SIRT1 Regulates Sphingolipid Metabolism and Neural
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

National Lipid Association Annual Summary of Clinical Lipidology 2015
Journal of Clinical Lipidology (2014) 8, S1-S36 Original Contribution National Lipid Association Annual Summary of Clinical Lipidology 2015 Harold E. Bays, MD, FTOS, FACC, FACE, FNLA*, Peter H. Jones, MD, FACP, FNLA, W. Virgil Brown, MD, FNLA, Terry A. Jacobson, MD, FACP, FNLA Louisville Metabolic and Atherosclerosis Research Center, Louisville, KY, USA (Dr Bays); Baylor College of Medicine, Houston, TX, USA (Dr Jones); Emory University School of Medicine, Atlanta, GA, USA (Dr Brown); and Department of Medicine, Emory University, Atlanta, GA, USA (Dr Jacobson) KEYWORDS: Abstract: The National Lipid Association (NLA) Annual Summary of Clinical Lipidology 2015 is a Clinical Lipidology; summary of principles important to the patient-centered evaluation, management, and care of patients Dyslipidemia; with dyslipidemia. This summary is intended to be a ‘‘living document,’’ with future annual updates National Lipid based on emerging science, clinical considerations, and new NLA Position and Consensus Statements. Association; The goal is to provide clinicians an ongoing resource that translates the latest advances in medical Annual Summary; science toward the evaluation and treatment of patients with dyslipidemia. The 2015 NLA Annual Cholesterol; Summary of Clinical Lipidology was founded on the principles of evidence-based medicine and is Recommendations; generally consistent with established national and international lipid guidelines. Topics include a Guidelines general discussion of the 2014 NLA Recommendations for Patient-Centered Management of Dyslipidemia, genetics, secondary causes of dyslipidemia, biomarkers and ‘‘advanced lipid testing,’’ medical nutrition, physical activity, obesity, pharmacotherapy, statin safety, lipid-altering drug interactions, lipoprotein apheresis, dyslipidemia in children and adolescence, dyslipidemia in older individuals, race/ethnicity, and women, health information technology and electronic medical records, as well as investigational lipid-altering drugs in development. -

Geeta Sikand, MS
Geeta Sikand, MA, RDN, FAND, CDE, CLS, FNLA Geeta Sikand is an Associate Clinical Professor of Medicine in Cardiology at the University of California, Irvine School of Medicine. Registered Dietitian Nutritionist, Clinical Lipid Specialist, Certified Diabetes Educator, Fellow of the National Lipid Association and Fellow of the Academy of Nutrition and Dietetics. Ms Sikand is Director of Nutrition at University of California Irvine Preventive Cardiology Program. Ms Sikand is the recipient of the 2016 Lifetime Achievement Award from the Academy of Nutrition and Dietetics Member Interest Group, the 2013 Distinguished Service Award from the Dietitians’ in Health Care Communities Practice Group. For her research on the effectiveness of medical nutrition therapy in dyslipidemia, Geeta was the 2001 recipient of the Academy of Nutrition & Dietetics prestigious "Mary Huddleson Award” and the 1997 Excellence in Research Award from the California Academy of Nutrition and Dietetics. Ms Sikand is Chair, Medical Nutrition Therapy Effectiveness Expert Workgroup of the Academy of Nutrition & Dietetics and served as Expert Member of the Disorders of Lipid Metabolism Expert Panel. Ms Sikand has served on the Board of Directors of the American Heart Association and as Vice President of Public Policy for California Dietetic Association. As President of California Dietetic Association: Orange District and Chair of Southern California Legislative Steering Committee Ms Sikand organized the first ever onsite Congressional visit with Congressman Ron Packard to seek his support for the Medicare medical nutrition therapy legislation which passed in 1996. Ms Sikand is a member of the Pacific Lipid Association (PLA) and has served on the National Lipid Association (NLA) Expert Panel “Recommendations for Patient Centered Management of Dyslipidemia” published in December 2015. -

Watsonjn2018.Pdf (1.780Mb)
UNIVERSITY OF CENTRAL OKLAHOMA Edmond, Oklahoma Department of Biology Investigating Differential Gene Expression in vivo of Cardiac Birth Defects in an Avian Model of Maternal Phenylketonuria A THESIS SUBMITTED TO THE GRADUATE FACULTY In partial fulfillment of the requirements For the degree of MASTER OF SCIENCE IN BIOLOGY By Jamie N. Watson Edmond, OK June 5, 2018 J. Watson/Dr. Nikki Seagraves ii J. Watson/Dr. Nikki Seagraves Acknowledgements It is difficult to articulate the amount of gratitude I have for the support and encouragement I have received throughout my master’s thesis. Many people have added value and support to my life during this time. I am thankful for the education, experience, and friendships I have gained at the University of Central Oklahoma. First, I would like to thank Dr. Nikki Seagraves for her mentorship and friendship. I lucked out when I met her. I have enjoyed working on this project and I am very thankful for her support. I would like thank Thomas Crane for his support and patience throughout my master’s degree. I would like to thank Dr. Shannon Conley for her continued mentorship and support. I would like to thank Liz Bullen and Dr. Eric Howard for their training and help on this project. I would like to thank Kristy Meyer for her friendship and help throughout graduate school. I would like to thank my committee members Dr. Robert Brennan and Dr. Lilian Chooback for their advisement on this project. Also, I would like to thank the biology faculty and staff. I would like to thank the Seagraves lab members: Jailene Canales, Kayley Pate, Mckayla Muse, Grace Thetford, Kody Harvey, Jordan Guffey, and Kayle Patatanian for their hard work and support. -

743914V1.Full.Pdf
bioRxiv preprint doi: https://doi.org/10.1101/743914; this version posted August 24, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. 1 Cross-talks of glycosylphosphatidylinositol biosynthesis with glycosphingolipid biosynthesis 2 and ER-associated degradation 3 4 Yicheng Wang1,2, Yusuke Maeda1, Yishi Liu3, Yoko Takada2, Akinori Ninomiya1, Tetsuya 5 Hirata1,2,4, Morihisa Fujita3, Yoshiko Murakami1,2, Taroh Kinoshita1,2,* 6 7 1Research Institute for Microbial Diseases, Osaka University, Suita, Osaka 565-0871, Japan 8 2WPI Immunology Frontier Research Center, Osaka University, Suita, Osaka 565-0871, 9 Japan 10 3Key Laboratory of Carbohydrate Chemistry and Biotechnology, Ministry of Education, 11 School of Biotechnology, Jiangnan University, Wuxi, Jiangsu 214122, China 12 4Current address: Center for Highly Advanced Integration of Nano and Life Sciences (G- 13 CHAIN), Gifu University, 1-1 Yanagido, Gifu-City, Gifu 501-1193, Japan 14 15 *Correspondence and requests for materials should be addressed to T.K. (email: 16 [email protected]) 17 18 19 Glycosylphosphatidylinositol (GPI)-anchored proteins and glycosphingolipids interact with 20 each other in the mammalian plasma membranes, forming dynamic microdomains. How their 21 interaction starts in the cells has been unclear. Here, based on a genome-wide CRISPR-Cas9 22 genetic screen for genes required for GPI side-chain modification by galactose in the Golgi 23 apparatus, we report that b1,3-galactosyltransferase 4 (B3GALT4), also called GM1 24 ganglioside synthase, additionally functions in transferring galactose to the N- 25 acetylgalactosamine side-chain of GPI. -

Age-Driven Developmental Drift in the Pathogenesis of Idiopathic Pulmonary Fibrosis
BACK TO BASICS INTERSTITIAL LUNG DISEASES | Age-driven developmental drift in the pathogenesis of idiopathic pulmonary fibrosis Moisés Selman1, Carlos López-Otín2 and Annie Pardo3 Affiliations: 1Instituto Nacional de Enfermedades Respiratorias Ismael Cosío Villegas, Mexico city, Mexico. 2Departamento de Bioquímica y Biología Molecular, Facultad de Medicina, Instituto Universitario de Oncología, Universidad de Oviedo, Oviedo, Spain. 3Facultad de Ciencias, Universidad Nacional Autónoma de México, Mexico city, Mexico. Correspondence: Moisés Selman, Instituto Nacional de Enfermedades Respiratorias, Tlalpan 4502, CP 14080, México DF, México. E-mail: [email protected] ABSTRACT Idiopathic pulmonary fibrosis (IPF) is a progressive and usually lethal disease of unknown aetiology. A growing body of evidence supports that IPF represents an epithelial-driven process characterised by aberrant epithelial cell behaviour, fibroblast/myofibroblast activation and excessive accumulation of extracellular matrix with the subsequent destruction of the lung architecture. The mechanisms involved in the abnormal hyper-activation of the epithelium are unclear, but we propose that recapitulation of pathways and processes critical to embryological development associated with a tissue specific age-related stochastic epigenetic drift may be implicated. These pathways may also contribute to the distinctive behaviour of IPF fibroblasts. Genomic and epigenomic studies have revealed that wingless/ Int, sonic hedgehog and other developmental signalling pathways are reactivated and deregulated in IPF. Moreover, some of these pathways cross-talk with transforming growth factor-β activating a profibrotic feedback loop. The expression pattern of microRNAs is also dysregulated in IPF and exhibits a similar expression profile to embryonic lungs. In addition, senescence, a process usually associated with ageing, which occurs early in alveolar epithelial cells of IPF lungs, likely represents a conserved programmed developmental mechanism. -

SUPPLEMENTARY MATERIAL Bone Morphogenetic Protein 4 Promotes
www.intjdevbiol.com doi: 10.1387/ijdb.160040mk SUPPLEMENTARY MATERIAL corresponding to: Bone morphogenetic protein 4 promotes craniofacial neural crest induction from human pluripotent stem cells SUMIYO MIMURA, MIKA SUGA, KAORI OKADA, MASAKI KINEHARA, HIROKI NIKAWA and MIHO K. FURUE* *Address correspondence to: Miho Kusuda Furue. Laboratory of Stem Cell Cultures, National Institutes of Biomedical Innovation, Health and Nutrition, 7-6-8, Saito-Asagi, Ibaraki, Osaka 567-0085, Japan. Tel: 81-72-641-9819. Fax: 81-72-641-9812. E-mail: [email protected] Full text for this paper is available at: http://dx.doi.org/10.1387/ijdb.160040mk TABLE S1 PRIMER LIST FOR QRT-PCR Gene forward reverse AP2α AATTTCTCAACCGACAACATT ATCTGTTTTGTAGCCAGGAGC CDX2 CTGGAGCTGGAGAAGGAGTTTC ATTTTAACCTGCCTCTCAGAGAGC DLX1 AGTTTGCAGTTGCAGGCTTT CCCTGCTTCATCAGCTTCTT FOXD3 CAGCGGTTCGGCGGGAGG TGAGTGAGAGGTTGTGGCGGATG GAPDH CAAAGTTGTCATGGATGACC CCATGGAGAAGGCTGGGG MSX1 GGATCAGACTTCGGAGAGTGAACT GCCTTCCCTTTAACCCTCACA NANOG TGAACCTCAGCTACAAACAG TGGTGGTAGGAAGAGTAAAG OCT4 GACAGGGGGAGGGGAGGAGCTAGG CTTCCCTCCAACCAGTTGCCCCAAA PAX3 TTGCAATGGCCTCTCAC AGGGGAGAGCGCGTAATC PAX6 GTCCATCTTTGCTTGGGAAA TAGCCAGGTTGCGAAGAACT p75 TCATCCCTGTCTATTGCTCCA TGTTCTGCTTGCAGCTGTTC SOX9 AATGGAGCAGCGAAATCAAC CAGAGAGATTTAGCACACTGATC SOX10 GACCAGTACCCGCACCTG CGCTTGTCACTTTCGTTCAG Suppl. Fig. S1. Comparison of the gene expression profiles of the ES cells and the cells induced by NC and NC-B condition. Scatter plots compares the normalized expression of every gene on the array (refer to Table S3). The central line -
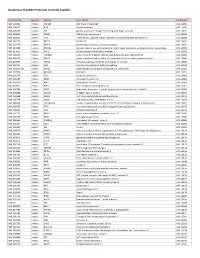
Quantigene Flowrna Probe Sets Currently Available
QuantiGene FlowRNA Probe Sets Currently Available Accession No. Species Symbol Gene Name Catalog No. NM_003452 Human ZNF189 zinc finger protein 189 VA1-10009 NM_000057 Human BLM Bloom syndrome VA1-10010 NM_005269 Human GLI glioma-associated oncogene homolog (zinc finger protein) VA1-10011 NM_002614 Human PDZK1 PDZ domain containing 1 VA1-10015 NM_003225 Human TFF1 Trefoil factor 1 (breast cancer, estrogen-inducible sequence expressed in) VA1-10016 NM_002276 Human KRT19 keratin 19 VA1-10022 NM_002659 Human PLAUR plasminogen activator, urokinase receptor VA1-10025 NM_017669 Human ERCC6L excision repair cross-complementing rodent repair deficiency, complementation group 6-like VA1-10029 NM_017699 Human SIDT1 SID1 transmembrane family, member 1 VA1-10032 NM_000077 Human CDKN2A cyclin-dependent kinase inhibitor 2A (melanoma, p16, inhibits CDK4) VA1-10040 NM_003150 Human STAT3 signal transducer and activator of transcripton 3 (acute-phase response factor) VA1-10046 NM_004707 Human ATG12 ATG12 autophagy related 12 homolog (S. cerevisiae) VA1-10047 NM_000737 Human CGB chorionic gonadotropin, beta polypeptide VA1-10048 NM_001017420 Human ESCO2 establishment of cohesion 1 homolog 2 (S. cerevisiae) VA1-10050 NM_197978 Human HEMGN hemogen VA1-10051 NM_001738 Human CA1 Carbonic anhydrase I VA1-10052 NM_000184 Human HBG2 Hemoglobin, gamma G VA1-10053 NM_005330 Human HBE1 Hemoglobin, epsilon 1 VA1-10054 NR_003367 Human PVT1 Pvt1 oncogene homolog (mouse) VA1-10061 NM_000454 Human SOD1 Superoxide dismutase 1, soluble (amyotrophic lateral sclerosis 1 (adult)) -

140503 IPF Signatures Supplement Withfigs Thorax
Supplementary material for Heterogeneous gene expression signatures correspond to distinct lung pathologies and biomarkers of disease severity in idiopathic pulmonary fibrosis Daryle J. DePianto1*, Sanjay Chandriani1⌘*, Alexander R. Abbas1, Guiquan Jia1, Elsa N. N’Diaye1, Patrick Caplazi1, Steven E. Kauder1, Sabyasachi Biswas1, Satyajit K. Karnik1#, Connie Ha1, Zora Modrusan1, Michael A. Matthay2, Jasleen Kukreja3, Harold R. Collard2, Jackson G. Egen1, Paul J. Wolters2§, and Joseph R. Arron1§ 1Genentech Research and Early Development, South San Francisco, CA 2Department of Medicine, University of California, San Francisco, CA 3Department of Surgery, University of California, San Francisco, CA ⌘Current address: Novartis Institutes for Biomedical Research, Emeryville, CA. #Current address: Gilead Sciences, Foster City, CA. *DJD and SC contributed equally to this manuscript §PJW and JRA co-directed this project Address correspondence to Paul J. Wolters, MD University of California, San Francisco Department of Medicine Box 0111 San Francisco, CA 94143-0111 [email protected] or Joseph R. Arron, MD, PhD Genentech, Inc. MS 231C 1 DNA Way South San Francisco, CA 94080 [email protected] 1 METHODS Human lung tissue samples Tissues were obtained at UCSF from clinical samples from IPF patients at the time of biopsy or lung transplantation. All patients were seen at UCSF and the diagnosis of IPF was established through multidisciplinary review of clinical, radiological, and pathological data according to criteria established by the consensus classification of the American Thoracic Society (ATS) and European Respiratory Society (ERS), Japanese Respiratory Society (JRS), and the Latin American Thoracic Association (ALAT) (ref. 5 in main text). Non-diseased normal lung tissues were procured from lungs not used by the Northern California Transplant Donor Network. -

Appendix 2. Significantly Differentially Regulated Genes in Term Compared with Second Trimester Amniotic Fluid Supernatant
Appendix 2. Significantly Differentially Regulated Genes in Term Compared With Second Trimester Amniotic Fluid Supernatant Fold Change in term vs second trimester Amniotic Affymetrix Duplicate Fluid Probe ID probes Symbol Entrez Gene Name 1019.9 217059_at D MUC7 mucin 7, secreted 424.5 211735_x_at D SFTPC surfactant protein C 416.2 206835_at STATH statherin 363.4 214387_x_at D SFTPC surfactant protein C 295.5 205982_x_at D SFTPC surfactant protein C 288.7 1553454_at RPTN repetin solute carrier family 34 (sodium 251.3 204124_at SLC34A2 phosphate), member 2 238.9 206786_at HTN3 histatin 3 161.5 220191_at GKN1 gastrokine 1 152.7 223678_s_at D SFTPA2 surfactant protein A2 130.9 207430_s_at D MSMB microseminoprotein, beta- 99.0 214199_at SFTPD surfactant protein D major histocompatibility complex, class II, 96.5 210982_s_at D HLA-DRA DR alpha 96.5 221133_s_at D CLDN18 claudin 18 94.4 238222_at GKN2 gastrokine 2 93.7 1557961_s_at D LOC100127983 uncharacterized LOC100127983 93.1 229584_at LRRK2 leucine-rich repeat kinase 2 HOXD cluster antisense RNA 1 (non- 88.6 242042_s_at D HOXD-AS1 protein coding) 86.0 205569_at LAMP3 lysosomal-associated membrane protein 3 85.4 232698_at BPIFB2 BPI fold containing family B, member 2 84.4 205979_at SCGB2A1 secretoglobin, family 2A, member 1 84.3 230469_at RTKN2 rhotekin 2 82.2 204130_at HSD11B2 hydroxysteroid (11-beta) dehydrogenase 2 81.9 222242_s_at KLK5 kallikrein-related peptidase 5 77.0 237281_at AKAP14 A kinase (PRKA) anchor protein 14 76.7 1553602_at MUCL1 mucin-like 1 76.3 216359_at D MUC7 mucin 7, -

Cholesterol-Dependent Increases in Glucosylceramide
Neuropharmacology 110 (2016) 458e469 Contents lists available at ScienceDirect Neuropharmacology journal homepage: www.elsevier.com/locate/neuropharm Cholesterol-dependent increases in glucosylceramide synthase activity in Niemann-Pick disease type C model cells: Abnormal trafficking of endogenously formed ceramide metabolites by inhibition of the enzyme Naohiro Hashimoto 1, Ikiru Matsumoto 1, Hiromasa Takahashi, Hitomi Ashikawa, * Hiroyuki Nakamura , Toshihiko Murayama Laboratory of Chemical Pharmacology, Graduate School of Pharmaceutical Sciences, Chiba University, Chiba 260-8675, Japan article info abstract Article history: Sphingolipids such as sphingomyelin and glycosphingolipids (GSLs) derived from glucosylceramide Received 1 February 2016 (GlcCer), in addition to cholesterol, accumulate in cells/neurons in Niemann-Pick disease type C (NPC). Received in revised form The activities of acid sphingomyelinase and lysosomal glucocerebrosidase (GCase), which degrade 10 August 2016 sphingomyelin and GlcCer, respectively, are down-regulated in NPC cells, however, changes in GlcCer Accepted 12 August 2016 synthase activity have not yet been elucidated. We herein demonstrated for the first time that GlcCer Available online 15 August 2016 synthase activity for the fluorescent ceramide, 4-nitrobenzo-2-oxa-1,3-diazole-labeled C6-ceramide À À (NBD-ceramide) increased in intact NPC1( / ) cells and cell lysates without affecting the protein levels. Keywords: (À/À) fl Niemann-Pick disease type C In NBD-ceramide-labeled NPC1 cells, NBD- uorescence preferentially accumulated in the Golgi Cholesterol complex and vesicular specks in the cytoplasm 40 and 150 min, respectively, after labeling, while a Glucosylceramide synthase treatment for 48 h with the GlcCer synthase inhibitors, N-butyldeoxynojirimycin (NB-DNJ) and 1-phenyl- Sphingomyelin 2-palmitoylamino-3-morpholino-1-propanol, accelerated the appearance of vesicular specks emitting À À Vesicular trafficking NBD-fluorescence within 40 min. -

Supplementary Methods
Supplementary methods Human lung tissues and tissue microarray (TMA) All human tissues were obtained from the Lung Cancer Specialized Program of Research Excellence (SPORE) Tissue Bank at the M.D. Anderson Cancer Center (Houston, TX). A collection of 26 lung adenocarcinomas and 24 non-tumoral paired tissues were snap-frozen and preserved in liquid nitrogen for total RNA extraction. For each tissue sample, the percentage of malignant tissue was calculated and the cellular composition of specimens was determined by histological examination (I.I.W.) following Hematoxylin-Eosin (H&E) staining. All malignant samples retained contained more than 50% tumor cells. Specimens resected from NSCLC stages I-IV patients who had no prior chemotherapy or radiotherapy were used for TMA analysis by immunohistochemistry. Patients who had smoked at least 100 cigarettes in their lifetime were defined as smokers. Samples were fixed in formalin, embedded in paraffin, stained with H&E, and reviewed by an experienced pathologist (I.I.W.). The 413 tissue specimens collected from 283 patients included 62 normal bronchial epithelia, 61 bronchial hyperplasias (Hyp), 15 squamous metaplasias (SqM), 9 squamous dysplasias (Dys), 26 carcinomas in situ (CIS), as well as 98 squamous cell carcinomas (SCC) and 141 adenocarcinomas. Normal bronchial epithelia, hyperplasia, squamous metaplasia, dysplasia, CIS, and SCC were considered to represent different steps in the development of SCCs. All tumors and lesions were classified according to the World Health Organization (WHO) 2004 criteria. The TMAs were prepared with a manual tissue arrayer (Advanced Tissue Arrayer ATA100, Chemicon International, Temecula, CA) using 1-mm-diameter cores in triplicate for tumors and 1.5 to 2-mm cores for normal epithelial and premalignant lesions. -

Whole-Exome Sequencing of Metastatic Cancer and Biomarkers of Treatment Response
Supplementary Online Content Beltran H, Eng K, Mosquera JM, et al. Whole-exome sequencing of metastatic cancer and biomarkers of treatment response. JAMA Oncol. Published online May 28, 2015. doi:10.1001/jamaoncol.2015.1313 eMethods eFigure 1. A schematic of the IPM Computational Pipeline eFigure 2. Tumor purity analysis eFigure 3. Tumor purity estimates from Pathology team versus computationally (CLONET) estimated tumor purities values for frozen tumor specimens (Spearman correlation 0.2765327, p- value = 0.03561) eFigure 4. Sequencing metrics Fresh/frozen vs. FFPE tissue eFigure 5. Somatic copy number alteration profiles by tumor type at cytogenetic map location resolution; for each cytogenetic map location the mean genes aberration frequency is reported eFigure 6. The 20 most frequently aberrant genes with respect to copy number gains/losses detected per tumor type eFigure 7. Top 50 genes with focal and large scale copy number gains (A) and losses (B) across the cohort eFigure 8. Summary of total number of copy number alterations across PM tumors eFigure 9. An example of tumor evolution looking at serial biopsies from PM222, a patient with metastatic bladder carcinoma eFigure 10. PM12 somatic mutations by coverage and allele frequency (A) and (B) mutation correlation between primary (y- axis) and brain metastasis (x-axis) eFigure 11. Point mutations across 5 metastatic sites of a 55 year old patient with metastatic prostate cancer at time of rapid autopsy eFigure 12. CT scans from patient PM137, a patient with recurrent platinum refractory metastatic urothelial carcinoma eFigure 13. Tracking tumor genomics between primary and metastatic samples from patient PM12 eFigure 14.