In Silico Analysis of Single Nucleotide Polymorphisms (Snps) in Human MPL Gene
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Microdeletions in 16P11.2 and 13Q31.3 Associated with Developmental Delay and Generalized Overgrowth
Microdeletions in 16p11.2 and 13q31.3 associated with developmental delay and generalized overgrowth A.M. George1, J. Taylor2 and D.R. Love1 1Diagnostic Genetics, LabPlus, Auckland City Hospital, Auckland, New Zealand 2Northern Regional Genetic Service, Auckland City Hospital, Auckland, New Zealand Corresponding author: D.R. Love E-mail: [email protected] Genet. Mol. Res. 11 (3): 3133-3137 (2012) Received November 28, 2011 Accepted July 18, 2012 Published September 3, 2012 DOI http://dx.doi.org/10.4238/2012.September.3.1 ABSTRACT. Chromosome microarray analysis of patients with developmental delay has provided evidence of small deletions or duplications associated with this clinical phenotype. In this context, a 7.1- to 8.7-Mb interstitial deletion of chromosome 16 is well documented, but within this interval a rare 200-kb deletion has recently been defined that appears to be associated with obesity, or developmental delay together with overgrowth. We report a patient carrying this rare deletion, who falls into the latter clinical category, but who also carries a second very rare deletion in 13q31.3. It remains unclear if this maternally inherited deletion acts as a second copy number variation leading to pathogenic variation, or is non-causal and the true modifiers are yet to be determined. Key words: Developmental delay; Obesity; Overgrowth; GPC5; SH2B1 Genetics and Molecular Research 11 (3): 3133-3137 (2012) ©FUNPEC-RP www.funpecrp.com.br A.M. George et al. 3134 INTRODUCTION Current referrals for chromosome microarray analysis (CMA) are primarily for de- termining the molecular basis of developmental delay and autistic spectrum disorder in child- hood. -

Autism Multiplex Family with 16P11.2P12.2 Microduplication Syndrome in Monozygotic Twins and Distal 16P11.2 Deletion in Their Brother
European Journal of Human Genetics (2012) 20, 540–546 & 2012 Macmillan Publishers Limited All rights reserved 1018-4813/12 www.nature.com/ejhg ARTICLE Autism multiplex family with 16p11.2p12.2 microduplication syndrome in monozygotic twins and distal 16p11.2 deletion in their brother Anne-Claude Tabet1,2,3,4, Marion Pilorge2,3,4, Richard Delorme5,6,Fre´de´rique Amsellem5,6, Jean-Marc Pinard7, Marion Leboyer6,8,9, Alain Verloes10, Brigitte Benzacken1,11,12 and Catalina Betancur*,2,3,4 The pericentromeric region of chromosome 16p is rich in segmental duplications that predispose to rearrangements through non-allelic homologous recombination. Several recurrent copy number variations have been described recently in chromosome 16p. 16p11.2 rearrangements (29.5–30.1 Mb) are associated with autism, intellectual disability (ID) and other neurodevelopmental disorders. Another recognizable but less common microdeletion syndrome in 16p11.2p12.2 (21.4 to 28.5–30.1 Mb) has been described in six individuals with ID, whereas apparently reciprocal duplications, studied by standard cytogenetic and fluorescence in situ hybridization techniques, have been reported in three patients with autism spectrum disorders. Here, we report a multiplex family with three boys affected with autism, including two monozygotic twins carrying a de novo 16p11.2p12.2 duplication of 8.95 Mb (21.28–30.23 Mb) characterized by single-nucleotide polymorphism array, encompassing both the 16p11.2 and 16p11.2p12.2 regions. The twins exhibited autism, severe ID, and dysmorphic features, including a triangular face, deep-set eyes, large and prominent nasal bridge, and tall, slender build. The eldest brother presented with autism, mild ID, early-onset obesity and normal craniofacial features, and carried a smaller, overlapping 16p11.2 microdeletion of 847 kb (28.40–29.25 Mb), inherited from his apparently healthy father. -

1 Supporting Information for a Microrna Network Regulates
Supporting Information for A microRNA Network Regulates Expression and Biosynthesis of CFTR and CFTR-ΔF508 Shyam Ramachandrana,b, Philip H. Karpc, Peng Jiangc, Lynda S. Ostedgaardc, Amy E. Walza, John T. Fishere, Shaf Keshavjeeh, Kim A. Lennoxi, Ashley M. Jacobii, Scott D. Rosei, Mark A. Behlkei, Michael J. Welshb,c,d,g, Yi Xingb,c,f, Paul B. McCray Jr.a,b,c Author Affiliations: Department of Pediatricsa, Interdisciplinary Program in Geneticsb, Departments of Internal Medicinec, Molecular Physiology and Biophysicsd, Anatomy and Cell Biologye, Biomedical Engineeringf, Howard Hughes Medical Instituteg, Carver College of Medicine, University of Iowa, Iowa City, IA-52242 Division of Thoracic Surgeryh, Toronto General Hospital, University Health Network, University of Toronto, Toronto, Canada-M5G 2C4 Integrated DNA Technologiesi, Coralville, IA-52241 To whom correspondence should be addressed: Email: [email protected] (M.J.W.); yi- [email protected] (Y.X.); Email: [email protected] (P.B.M.) This PDF file includes: Materials and Methods References Fig. S1. miR-138 regulates SIN3A in a dose-dependent and site-specific manner. Fig. S2. miR-138 regulates endogenous SIN3A protein expression. Fig. S3. miR-138 regulates endogenous CFTR protein expression in Calu-3 cells. Fig. S4. miR-138 regulates endogenous CFTR protein expression in primary human airway epithelia. Fig. S5. miR-138 regulates CFTR expression in HeLa cells. Fig. S6. miR-138 regulates CFTR expression in HEK293T cells. Fig. S7. HeLa cells exhibit CFTR channel activity. Fig. S8. miR-138 improves CFTR processing. Fig. S9. miR-138 improves CFTR-ΔF508 processing. Fig. S10. SIN3A inhibition yields partial rescue of Cl- transport in CF epithelia. -

A Novel Resveratrol Analog: Its Cell Cycle Inhibitory, Pro-Apoptotic and Anti-Inflammatory Activities on Human Tumor Cells
A NOVEL RESVERATROL ANALOG : ITS CELL CYCLE INHIBITORY, PRO-APOPTOTIC AND ANTI-INFLAMMATORY ACTIVITIES ON HUMAN TUMOR CELLS A dissertation submitted to Kent State University in partial fulfillment of the requirements for the degree of Doctor of Philosophy by Boren Lin May 2006 Dissertation written by Boren Lin B.S., Tunghai University, 1996 M.S., Kent State University, 2003 Ph. D., Kent State University, 2006 Approved by Dr. Chun-che Tsai , Chair, Doctoral Dissertation Committee Dr. Bryan R. G. Williams , Co-chair, Doctoral Dissertation Committee Dr. Johnnie W. Baker , Members, Doctoral Dissertation Committee Dr. James L. Blank , Dr. Bansidhar Datta , Dr. Gail C. Fraizer , Accepted by Dr. Robert V. Dorman , Director, School of Biomedical Sciences Dr. John R. Stalvey , Dean, College of Arts and Sciences ii TABLE OF CONTENTS LIST OF FIGURES……………………………………………………………….………v LIST OF TABLES……………………………………………………………………….vii ACKNOWLEDGEMENTS….………………………………………………………….viii I INTRODUCTION….………………………………………………….1 Background and Significance……………………………………………………..1 Specific Aims………………………………………………………………………12 II MATERIALS AND METHODS.…………………………………………….16 Cell Culture and Compounds…….……………….…………………………….….16 MTT Cell Viability Assay………………………………………………………….16 Trypan Blue Exclusive Assay……………………………………………………...18 Flow Cytometry for Cell Cycle Analysis……………..……………....……………19 DNA Fragmentation Assay……………………………………………...…………23 Caspase-3 Activity Assay………………………………...……….….…….………24 Annexin V-FITC Staining Assay…………………………………..…...….………28 NF-kappa B p65 Activity Assay……………………………………..………….…29 -

The I-Motif As a Molecular Target: More Than a Complementary DNA Secondary Structure
pharmaceuticals Review The i-Motif as a Molecular Target: More Than a Complementary DNA Secondary Structure Susie L. Brown and Samantha Kendrick * Biochemistry and Molecular Biology Department, University of Arkansas for Medical Sciences, Little Rock, AR 72205, USA; [email protected] * Correspondence: [email protected] Abstract: Stretches of cytosine-rich DNA are capable of adopting a dynamic secondary structure, the i-motif. When within promoter regions, the i-motif has the potential to act as a molecular switch for controlling gene expression. However, i-motif structures in genomic areas of repetitive nucleotide sequences may play a role in facilitating or hindering expansion of these DNA elements. Despite research on the i-motif trailing behind the complementary G-quadruplex structure, recent discoveries including the identification of a specific i-motif antibody are pushing this field forward. This perspective reviews initial and current work characterizing the i-motif and providing insight into the biological function of this DNA structure, with a focus on how the i-motif can serve as a molecular target for developing new therapeutic approaches to modulate gene expression and extension of repetitive DNA. Keywords: i-motif; DNA quadruplex structures; transcription regulation; cancer therapeutics 1. Introduction Citation: Brown, S.L; Kendrick, S. In 1953, Watson and Crick first published the structure of the DNA double helix [1]. The i-Motif as a Molecular Target: They described a DNA molecule as a right-handed, twisted coil composed of a purine and More Than a Complementary DNA pyrimidine inner core held together by hydrogen bonds, with a sugar–phosphate back- Secondary Structure. -

Structure and Expression Analyses of SVA Elements in Relation to Functional Genes
pISSN 1598-866X eISSN 2234-0742 Genomics Inform 2013;11(3):142-148 G&I Genomics & Informatics http://dx.doi.org/10.5808/GI.2013.11.3.142 ORIGINAL ARTICLE Structure and Expression Analyses of SVA Elements in Relation to Functional Genes Yun-Jeong Kwon, Yuri Choi, Jungwoo Eo, Yu-Na Noh, Jeong-An Gim, Yi-Deun Jung, Ja-Rang Lee, Heui-Soo Kim* Department of Biological Sciences, College of Natural Sciences, Pusan National University, Busan 609-735, Korea SINE-VNTR-Alu (SVA) elements are present in hominoid primates and are divided into 6 subfamilies (SVA-A to SVA-F) and active in the human population. Using a bioinformatic tool, 22 SVA element-associated genes are identified in the human genome. In an analysis of genomic structure, SVA elements are detected in the 5′ untranslated region (UTR) of HGSNAT (SVA-B), MRGPRX3 (SVA-D), HYAL1 (SVA-F), TCHH (SVA-F), and ATXN2L (SVA-F) genes, while some elements are observed in the 3′UTR of SPICE1 (SVA-B), TDRKH (SVA-C), GOSR1 (SVA-D), BBS5 (SVA-D), NEK5 (SVA-D), ABHD2 (SVA-F), C1QTNF7 (SVA-F), ORC6L (SVA-F), TMEM69 (SVA-F), and CCDC137 (SVA-F) genes. They could contribute to exon extension or supplying poly A signals. LEPR (SVA-C), ALOX5 (SVA-D), PDS5B (SVA-D), and ABCA10 (SVA-F) genes also showed alternative transcripts by SVA exonization events. Dominant expression of HYAL1_SVA appeared in lung tissues, while HYAL1_noSVA showed ubiquitous expression in various human tissues. Expression of both transcripts (TDRKH_SVA and TDRKH_noSVA) of the TDRKH gene appeared to be ubiquitous. -

Rabbit Anti-TPOR/FITC Conjugated Antibody-SL10362R-FITC
SunLong Biotech Co.,LTD Tel: 0086-571- 56623320 Fax:0086-571- 56623318 E-mail:[email protected] www.sunlongbiotech.com Rabbit Anti-TPOR/FITC Conjugated antibody SL10362R-FITC Product Name: Anti-TPOR/FITC Chinese Name: FITC标记的血小板生成素受体抗体 C MPL; C-MPL; CMPL; CD110; CD 110; MPL; MPLV; Myeloproliferative leukemia Alias: protein; Myeloproliferative leukemia virus oncogene; Proto-oncogene c-Mpl; THCYT2; Thrombopoietin receptor; TPO R; TPO-R; TPOR; TPOR_HUMAN. Organism Species: Rabbit Clonality: Polyclonal React Species: Human,Mouse,Rat,Dog,Sheep, ICC=1:50-200IF=1:50-200 Applications: not yet tested in other applications. optimal dilutions/concentrations should be determined by the end user. Molecular weight: 68kDa Form: Lyophilized or Liquid Concentration: 1mg/ml immunogen: KLH conjugated synthetic peptide derived from human TPOR Lsotype: IgG Purification: affinity purified by Protein A Storage Buffer: 0.01Mwww.sunlongbiotech.com TBS(pH7.4) with 1% BSA, 0.03% Proclin300 and 50% Glycerol. Store at -20 °C for one year. Avoid repeated freeze/thaw cycles. The lyophilized antibody is stable at room temperature for at least one month and for greater than a year Storage: when kept at -20°C. When reconstituted in sterile pH 7.4 0.01M PBS or diluent of antibody the antibody is stable for at least two weeks at 2-4 °C. background: In 1990 an oncogene, v-mpl, was identified from the murine myeloproliferative leukemia virus that was capable of immortalizing bone marrow hematopoietic cells from different lineages. In 1992 the human homologue, named, c-mpl, was cloned. Product Detail: Sequence data revealed that c-mpl encoded a protein that was homologous with members of the hematopoietic receptor superfamily. -

Supplementary Table 1
Supplementary Table 1. 492 genes are unique to 0 h post-heat timepoint. The name, p-value, fold change, location and family of each gene are indicated. Genes were filtered for an absolute value log2 ration 1.5 and a significance value of p ≤ 0.05. Symbol p-value Log Gene Name Location Family Ratio ABCA13 1.87E-02 3.292 ATP-binding cassette, sub-family unknown transporter A (ABC1), member 13 ABCB1 1.93E-02 −1.819 ATP-binding cassette, sub-family Plasma transporter B (MDR/TAP), member 1 Membrane ABCC3 2.83E-02 2.016 ATP-binding cassette, sub-family Plasma transporter C (CFTR/MRP), member 3 Membrane ABHD6 7.79E-03 −2.717 abhydrolase domain containing 6 Cytoplasm enzyme ACAT1 4.10E-02 3.009 acetyl-CoA acetyltransferase 1 Cytoplasm enzyme ACBD4 2.66E-03 1.722 acyl-CoA binding domain unknown other containing 4 ACSL5 1.86E-02 −2.876 acyl-CoA synthetase long-chain Cytoplasm enzyme family member 5 ADAM23 3.33E-02 −3.008 ADAM metallopeptidase domain Plasma peptidase 23 Membrane ADAM29 5.58E-03 3.463 ADAM metallopeptidase domain Plasma peptidase 29 Membrane ADAMTS17 2.67E-04 3.051 ADAM metallopeptidase with Extracellular other thrombospondin type 1 motif, 17 Space ADCYAP1R1 1.20E-02 1.848 adenylate cyclase activating Plasma G-protein polypeptide 1 (pituitary) receptor Membrane coupled type I receptor ADH6 (includes 4.02E-02 −1.845 alcohol dehydrogenase 6 (class Cytoplasm enzyme EG:130) V) AHSA2 1.54E-04 −1.6 AHA1, activator of heat shock unknown other 90kDa protein ATPase homolog 2 (yeast) AK5 3.32E-02 1.658 adenylate kinase 5 Cytoplasm kinase AK7 -

Mid-Gestation Lethality of Atxn2l-Ablated Mice
International Journal of Molecular Sciences Article Mid-Gestation lethality of Atxn2l-Ablated Mice Jana Key 1,2, Patrick N. Harter 3, Nesli-Ece Sen 1,2, Elise Gradhand 4, Georg Auburger 1,* and Suzana Gispert 1,* 1 Exp. Neurology, Medical Faculty, Goethe University, Theodor Stern Kai 7, 60590 Frankfurt am Main, Germany; [email protected] (J.K.); [email protected] (N.-E.S.) 2 Faculty of Biosciences, Goethe-University, Altenhöferallee 1, 60438 Frankfurt am Main, Germany 3 Institute of Neurology (Edinger-Institute), University Hospital Frankfurt, Goethe University, Heinrich-Hoffmann-Strasse 7, 60528 Frankfurt am Main, Germany; [email protected] 4 Dr. Senckenberg Institute for Pathology, University Hospital, Goethe University, Theodor-Stern-Kai-7, 60590 Frankfurt am Main, Germany; [email protected] * Correspondence: [email protected] (G.A.); [email protected] (S.G.); Tel.: +49-69-6301-7428 (G.A.); +49-69-6301-7417 (S.G.) Received: 19 June 2020; Accepted: 16 July 2020; Published: 20 July 2020 Abstract: Depletion of yeast/fly Ataxin-2 rescues TDP-43 overexpression toxicity. In mouse models of Amyotrophic Lateral Sclerosis via TDP-43 overexpression, depletion of its ortholog ATXN2 mitigated motor neuron degeneration and extended lifespan from 25 days to >300 days. There is another ortholog in mammals, named ATXN2L (Ataxin-2-like), which is almost uncharacterized but also functions in RNA surveillance at stress granules. We generated mice with Crispr/Cas9-mediated deletion of Atxn2l exons 5-8, studying homozygotes prenatally and heterozygotes during aging. Our novel findings indicate that ATXN2L absence triggers mid-gestational embryonic lethality, affecting female animals more strongly. -

A Yeast Bifc-Seq Method for Genome-Wide Interactome Mapping
bioRxiv preprint doi: https://doi.org/10.1101/2020.06.16.154146; this version posted June 17, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. 1 A yeast BiFC-seq method for genome-wide interactome mapping 2 3 Limin Shang1,a, Yuehui Zhang1,b, Yuchen Liu1,c, Chaozhi Jin1,d, Yanzhi Yuan1,e, 4 Chunyan Tian1,f, Ming Ni2,g, Xiaochen Bo2,h, Li Zhang3,i, Dong Li1,j, Fuchu He1,*,k & 5 Jian Wang1,*,l 6 1State Key Laboratory of Proteomics, Beijing Proteome Research Center, National 7 Center for Protein Sciences (Beijing), Beijing Institute of Lifeomics, Beijing 102206, 8 China; 2Department of Biotechnology, Beijing Institute of Radiation Medicine, 9 Beijing 100850, China; 3Department of Rehabilitation Medicine, Nan Lou; 10 Department of Key Laboratory of Wound Repair and Regeneration of PLA, College 11 of Life Sciences, Chinese PLA General Hospital, Beijing 100853, China; 12 Correspondence should be addressed to F.H. ([email protected]) and J.W. 13 ([email protected]). 14 * Corresponding authors. 15 E-mail:[email protected] (Fuchu He),[email protected] (Jian Wang) 16 a ORCID: 0000-0002-6371-1956. 17 b ORCID: 0000-0001-5257-1671 18 c ORCID: 0000-0003-4691-4951 19 d ORCID: 0000-0002-1477-0255 20 e ORCID: 0000-0002-6576-8112 21 f ORCID: 0000-0003-1589-293X 22 g ORCID: 0000-0001-9465-2787 23 h ORCID: 0000-0003-3490-5812 24 i ORCID: 0000-0002-3477-8860 25 j ORCID: 0000-0002-8680-0468 26 k ORCID: 0000-0002-8571-2351 27 l ORCID: 0000-0002-8116-7691 28 Total word counts:4398 (from “Introduction” to “Conclusions”) 29 Total Keywords: 5 bioRxiv preprint doi: https://doi.org/10.1101/2020.06.16.154146; this version posted June 17, 2020. -

ATXN2L Sirna (Mouse)
For research purposes only, not for human use Product Data Sheet ATXN2L siRNA (Mouse) Catalog # Source Reactivity Applications CRN3114 Synthetic M RNAi Description siRNA to inhibit ATXN2L expression using RNA interference Specificity ATXN2L siRNA (Mouse) is a target-specific 19-23 nt siRNA oligo duplexes designed to knock down gene expression. Form Lyophilized powder Gene Symbol ATXN2L Alternative Names A2LP; Ataxin-2-like protein Entrez Gene 233871 (Mouse) SwissProt Q7TQH0 (Mouse) Purity > 97% Quality Control Oligonucleotide synthesis is monitored base by base through trityl analysis to ensure appropriate coupling efficiency. The oligo is subsequently purified by affinity-solid phase extraction. The annealed RNA duplex is further analyzed by mass spectrometry to verify the exact composition of the duplex. Each lot is compared to the previous lot by mass spectrometry to ensure maximum lot-to-lot consistency. Components We offers pre-designed sets of 3 different target-specific siRNA oligo duplexes of mouse ATXN2L gene. Each vial contains 5 nmol of lyophilized siRNA. The duplexes can be transfected individually or pooled together to achieve knockdown of the target gene, which is most commonly assessed by qPCR or western blot. Our siRNA oligos are also chemically modified (2’-OMe) at no extra charge for increased stability and enhanced knockdown in vitro and in vivo. Directions for Use We recommends transfection with 100 nM siRNA 48 to 72 hours prior to cell lysis. Application key: E- ELISA, WB- Western blot, IH- Immunohistochemistry, IF- -
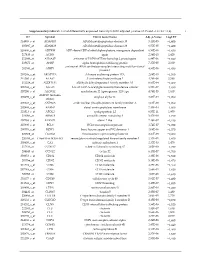
Supplementary Table S1. List of Differentially Expressed
Supplementary table S1. List of differentially expressed transcripts (FDR adjusted p‐value < 0.05 and −1.4 ≤ FC ≥1.4). 1 ID Symbol Entrez Gene Name Adj. p‐Value Log2 FC 214895_s_at ADAM10 ADAM metallopeptidase domain 10 3,11E‐05 −1,400 205997_at ADAM28 ADAM metallopeptidase domain 28 6,57E‐05 −1,400 220606_s_at ADPRM ADP‐ribose/CDP‐alcohol diphosphatase, manganese dependent 6,50E‐06 −1,430 217410_at AGRN agrin 2,34E‐10 1,420 212980_at AHSA2P activator of HSP90 ATPase homolog 2, pseudogene 6,44E‐06 −1,920 219672_at AHSP alpha hemoglobin stabilizing protein 7,27E‐05 2,330 aminoacyl tRNA synthetase complex interacting multifunctional 202541_at AIMP1 4,91E‐06 −1,830 protein 1 210269_s_at AKAP17A A‐kinase anchoring protein 17A 2,64E‐10 −1,560 211560_s_at ALAS2 5ʹ‐aminolevulinate synthase 2 4,28E‐06 3,560 212224_at ALDH1A1 aldehyde dehydrogenase 1 family member A1 8,93E‐04 −1,400 205583_s_at ALG13 ALG13 UDP‐N‐acetylglucosaminyltransferase subunit 9,50E‐07 −1,430 207206_s_at ALOX12 arachidonate 12‐lipoxygenase, 12S type 4,76E‐05 1,630 AMY1C (includes 208498_s_at amylase alpha 1C 3,83E‐05 −1,700 others) 201043_s_at ANP32A acidic nuclear phosphoprotein 32 family member A 5,61E‐09 −1,760 202888_s_at ANPEP alanyl aminopeptidase, membrane 7,40E‐04 −1,600 221013_s_at APOL2 apolipoprotein L2 6,57E‐11 1,600 219094_at ARMC8 armadillo repeat containing 8 3,47E‐08 −1,710 207798_s_at ATXN2L ataxin 2 like 2,16E‐07 −1,410 215990_s_at BCL6 BCL6 transcription repressor 1,74E‐07 −1,700 200776_s_at BZW1 basic leucine zipper and W2 domains 1 1,09E‐06 −1,570 222309_at