AMY1A ELISA Kit (Human) (OKEH01050)
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Structural Forms of the Human Amylase Locus and Their Relationships to Snps, Haplotypes, and Obesity
Structural Forms of the Human Amylase Locus and Their Relationships to SNPs, Haplotypes, and Obesity The Harvard community has made this article openly available. Please share how this access benefits you. Your story matters Citation Usher, Christina Leigh. 2015. Structural Forms of the Human Amylase Locus and Their Relationships to SNPs, Haplotypes, and Obesity. Doctoral dissertation, Harvard University, Graduate School of Arts & Sciences. Citable link http://nrs.harvard.edu/urn-3:HUL.InstRepos:17467224 Terms of Use This article was downloaded from Harvard University’s DASH repository, and is made available under the terms and conditions applicable to Other Posted Material, as set forth at http:// nrs.harvard.edu/urn-3:HUL.InstRepos:dash.current.terms-of- use#LAA Structural forms of the human amylase locus and their relationships to SNPs, haplotypes, and obesity A dissertation presented by Christina Leigh Usher to The Division of Medical Sciences in partial fulfillment of the requirements for the degree of Doctor of Philosophy in the subject of Genetics and Genomics Harvard University Cambridge, Massachusetts March 2015 © 2015 Christina Leigh Usher All rights reserved. Dissertation Advisor: Professor Steven McCarroll Christina Leigh Usher Structural forms of the human amylase locus and their relationships to SNPs, haplotypes, and obesity Abstract Hundreds of human genes reside in structurally complex loci that elude molecular analysis and assessment in genome-wide association studies (GWAS). One such locus contains the three different amylase genes (AMY2B, AMY2A, and AMY1) responsible for digesting starch into sugar. The copy number of AMY1 is reported to be the genome’s largest influence on obesity, yet has gone undetected in GWAS. -

Renal Cell Neoplasms Contain Shared Tumor Type–Specific Copy Number Variations
The American Journal of Pathology, Vol. 180, No. 6, June 2012 Copyright © 2012 American Society for Investigative Pathology. Published by Elsevier Inc. All rights reserved. http://dx.doi.org/10.1016/j.ajpath.2012.01.044 Tumorigenesis and Neoplastic Progression Renal Cell Neoplasms Contain Shared Tumor Type–Specific Copy Number Variations John M. Krill-Burger,* Maureen A. Lyons,*† The annual incidence of renal cell carcinoma (RCC) has Lori A. Kelly,*† Christin M. Sciulli,*† increased steadily in the United States for the past three Patricia Petrosko,*† Uma R. Chandran,†‡ decades, with approximately 58,000 new cases diag- 1,2 Michael D. Kubal,§ Sheldon I. Bastacky,*† nosed in 2010, representing 3% of all malignancies. Anil V. Parwani,*†‡ Rajiv Dhir,*†‡ and Treatment of RCC is complicated by the fact that it is not a single disease but composes multiple tumor types with William A. LaFramboise*†‡ different morphological characteristics, clinical courses, From the Departments of Pathology* and Biomedical and outcomes (ie, clear-cell carcinoma, 82% of RCC ‡ Informatics, University of Pittsburgh, Pittsburgh, Pennsylvania; cases; type 1 or 2 papillary tumors, 11% of RCC cases; † the University of Pittsburgh Cancer Institute, Pittsburgh, chromophobe tumors, 5% of RCC cases; and collecting § Pennsylvania; and Life Technologies, Carlsbad, California duct carcinoma, approximately 1% of RCC cases).2,3 Benign renal neoplasms are subdivided into papillary adenoma, renal oncocytoma, and metanephric ade- Copy number variant (CNV) analysis was performed on noma.2,3 Treatment of RCC often involves surgical resec- renal cell carcinoma (RCC) specimens (chromophobe, tion of a large renal tissue component or removal of the clear cell, oncocytoma, papillary type 1, and papillary entire affected kidney because of the relatively large size of type 2) using high-resolution arrays (1.85 million renal tumors on discovery and the availability of a life-sus- probes). -

Salivary Alpha Amylase (AMY1C) (NM 001008219) Human Tagged ORF Clone Product Data
OriGene Technologies, Inc. 9620 Medical Center Drive, Ste 200 Rockville, MD 20850, US Phone: +1-888-267-4436 [email protected] EU: [email protected] CN: [email protected] Product datasheet for RG215827 Salivary alpha amylase (AMY1C) (NM_001008219) Human Tagged ORF Clone Product data: Product Type: Expression Plasmids Product Name: Salivary alpha amylase (AMY1C) (NM_001008219) Human Tagged ORF Clone Tag: TurboGFP Symbol: AMY1C Synonyms: AMY1 Vector: pCMV6-AC-GFP (PS100010) E. coli Selection: Ampicillin (100 ug/mL) Cell Selection: Neomycin This product is to be used for laboratory only. Not for diagnostic or therapeutic use. View online » ©2021 OriGene Technologies, Inc., 9620 Medical Center Drive, Ste 200, Rockville, MD 20850, US 1 / 5 Salivary alpha amylase (AMY1C) (NM_001008219) Human Tagged ORF Clone – RG215827 ORF Nucleotide >RG215827 representing NM_001008219 Sequence: Red=Cloning site Blue=ORF Green=Tags(s) TTTTGTAATACGACTCACTATAGGGCGGCCGGGAATTCGTCGACTGGATCCGGTACCGAGGAGATCTGCC GCCGCGATCGCC ATGAAGCTCTTTTGGTTGCTTTTCACCATTGGGTTCTGCTGGGCTCAGTATTCCTCAAATACACAACAAG GACGAACATCTATTGTTCATCTGTTTGAATGGCGATGGGTTGATATTGCTCTTGAATGTGAGCGATATTT AGCTCCCAAGGGATTTGGAGGGGTTCAGGTCTCTCCACCAAATGAAAATGTTGCCATTCACAACCCTTTC AGACCTTGGTGGGAAAGATACCAACCAGTTAGCTATAAATTATGCACAAGATCTGGAAATGAAGATGAAT TTAGAAACATGGTGACTAGATGCAACAATGTTGGGGTTCGTATTTATGTGGATGCTGTAATTAATCATAT GTGTGGTAATGCTGTGAGTGCAGGAACAAGCAGTACCTGTGGAAGTTACTTCAACCCTGGAAGTAGGGAC TTTCCAGCAGTCCCATATTCTGGATGGGATTTTAATGATGGTAAATGTAAAACTGGAAGTGGAGATATCG AGAACTATAATGATGCTACTCAGGTCAGAGATTGTCGTCTGTCTGGTCTTCTCGATCTTGCACTGGGGAA -

Chuanxiong Rhizoma Compound on HIF-VEGF Pathway and Cerebral Ischemia-Reperfusion Injury’S Biological Network Based on Systematic Pharmacology
ORIGINAL RESEARCH published: 25 June 2021 doi: 10.3389/fphar.2021.601846 Exploring the Regulatory Mechanism of Hedysarum Multijugum Maxim.-Chuanxiong Rhizoma Compound on HIF-VEGF Pathway and Cerebral Ischemia-Reperfusion Injury’s Biological Network Based on Systematic Pharmacology Kailin Yang 1†, Liuting Zeng 1†, Anqi Ge 2†, Yi Chen 1†, Shanshan Wang 1†, Xiaofei Zhu 1,3† and Jinwen Ge 1,4* Edited by: 1 Takashi Sato, Key Laboratory of Hunan Province for Integrated Traditional Chinese and Western Medicine on Prevention and Treatment of 2 Tokyo University of Pharmacy and Life Cardio-Cerebral Diseases, Hunan University of Chinese Medicine, Changsha, China, Galactophore Department, The First 3 Sciences, Japan Hospital of Hunan University of Chinese Medicine, Changsha, China, School of Graduate, Central South University, Changsha, China, 4Shaoyang University, Shaoyang, China Reviewed by: Hui Zhao, Capital Medical University, China Background: Clinical research found that Hedysarum Multijugum Maxim.-Chuanxiong Maria Luisa Del Moral, fi University of Jaén, Spain Rhizoma Compound (HCC) has de nite curative effect on cerebral ischemic diseases, *Correspondence: such as ischemic stroke and cerebral ischemia-reperfusion injury (CIR). However, its Jinwen Ge mechanism for treating cerebral ischemia is still not fully explained. [email protected] †These authors share first authorship Methods: The traditional Chinese medicine related database were utilized to obtain the components of HCC. The Pharmmapper were used to predict HCC’s potential targets. Specialty section: The CIR genes were obtained from Genecards and OMIM and the protein-protein This article was submitted to interaction (PPI) data of HCC’s targets and IS genes were obtained from String Ethnopharmacology, a section of the journal database. -

Role of Amylase in Ovarian Cancer Mai Mohamed University of South Florida, [email protected]
University of South Florida Scholar Commons Graduate Theses and Dissertations Graduate School July 2017 Role of Amylase in Ovarian Cancer Mai Mohamed University of South Florida, [email protected] Follow this and additional works at: http://scholarcommons.usf.edu/etd Part of the Pathology Commons Scholar Commons Citation Mohamed, Mai, "Role of Amylase in Ovarian Cancer" (2017). Graduate Theses and Dissertations. http://scholarcommons.usf.edu/etd/6907 This Dissertation is brought to you for free and open access by the Graduate School at Scholar Commons. It has been accepted for inclusion in Graduate Theses and Dissertations by an authorized administrator of Scholar Commons. For more information, please contact [email protected]. Role of Amylase in Ovarian Cancer by Mai Mohamed A dissertation submitted in partial fulfillment of the requirements for the degree of Doctor of Philosophy Department of Pathology and Cell Biology Morsani College of Medicine University of South Florida Major Professor: Patricia Kruk, Ph.D. Paula C. Bickford, Ph.D. Meera Nanjundan, Ph.D. Marzenna Wiranowska, Ph.D. Lauri Wright, Ph.D. Date of Approval: June 29, 2017 Keywords: ovarian cancer, amylase, computational analyses, glycocalyx, cellular invasion Copyright © 2017, Mai Mohamed Dedication This dissertation is dedicated to my parents, Ahmed and Fatma, who have always stressed the importance of education, and, throughout my education, have been my strongest source of encouragement and support. They always believed in me and I am eternally grateful to them. I would also like to thank my brothers, Mohamed and Hussien, and my sister, Mariam. I would also like to thank my husband, Ahmed. -

Early Growth Response 1 Regulates Hematopoietic Support and Proliferation in Human Primary Bone Marrow Stromal Cells
Hematopoiesis SUPPLEMENTARY APPENDIX Early growth response 1 regulates hematopoietic support and proliferation in human primary bone marrow stromal cells Hongzhe Li, 1,2 Hooi-Ching Lim, 1,2 Dimitra Zacharaki, 1,2 Xiaojie Xian, 2,3 Keane J.G. Kenswil, 4 Sandro Bräunig, 1,2 Marc H.G.P. Raaijmakers, 4 Niels-Bjarne Woods, 2,3 Jenny Hansson, 1,2 and Stefan Scheding 1,2,5 1Division of Molecular Hematology, Department of Laboratory Medicine, Lund University, Lund, Sweden; 2Lund Stem Cell Center, Depart - ment of Laboratory Medicine, Lund University, Lund, Sweden; 3Division of Molecular Medicine and Gene Therapy, Department of Labora - tory Medicine, Lund University, Lund, Sweden; 4Department of Hematology, Erasmus MC Cancer Institute, Rotterdam, the Netherlands and 5Department of Hematology, Skåne University Hospital Lund, Skåne, Sweden ©2020 Ferrata Storti Foundation. This is an open-access paper. doi:10.3324/haematol. 2019.216648 Received: January 14, 2019. Accepted: July 19, 2019. Pre-published: August 1, 2019. Correspondence: STEFAN SCHEDING - [email protected] Li et al.: Supplemental data 1. Supplemental Materials and Methods BM-MNC isolation Bone marrow mononuclear cells (BM-MNC) from BM aspiration samples were isolated by density gradient centrifugation (LSM 1077 Lymphocyte, PAA, Pasching, Austria) either with or without prior incubation with RosetteSep Human Mesenchymal Stem Cell Enrichment Cocktail (STEMCELL Technologies, Vancouver, Canada) for lineage depletion (CD3, CD14, CD19, CD38, CD66b, glycophorin A). BM-MNCs from fetal long bones and adult hip bones were isolated as reported previously 1 by gently crushing bones (femora, tibiae, fibulae, humeri, radii and ulna) in PBS+0.5% FCS subsequent passing of the cell suspension through a 40-µm filter. -

Research Article Sex Difference of Ribosome in Stroke-Induced Peripheral Immunosuppression by Integrated Bioinformatics Analysis
Hindawi BioMed Research International Volume 2020, Article ID 3650935, 15 pages https://doi.org/10.1155/2020/3650935 Research Article Sex Difference of Ribosome in Stroke-Induced Peripheral Immunosuppression by Integrated Bioinformatics Analysis Jian-Qin Xie ,1,2,3 Ya-Peng Lu ,1,3 Hong-Li Sun ,1,3 Li-Na Gao ,2,3 Pei-Pei Song ,2,3 Zhi-Jun Feng ,3 and Chong-Ge You 2,3 1Department of Anesthesiology, Lanzhou University Second Hospital, Lanzhou, Gansu 730030, China 2Laboratory Medicine Center, Lanzhou University Second Hospital, Lanzhou, Gansu 730030, China 3The Second Clinical Medical College of Lanzhou University, Lanzhou, Gansu 730030, China Correspondence should be addressed to Chong-Ge You; [email protected] Received 13 April 2020; Revised 8 October 2020; Accepted 18 November 2020; Published 3 December 2020 Academic Editor: Rudolf K. Braun Copyright © 2020 Jian-Qin Xie et al. This is an open access article distributed under the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited. Ischemic stroke (IS) greatly threatens human health resulting in high mortality and substantial loss of function. Recent studies have shown that the outcome of IS has sex specific, but its mechanism is still unclear. This study is aimed at identifying the sexually dimorphic to peripheral immune response in IS progression, predicting potential prognostic biomarkers that can lead to sex- specific outcome, and revealing potential treatment targets. Gene expression dataset GSE37587, including 68 peripheral whole blood samples which were collected within 24 hours from known onset of symptom and again at 24-48 hours after onset (20 women and 14 men), was downloaded from the Gene Expression Omnibus (GEO) datasets. -

The Genome-Wide Landscape of Copy Number Variations in the MUSGEN Study Provides Evidence for a Founder Effect in the Isolated Finnish Population
European Journal of Human Genetics (2013) 21, 1411–1416 & 2013 Macmillan Publishers Limited All rights reserved 1018-4813/13 www.nature.com/ejhg ARTICLE The genome-wide landscape of copy number variations in the MUSGEN study provides evidence for a founder effect in the isolated Finnish population Chakravarthi Kanduri1, Liisa Ukkola-Vuoti1, Jaana Oikkonen1, Gemma Buck2, Christine Blancher2, Pirre Raijas3, Kai Karma4, Harri La¨hdesma¨ki5 and Irma Ja¨rvela¨*,1 Here we characterized the genome-wide architecture of copy number variations (CNVs) in 286 healthy, unrelated Finnish individuals belonging to the MUSGEN study, where molecular background underlying musical aptitude and related traits are studied. By using Illumina HumanOmniExpress-12v.1.0 beadchip, we identified 5493 CNVs that were spread across 467 different cytogenetic regions, spanning a total size of 287.83 Mb (B9.6% of the human genome). Merging the overlapping CNVs across samples resulted in 999 discrete copy number variable regions (CNVRs), of which B6.9% were putatively novel. The average number of CNVs per person was 20, whereas the average size of CNV per locus was 52.39 kb. Large CNVs (41 Mb) were present in 4% of the samples. The proportion of homozygous deletions in this data set (B12.4%) seemed to be higher when compared with three other populations. Interestingly, several CNVRs were significantly enriched in this sample set, whereas several others were totally depleted. For example, a CNVR at chr2p22.1 intersecting GALM was more common in this population (P ¼ 3.3706 Â 10 À44) than in African and other European populations. The enriched CNVRs, however, showed no significant association with music-related phenotypes. -

(12) Patent Application Publication (10) Pub. No.: US 2001/0034023 A1 Stanton, JR
US 2001.0034O23A1. (19) United States (12) Patent Application Publication (10) Pub. No.: US 2001/0034023 A1 Stanton, JR. et al. (43) Pub. Date: Oct. 25, 2001 (54) GENE SEQUENCE VARIATIONS WITH Related U.S. Application Data UTILITY IN DETERMINING THE (63) Continuation-in-part of application No. 09/710,467, TREATMENT OF DISEASE, INGENES filed on Nov. 8, 2000, which is a continuation-in-part RELATING TO DRUG PROCESSING of application No. 09/696,482, filed on Oct. 24, 2000. Non-provisional of provisional application No. 60/131,334, filed on Apr. 26, 1999. Non-provisional (76) Inventors: Vincent P. Stanton JR., Belmont, MA of provisional application No. 60/139,440, filed on (US); Martin Zillmann, Shrewsbury, Jun. 15, 1999. MA (US) (30) Foreign Application Priority Data Correspondence Address: Jan. 20, 2000 (WO)........................... PCT/USOO/O1392 EDWARD O. KRUESSER BROBECK PHLEGER & HARRISON Publication Classification 12390 EL CAMINO REAL (51) Int. Cl. ............................. C12O 1/68; G06F 19/00 SAN DIEGO, CA 92130 (US) (52) U.S. Cl. ................................................... 435/6; 702/20 (57) ABSTRACT (21) Appl. No.: 09/733,000 Methods for identifying and utilizing variances in genes relating to efficacy and Safety of medical therapy and other aspects of medical therapy are described, including methods (22) Filed: Dec. 7, 2000 for Selecting an effective treatment. US 2001/0034023 A1 Oct. 25, 2001 GENE SEQUENCE WARIATIONS WITH UTILITY 0005 For some drugs, over 90% of the measurable IN DETERMINING THE TREATMENT OF interSubject variation in Selected pharmacokinetic param DISEASE, INGENES RELATING TO DRUG eters has been shown to be heritable. For a limited number PROCESSING of drugs, DNA sequence variances have been identified in Specific genes that are involved in drug action or metabo RELATED APPLICATIONS lism, and these variances have been shown to account for the 0001. -
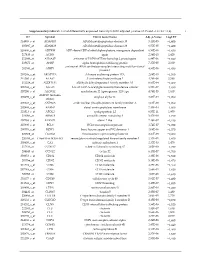
Supplementary Table S1. List of Differentially Expressed
Supplementary table S1. List of differentially expressed transcripts (FDR adjusted p‐value < 0.05 and −1.4 ≤ FC ≥1.4). 1 ID Symbol Entrez Gene Name Adj. p‐Value Log2 FC 214895_s_at ADAM10 ADAM metallopeptidase domain 10 3,11E‐05 −1,400 205997_at ADAM28 ADAM metallopeptidase domain 28 6,57E‐05 −1,400 220606_s_at ADPRM ADP‐ribose/CDP‐alcohol diphosphatase, manganese dependent 6,50E‐06 −1,430 217410_at AGRN agrin 2,34E‐10 1,420 212980_at AHSA2P activator of HSP90 ATPase homolog 2, pseudogene 6,44E‐06 −1,920 219672_at AHSP alpha hemoglobin stabilizing protein 7,27E‐05 2,330 aminoacyl tRNA synthetase complex interacting multifunctional 202541_at AIMP1 4,91E‐06 −1,830 protein 1 210269_s_at AKAP17A A‐kinase anchoring protein 17A 2,64E‐10 −1,560 211560_s_at ALAS2 5ʹ‐aminolevulinate synthase 2 4,28E‐06 3,560 212224_at ALDH1A1 aldehyde dehydrogenase 1 family member A1 8,93E‐04 −1,400 205583_s_at ALG13 ALG13 UDP‐N‐acetylglucosaminyltransferase subunit 9,50E‐07 −1,430 207206_s_at ALOX12 arachidonate 12‐lipoxygenase, 12S type 4,76E‐05 1,630 AMY1C (includes 208498_s_at amylase alpha 1C 3,83E‐05 −1,700 others) 201043_s_at ANP32A acidic nuclear phosphoprotein 32 family member A 5,61E‐09 −1,760 202888_s_at ANPEP alanyl aminopeptidase, membrane 7,40E‐04 −1,600 221013_s_at APOL2 apolipoprotein L2 6,57E‐11 1,600 219094_at ARMC8 armadillo repeat containing 8 3,47E‐08 −1,710 207798_s_at ATXN2L ataxin 2 like 2,16E‐07 −1,410 215990_s_at BCL6 BCL6 transcription repressor 1,74E‐07 −1,700 200776_s_at BZW1 basic leucine zipper and W2 domains 1 1,09E‐06 −1,570 222309_at -

Cancer Sequencing Service Data File Formats File Format V2.4 Software V2.4 December 2012
Cancer Sequencing Service Data File Formats File format v2.4 Software v2.4 December 2012 CGA Tools, cPAL, and DNB are trademarks of Complete Genomics, Inc. in the US and certain other countries. All other trademarks are the property of their respective owners. Disclaimer of Warranties. COMPLETE GENOMICS, INC. PROVIDES THESE DATA IN GOOD FAITH TO THE RECIPIENT “AS IS.” COMPLETE GENOMICS, INC. MAKES NO REPRESENTATION OR WARRANTY, EXPRESS OR IMPLIED, INCLUDING WITHOUT LIMITATION ANY IMPLIED WARRANTY OF MERCHANTABILITY OR FITNESS FOR A PARTICULAR PURPOSE OR USE, OR ANY OTHER STATUTORY WARRANTY. COMPLETE GENOMICS, INC. ASSUMES NO LEGAL LIABILITY OR RESPONSIBILITY FOR ANY PURPOSE FOR WHICH THE DATA ARE USED. Any permitted redistribution of the data should carry the Disclaimer of Warranties provided above. Data file formats are expected to evolve over time. Backward compatibility of any new file format is not guaranteed. Complete Genomics data is for Research Use Only and not for use in the treatment or diagnosis of any human subject. Information, descriptions and specifications in this publication are subject to change without notice. Copyright © 2011-2012 Complete Genomics Incorporated. All rights reserved. RM_DFFCS_2.4-01 Table of Contents Table of Contents Preface ...........................................................................................................................................................................................1 Conventions ................................................................................................................................................................................................. -

Mapping Mrna Libraries
Mapping mRNA libraries STAR-2.5.2b was used with options shown below. Index generation: $STAR --runThreadN 16 \ --runMode genomeGenerate \ --genomeDir $IndexPref \ --genomeFastaFiles $GenomePref/Mus_musculus.GRCm38.dna.primary_assembly.fa \ --sjdbGTFfile $AnnoPref/Mus_musculus.GRCm38.86.gtf \ --sjdbOverhang 100 Aligning and counting reads with the genes (annotated transcripts): $STAR --runThreadN 8 \ --runMode alignReads \ --genomeDir $IndexPref \ --readFilesIn $FileOne $FileTwo \ --outFileNamePrefix $OutDir \ --outFilterType BySJout \ --outFilterMismatchNoverLmax 0.05 \ --alignSJoverhangMin 8 \ --alignSJDBoverhangMin 1 \ --alignIntronMin 20 \ --alignMatesGapMax 1000000 \ --outSAMtype BAM SortedByCoordinate Complex homology groups discussed in this paper Modencode project declares more homology pairs than homologene. When we combine homology pairs from different sources there exists a potential for amplifying the number of incorrect homology pairs, therefore here we review the complex homology groups that are mentioned in the paper. Homology of genes is defined on the basis of the evolutionary history of genomes which cannot be easily assessed by looking at the data from two species, but we can compare clues from synteny and expression profiles. While we see many differences in expression profiles among members of the same homology group, given a choice of different sets of homology pairs we prefer one that explains more cases of high gene expressions with homology. These homology groups were included in our specific findings while they are not consistently identified by modencode and homologene: • Alkaline phosphatases • Amylases • Kallikrein 1-related peptidases • Defensins with high expression in mouse skin and vagina More authoritative methods to determine homology collect information from many species. However, we use a method which allows to choose most plausible homology when we have two-three alternatives.