6, No. 25 ANGPTL2/LILRB2 signaling promotes the propagation of lung cancer cells Xiaoye Liu1,2,3,*, Xiaoting Yu4,*, Jingjing Xie5, Mengna Zhan6, Zhuo Yu3, Li Xie3, Hongxiang Zeng3, Feifei Zhang3, Guoqiang Chen1,3, Xianghua Yi4 and Junke Zheng2,3 1 Institute of Health Sciences, Shanghai Institute for Biological Sciences, University of Chinese Academy of Science, Chinese Academy of Sciences and Shanghai Jiao Tong University School of Medicine, Shanghai, China 2 Hongqiao International Institute of Medicine, Shanghai Tongren Hospital, Faculty of Basic Medicine, Shanghai Jiao Tong University School of Medicine, Shanghai, China 3 Key Laboratory of Cell Differentiation and Apoptosis of Chinese Ministry of Education, Shanghai Jiao Tong University School of Medicine, Shanghai, China 4 Department of Pathology, Tongji Hospital, Tongji University School of Medicine, Shanghai, China 5 Bingzhou Medical University, Taishan Scholar Program, Yantai, China 6 Ruijin Hospital, Shanghai Jiao Tong University School of Medicine, Shanghai, China * These authors have contributed equally to this paper Correspondence to: Junke Zheng, email: [email protected] Correspondence to: Xianghua Yi, email: [email protected] Correspondence to: Guoqiang Chen, email: [email protected] Keywords: ANGPTL2/LILRB2 signaling, lung cancer, metastasis, CaMK1 Received: December 31, 2014 Accepted: May 10, 2015 Published: May 20, 2015 This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited. ABSTRACT Immune inhibitory receptors expressed on various types of immune cells deliver inhibitory signals that maintain the homeostasis of the immune system.