The Landscape of Human STR Variation
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Identification of the Binding Partners for Hspb2 and Cryab Reveals
Brigham Young University BYU ScholarsArchive Theses and Dissertations 2013-12-12 Identification of the Binding arP tners for HspB2 and CryAB Reveals Myofibril and Mitochondrial Protein Interactions and Non- Redundant Roles for Small Heat Shock Proteins Kelsey Murphey Langston Brigham Young University - Provo Follow this and additional works at: https://scholarsarchive.byu.edu/etd Part of the Microbiology Commons BYU ScholarsArchive Citation Langston, Kelsey Murphey, "Identification of the Binding Partners for HspB2 and CryAB Reveals Myofibril and Mitochondrial Protein Interactions and Non-Redundant Roles for Small Heat Shock Proteins" (2013). Theses and Dissertations. 3822. https://scholarsarchive.byu.edu/etd/3822 This Thesis is brought to you for free and open access by BYU ScholarsArchive. It has been accepted for inclusion in Theses and Dissertations by an authorized administrator of BYU ScholarsArchive. For more information, please contact [email protected], [email protected]. Identification of the Binding Partners for HspB2 and CryAB Reveals Myofibril and Mitochondrial Protein Interactions and Non-Redundant Roles for Small Heat Shock Proteins Kelsey Langston A thesis submitted to the faculty of Brigham Young University in partial fulfillment of the requirements for the degree of Master of Science Julianne H. Grose, Chair William R. McCleary Brian Poole Department of Microbiology and Molecular Biology Brigham Young University December 2013 Copyright © 2013 Kelsey Langston All Rights Reserved ABSTRACT Identification of the Binding Partners for HspB2 and CryAB Reveals Myofibril and Mitochondrial Protein Interactors and Non-Redundant Roles for Small Heat Shock Proteins Kelsey Langston Department of Microbiology and Molecular Biology, BYU Master of Science Small Heat Shock Proteins (sHSP) are molecular chaperones that play protective roles in cell survival and have been shown to possess chaperone activity. -

Supplementary Table 1: Adhesion Genes Data Set
Supplementary Table 1: Adhesion genes data set PROBE Entrez Gene ID Celera Gene ID Gene_Symbol Gene_Name 160832 1 hCG201364.3 A1BG alpha-1-B glycoprotein 223658 1 hCG201364.3 A1BG alpha-1-B glycoprotein 212988 102 hCG40040.3 ADAM10 ADAM metallopeptidase domain 10 133411 4185 hCG28232.2 ADAM11 ADAM metallopeptidase domain 11 110695 8038 hCG40937.4 ADAM12 ADAM metallopeptidase domain 12 (meltrin alpha) 195222 8038 hCG40937.4 ADAM12 ADAM metallopeptidase domain 12 (meltrin alpha) 165344 8751 hCG20021.3 ADAM15 ADAM metallopeptidase domain 15 (metargidin) 189065 6868 null ADAM17 ADAM metallopeptidase domain 17 (tumor necrosis factor, alpha, converting enzyme) 108119 8728 hCG15398.4 ADAM19 ADAM metallopeptidase domain 19 (meltrin beta) 117763 8748 hCG20675.3 ADAM20 ADAM metallopeptidase domain 20 126448 8747 hCG1785634.2 ADAM21 ADAM metallopeptidase domain 21 208981 8747 hCG1785634.2|hCG2042897 ADAM21 ADAM metallopeptidase domain 21 180903 53616 hCG17212.4 ADAM22 ADAM metallopeptidase domain 22 177272 8745 hCG1811623.1 ADAM23 ADAM metallopeptidase domain 23 102384 10863 hCG1818505.1 ADAM28 ADAM metallopeptidase domain 28 119968 11086 hCG1786734.2 ADAM29 ADAM metallopeptidase domain 29 205542 11085 hCG1997196.1 ADAM30 ADAM metallopeptidase domain 30 148417 80332 hCG39255.4 ADAM33 ADAM metallopeptidase domain 33 140492 8756 hCG1789002.2 ADAM7 ADAM metallopeptidase domain 7 122603 101 hCG1816947.1 ADAM8 ADAM metallopeptidase domain 8 183965 8754 hCG1996391 ADAM9 ADAM metallopeptidase domain 9 (meltrin gamma) 129974 27299 hCG15447.3 ADAMDEC1 ADAM-like, -

Genome Sequences of Tropheus Moorii and Petrochromis Trewavasae, Two Eco‑Morphologically Divergent Cichlid Fshes Endemic to Lake Tanganyika C
www.nature.com/scientificreports OPEN Genome sequences of Tropheus moorii and Petrochromis trewavasae, two eco‑morphologically divergent cichlid fshes endemic to Lake Tanganyika C. Fischer1,2, S. Koblmüller1, C. Börger1, G. Michelitsch3, S. Trajanoski3, C. Schlötterer4, C. Guelly3, G. G. Thallinger2,5* & C. Sturmbauer1,5* With more than 1000 species, East African cichlid fshes represent the fastest and most species‑rich vertebrate radiation known, providing an ideal model to tackle molecular mechanisms underlying recurrent adaptive diversifcation. We add high‑quality genome reconstructions for two phylogenetic key species of a lineage that diverged about ~ 3–9 million years ago (mya), representing the earliest split of the so‑called modern haplochromines that seeded additional radiations such as those in Lake Malawi and Victoria. Along with the annotated genomes we analysed discriminating genomic features of the study species, each representing an extreme trophic morphology, one being an algae browser and the other an algae grazer. The genomes of Tropheus moorii (TM) and Petrochromis trewavasae (PT) comprise 911 and 918 Mbp with 40,300 and 39,600 predicted genes, respectively. Our DNA sequence data are based on 5 and 6 individuals of TM and PT, and the transcriptomic sequences of one individual per species and sex, respectively. Concerning variation, on average we observed 1 variant per 220 bp (interspecifc), and 1 variant per 2540 bp (PT vs PT)/1561 bp (TM vs TM) (intraspecifc). GO enrichment analysis of gene regions afected by variants revealed several candidates which may infuence phenotype modifcations related to facial and jaw morphology, such as genes belonging to the Hedgehog pathway (SHH, SMO, WNT9A) and the BMP and GLI families. -

Biallelic Alteration and Dysregulation of the Hippo Pathway in Mucinous Tubular and Spindle Cell Carcinoma of the Kidney
Published OnlineFirst September 7, 2016; DOI: 10.1158/2159-8290.CD-16-0267 RESEARCH BRIEF Biallelic Alteration and Dysregulation of the Hippo Pathway in Mucinous Tubular and Spindle Cell Carcinoma of the Kidney Rohit Mehra 1 , 2 , 3 , Pankaj Vats 1 , 3 , 4 , Marcin Cieslik 3 , Xuhong Cao 3 , 5 , Fengyun Su 3 , Sudhanshu Shukla 3 , Aaron M. Udager 1 , Rui Wang 3 , Jincheng Pan 6 , Katayoon Kasaian 3 , Robert Lonigro 3 , Javed Siddiqui 3 , Kumpati Premkumar 4 , Ganesh Palapattu 7 , Alon Weizer 2 , 7 , Khaled S. Hafez 7 , J. Stuart Wolf Jr 7 , Ankur R. Sangoi 8 , Kiril Trpkov 9 , Adeboye O. Osunkoya 10 , Ming Zhou 11 , Giovanna A. Giannico 12 , Jesse K. McKenney 13 , Saravana M. Dhanasekaran 1 , 3 , and Arul M. Chinnaiyan 1 , 2 , 3 , 5 , 7 ABSTRACT Mucinous tubular and spindle cell carcinoma (MTSCC) is a relatively rare subtype of renal cell carcinoma (RCC) with distinctive morphologic and cytogenetic fea- tures. Here, we carry out whole-exome and transcriptome sequencing of a multi-institutional cohort of MTSCC (n = 22). We demonstrate the presence of either biallelic loss of Hippo pathway tumor sup- pressor genes (TSG) and/or evidence of alteration of Hippo pathway genes in 85% of samples. PTPN14 (31%) and NF2 (22%) were the most commonly implicated Hippo pathway genes, whereas other genes such as SAV1 and HIPK2 were also involved in a mutually exclusive fashion. Mutations in the context of recurrent chromosomal losses amounted to biallelic alterations in these TSGs. As a readout of Hippo pathway inactivation, a majority of cases (90%) exhibited increased nuclear YAP1 protein expression. -

Insights Into the Genetic Architecture of the Human Face
bioRxiv preprint doi: https://doi.org/10.1101/2020.05.12.090555; this version posted May 14, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. Insights into the genetic architecture of the human face Julie D. White1†*, Karlijne Indencleef2,3,4†*, Sahin Naqvi5,6, Ryan J. Eller7, Jasmien Roosenboom8, Myoung Keun Lee8, Jiarui Li2,3, Jaaved Mohammed5,9, Stephen Richmond10, Ellen E. Quillen11,12, Heather L. Norton13, Eleanor Feingold14, Tomek Swigut5,9, Mary L. Marazita8,14, Hilde Peeters15, Greet Hens4, John R. Shaffer8,14, Joanna Wysocka5,9, Susan Walsh7, Seth M. Weinberg8,14,16, Mark D. Shriver1, Peter Claes2,3,15,17,18* Affiliations: 1Department of Anthropology, Pennsylvania State University, State College, PA, 16802, USA 2Department of Electrical Engineering, ESAT/PSI, KU Leuven, Leuven, 3000, Belgium 3Medical Imaging Research Center, UZ Leuven, Leuven, 3000, Belgium 4Department of Otorhinolaryngology, KU Leuven, Leuven, 3000, Belgium 5Department of Chemical and Systems Biology, Stanford University School of Medicine, Stanford, CA, 94305, USA 6Department of Genetics, Stanford University School of Medicine, Stanford, CA, 94305, USA 7Department of Biology, Indiana University Purdue University Indianapolis, Indianapolis, IN, 46202, USA 8Department of Oral Biology, Center for Craniofacial and Dental Genetics, University of Pittsburgh, Pittsburgh, -

Transcriptomic and Epigenomic Characterization of the Developing Bat Wing
ARTICLES OPEN Transcriptomic and epigenomic characterization of the developing bat wing Walter L Eckalbar1,2,9, Stephen A Schlebusch3,9, Mandy K Mason3, Zoe Gill3, Ash V Parker3, Betty M Booker1,2, Sierra Nishizaki1,2, Christiane Muswamba-Nday3, Elizabeth Terhune4,5, Kimberly A Nevonen4, Nadja Makki1,2, Tara Friedrich2,6, Julia E VanderMeer1,2, Katherine S Pollard2,6,7, Lucia Carbone4,8, Jeff D Wall2,7, Nicola Illing3 & Nadav Ahituv1,2 Bats are the only mammals capable of powered flight, but little is known about the genetic determinants that shape their wings. Here we generated a genome for Miniopterus natalensis and performed RNA-seq and ChIP-seq (H3K27ac and H3K27me3) analyses on its developing forelimb and hindlimb autopods at sequential embryonic stages to decipher the molecular events that underlie bat wing development. Over 7,000 genes and several long noncoding RNAs, including Tbx5-as1 and Hottip, were differentially expressed between forelimb and hindlimb, and across different stages. ChIP-seq analysis identified thousands of regions that are differentially modified in forelimb and hindlimb. Comparative genomics found 2,796 bat-accelerated regions within H3K27ac peaks, several of which cluster near limb-associated genes. Pathway analyses highlighted multiple ribosomal proteins and known limb patterning signaling pathways as differentially regulated and implicated increased forelimb mesenchymal condensation in differential growth. In combination, our work outlines multiple genetic components that likely contribute to bat wing formation, providing insights into this morphological innovation. The order Chiroptera, commonly known as bats, is the only group of To characterize the genetic differences that underlie divergence in mammals to have evolved the capability of flight. -

Nº Ref Uniprot Proteína Péptidos Identificados Por MS/MS 1 P01024
Document downloaded from http://www.elsevier.es, day 26/09/2021. This copy is for personal use. Any transmission of this document by any media or format is strictly prohibited. Nº Ref Uniprot Proteína Péptidos identificados 1 P01024 CO3_HUMAN Complement C3 OS=Homo sapiens GN=C3 PE=1 SV=2 por 162MS/MS 2 P02751 FINC_HUMAN Fibronectin OS=Homo sapiens GN=FN1 PE=1 SV=4 131 3 P01023 A2MG_HUMAN Alpha-2-macroglobulin OS=Homo sapiens GN=A2M PE=1 SV=3 128 4 P0C0L4 CO4A_HUMAN Complement C4-A OS=Homo sapiens GN=C4A PE=1 SV=1 95 5 P04275 VWF_HUMAN von Willebrand factor OS=Homo sapiens GN=VWF PE=1 SV=4 81 6 P02675 FIBB_HUMAN Fibrinogen beta chain OS=Homo sapiens GN=FGB PE=1 SV=2 78 7 P01031 CO5_HUMAN Complement C5 OS=Homo sapiens GN=C5 PE=1 SV=4 66 8 P02768 ALBU_HUMAN Serum albumin OS=Homo sapiens GN=ALB PE=1 SV=2 66 9 P00450 CERU_HUMAN Ceruloplasmin OS=Homo sapiens GN=CP PE=1 SV=1 64 10 P02671 FIBA_HUMAN Fibrinogen alpha chain OS=Homo sapiens GN=FGA PE=1 SV=2 58 11 P08603 CFAH_HUMAN Complement factor H OS=Homo sapiens GN=CFH PE=1 SV=4 56 12 P02787 TRFE_HUMAN Serotransferrin OS=Homo sapiens GN=TF PE=1 SV=3 54 13 P00747 PLMN_HUMAN Plasminogen OS=Homo sapiens GN=PLG PE=1 SV=2 48 14 P02679 FIBG_HUMAN Fibrinogen gamma chain OS=Homo sapiens GN=FGG PE=1 SV=3 47 15 P01871 IGHM_HUMAN Ig mu chain C region OS=Homo sapiens GN=IGHM PE=1 SV=3 41 16 P04003 C4BPA_HUMAN C4b-binding protein alpha chain OS=Homo sapiens GN=C4BPA PE=1 SV=2 37 17 Q9Y6R7 FCGBP_HUMAN IgGFc-binding protein OS=Homo sapiens GN=FCGBP PE=1 SV=3 30 18 O43866 CD5L_HUMAN CD5 antigen-like OS=Homo -

Genes Associated with the Progression of Neurofibrillary Tangles in Alzheimer’S Disease
OPEN Citation: Transl Psychiatry (2014) 4, e396; doi:10.1038/tp.2014.35 © 2014 Macmillan Publishers Limited All rights reserved 2158-3188/14 www.nature.com/tp ORIGINAL ARTICLE Genes associated with the progression of neurofibrillary tangles in Alzheimer’s disease A Miyashita1,10, H Hatsuta2,10, M Kikuchi3,10, A Nakaya4, Y Saito5, T Tsukie3, N Hara1, S Ogishima6, N Kitamura7, K Akazawa7, A Kakita8, H Takahashi8, S Murayama2, Y Ihara9, T Ikeuchi1, R Kuwano1 and Japanese Alzheimer’s Disease Neuroimaging Initiative The spreading of neurofibrillary tangles (NFTs), intraneuronal aggregates of highly phosphorylated microtubule-associated protein tau, across the human brain is correlated with the cognitive severity of Alzheimer’s disease (AD). To identify genes relevant to NFT expansion defined by the Braak stage, we conducted whole-genome exon array analysis with an exploratory sample set consisting of 213 human post-mortem brain tissue specimens from the entorinal, temporal and frontal cortices of 71 brain-donor subjects: Braak NFT stages 0 (N = 13), I–II (N = 20), III–IV (N = 19) and V–VI (N = 19). We identified eight genes, RELN, PTGS2, MYO5C, TRIL, DCHS2, GRB14, NPAS4 and PHYHD1, associated with the Braak stage. The expression levels of three genes, PHYHD1, MYO5C and GRB14, exhibited reproducible association on real-time quantitative PCR analysis. In another sample set, including control subjects (N = 30), and in patients with late-onset AD (N = 37), dementia with Lewy bodies (N = 17) and Parkinson disease (N = 36), the expression levels of two genes, PHYHD1 and MYO5C, were obviously associated with late-onset AD. Protein–protein interaction network analysis with a public database revealed that PHYHD1 interacts with MYO5C via POT1, and PHYHD1 directly interacts with amyloid beta-peptide 42. -

Genomic Portrait of a Sporadic Amyotrophic Lateral Sclerosis Case in a Large Spinocerebellar Ataxia Type 1 Family
Journal of Personalized Medicine Article Genomic Portrait of a Sporadic Amyotrophic Lateral Sclerosis Case in a Large Spinocerebellar Ataxia Type 1 Family Giovanna Morello 1,2, Giulia Gentile 1 , Rossella Spataro 3, Antonio Gianmaria Spampinato 1,4, 1 2 3 5, , Maria Guarnaccia , Salvatore Salomone , Vincenzo La Bella , Francesca Luisa Conforti * y 1, , and Sebastiano Cavallaro * y 1 Institute for Research and Biomedical Innovation (IRIB), Italian National Research Council (CNR), Via Paolo Gaifami, 18, 95125 Catania, Italy; [email protected] (G.M.); [email protected] (G.G.); [email protected] (A.G.S.); [email protected] (M.G.) 2 Department of Biomedical and Biotechnological Sciences, Section of Pharmacology, University of Catania, 95123 Catania, Italy; [email protected] 3 ALS Clinical Research Center and Neurochemistry Laboratory, BioNeC, University of Palermo, 90127 Palermo, Italy; [email protected] (R.S.); [email protected] (V.L.B.) 4 Department of Mathematics and Computer Science, University of Catania, 95123 Catania, Italy 5 Department of Pharmacy, Health and Nutritional Sciences, University of Calabria, Arcavacata di Rende, 87036 Rende, Italy * Correspondence: [email protected] (F.L.C.); [email protected] (S.C.); Tel.: +39-0984-496204 (F.L.C.); +39-095-7338111 (S.C.); Fax: +39-0984-496203 (F.L.C.); +39-095-7338110 (S.C.) F.L.C. and S.C. are co-last authors on this work. y Received: 6 November 2020; Accepted: 30 November 2020; Published: 2 December 2020 Abstract: Background: Repeat expansions in the spinocerebellar ataxia type 1 (SCA1) gene ATXN1 increases the risk for amyotrophic lateral sclerosis (ALS), supporting a relationship between these disorders. -

From Syndromes to Normal Variation: a Candidate Gene Study of Interorbital Distances
FROM SYNDROMES TO NORMAL VARIATION: A CANDIDATE GENE STUDY OF INTERORBITAL DISTANCES by Samantha Wesoly BS, Molecular and Cell Biology, University of Connecticut, 2014 Submitted to the Graduate Faculty of the Department of Human Genetics in the Graduate School of Public Health in partial fulfillment of the requirements for the degree of Master of Science University of Pittsburgh 2017 UNIVERSITY OF PITTSBURGH GRADUATE SCHOOL OF PUBLIC HEALTH This thesis was presented by Samantha Wesoly It was defended on April 14, 2017 and approved by Committee Chair: Seth Weinberg, PhD Associate Professor, Department of Oral Biology School of Dental Medicine, University of Pittsburgh Committee Member: Elizabeth J Leslie, PhD Assistant Professor, Department of Oral Biology School of Dental Medicine, University of Pittsburgh Committee Member: Robin Grubs, M.S., PhD, LCGC Assistant Professor and Director of Genetic Counseling Program Department of Human Genetics Graduate School of Public Health, University of Pittsburgh Committee Member: John R. Shaffer, PhD Assistant Professor, Department of Human Genetics Graduate School of Public Health, University of Pittsburgh ii Copyright © by Samantha Wesoly 2017 iii Seth Weinberg, PhD FROM SYNDROMES TO NORMAL VARIATION: A CANDIDATE GENE STUDY OF INTERORBITAL DISTANCES Samantha Wesoly, MS University of Pittsburgh, 2017 ABSTRACT Hypertelorism and telecanthus are clinical phenotypes associated with many genetic syndromes. To date, research is limited regarding whether disease-causing genes are related to normal craniofacial development in unaffected individuals. The aim of this study is to determine whether common genetic variation in forty selected genes implicated in hypertelorism/telecanthus- related syndromes contribute to normal variation of intercanthal and outer-canthal distances of the orbits. -
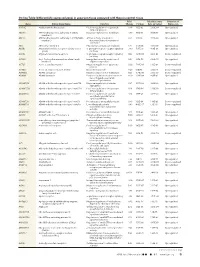
Differentially Expressed Genes in Aneurysm Tissue Compared With
On-line Table: Differentially expressed genes in aneurysm tissue compared with those in control tissue Fold False Discovery Direction of Gene Entrez Gene Name Function Change P Value Rate (q Value) Expression AADAC Arylacetamide deacetylase Positive regulation of triglyceride 4.46 1.33E-05 2.60E-04 Up-regulated catabolic process ABCA6 ATP-binding cassette, subfamily A (ABC1), Integral component of membrane 3.79 9.15E-14 8.88E-12 Up-regulated member 6 ABCC3 ATP-binding cassette, subfamily C (CFTR/MRP), ATPase activity, coupled to 6.63 1.21E-10 7.33E-09 Up-regulated member 3 transmembrane movement of substances ABI3 ABI family, member 3 Peptidyl-tyrosine phosphorylation 6.47 2.47E-05 4.56E-04 Up-regulated ACKR1 Atypical chemokine receptor 1 (Duffy blood G-protein–coupled receptor signaling 3.80 7.95E-10 4.18E-08 Up-regulated group) pathway ACKR2 Atypical chemokine receptor 2 G-protein–coupled receptor signaling 0.42 3.29E-04 4.41E-03 Down-regulated pathway ACSM1 Acyl-CoA synthetase medium-chain family Energy derivation by oxidation of 9.87 1.70E-08 6.52E-07 Up-regulated member 1 organic compounds ACTC1 Actin, ␣, cardiac muscle 1 Negative regulation of apoptotic 0.30 7.96E-06 1.65E-04 Down-regulated process ACTG2 Actin, ␥2, smooth muscle, enteric Blood microparticle 0.29 1.61E-16 2.36E-14 Down-regulated ADAM33 ADAM domain 33 Integral component of membrane 0.23 9.74E-09 3.95E-07 Down-regulated ADAM8 ADAM domain 8 Positive regulation of tumor necrosis 4.69 2.93E-04 4.01E-03 Up-regulated factor (ligand) superfamily member 11 production ADAMTS18 -

Detection of H3k4me3 Identifies Neurohiv Signatures, Genomic
viruses Article Detection of H3K4me3 Identifies NeuroHIV Signatures, Genomic Effects of Methamphetamine and Addiction Pathways in Postmortem HIV+ Brain Specimens that Are Not Amenable to Transcriptome Analysis Liana Basova 1, Alexander Lindsey 1, Anne Marie McGovern 1, Ronald J. Ellis 2 and Maria Cecilia Garibaldi Marcondes 1,* 1 San Diego Biomedical Research Institute, San Diego, CA 92121, USA; [email protected] (L.B.); [email protected] (A.L.); [email protected] (A.M.M.) 2 Departments of Neurosciences and Psychiatry, University of California San Diego, San Diego, CA 92103, USA; [email protected] * Correspondence: [email protected] Abstract: Human postmortem specimens are extremely valuable resources for investigating trans- lational hypotheses. Tissue repositories collect clinically assessed specimens from people with and without HIV, including age, viral load, treatments, substance use patterns and cognitive functions. One challenge is the limited number of specimens suitable for transcriptional studies, mainly due to poor RNA quality resulting from long postmortem intervals. We hypothesized that epigenomic Citation: Basova, L.; Lindsey, A.; signatures would be more stable than RNA for assessing global changes associated with outcomes McGovern, A.M.; Ellis, R.J.; of interest. We found that H3K27Ac or RNA Polymerase (Pol) were not consistently detected by Marcondes, M.C.G. Detection of H3K4me3 Identifies NeuroHIV Chromatin Immunoprecipitation (ChIP), while the enhancer H3K4me3 histone modification was Signatures, Genomic Effects of abundant and stable up to the 72 h postmortem. We tested our ability to use H3K4me3 in human Methamphetamine and Addiction prefrontal cortex from HIV+ individuals meeting criteria for methamphetamine use disorder or not Pathways in Postmortem HIV+ Brain (Meth +/−) which exhibited poor RNA quality and were not suitable for transcriptional profiling.