Package 'Rfast'
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

1 Introduction
TR:UPM-ETSINF/DIA/2015-1 1 Univariate and bivariate truncated von Mises distributions Pablo Fernandez-Gonzalez, Concha Bielza, Pedro Larra~naga Department of Artificial Intelligence, Universidad Polit´ecnica de Madrid Abstract: In this article we study the univariate and bivariate truncated von Mises distribution, as a generalization of the von Mises distribution (Jupp and Mardia (1989)), (Mardia and Jupp (2000)). This implies the addition of two or four new truncation parameters in the univariate and, bivariate cases, respectively. The re- sults include the definition, properties of the distribution and maximum likelihood estimators for the univariate and bivariate cases. Additionally, the analysis of the bivariate case shows how the conditional distribution is a truncated von Mises dis- tribution, whereas the marginal distribution that generalizes the distribution intro- duced in Singh (2002). From the viewpoint of applications, we test the distribution with simulated data, as well as with data regarding leaf inclination angles (Bowyer and Danson. (2005)) and dihedral angles in protein chains (Murzin AG (1995)). This research aims to assert this probability distribution as a potential option for modelling or simulating any kind of phenomena where circular distributions are applicable. Key words and phrases: Angular probability distributions, Directional statistics, von Mises distribution, Truncated probability distributions. 1 Introduction The von Mises distribution has received undisputed attention in the field of di- rectional statistics (Jupp and Mardia (1989)) and in other areas like supervised classification (Lopez-Cruz et al. (2013)). Thanks to desirable properties such as its symmetry, mathematical tractability and convergence to the wrapped nor- mal distribution (Mardia and Jupp (2000)) for high concentrations, it is a viable option for many statistical analyses. -
![Arxiv:1911.00962V1 [Cs.CV] 3 Nov 2019 Though the Label of Pose Itself Is Discrete](https://docslib.b-cdn.net/cover/7665/arxiv-1911-00962v1-cs-cv-3-nov-2019-though-the-label-of-pose-itself-is-discrete-507665.webp)
Arxiv:1911.00962V1 [Cs.CV] 3 Nov 2019 Though the Label of Pose Itself Is Discrete
Conservative Wasserstein Training for Pose Estimation Xiaofeng Liu1;2†∗, Yang Zou1y, Tong Che3y, Peng Ding4, Ping Jia4, Jane You5, B.V.K. Vijaya Kumar1 1Carnegie Mellon University; 2Harvard University; 3MILA 4CIOMP, Chinese Academy of Sciences; 5The Hong Kong Polytechnic University yContribute equally ∗Corresponding to: [email protected] Pose with true label Probability of Softmax prediction Abstract Expected probability (i.e., 1 in one-hot case) σ푁−1 푡 푠 푡푗∗ of each pose ( 푖=0 푠푖 = 1) 푗∗ 3 푠 푠 2 푠3 2 푠푗∗ 푠푗∗ This paper targets the task with discrete and periodic 푠1 푠1 class labels (e:g:; pose/orientation estimation) in the con- Same Cross-entropy text of deep learning. The commonly used cross-entropy or 푠0 푠0 regression loss is not well matched to this problem as they 푠푁−1 ignore the periodic nature of the labels and the class simi- More ideal Inferior 푠푁−1 larity, or assume labels are continuous value. We propose to Distribution 푠푁−2 Distribution 푠푁−2 incorporate inter-class correlations in a Wasserstein train- Figure 1. The limitation of CE loss for pose estimation. The ing framework by pre-defining (i:e:; using arc length of a ground truth direction of the car is tj ∗. Two possible softmax circle) or adaptively learning the ground metric. We extend predictions (green bar) of the pose estimator have the same proba- the ground metric as a linear, convex or concave increasing bility at tj ∗ position. Therefore, both predicted distributions have function w:r:t: arc length from an optimization perspective. the same CE loss. -

Chapter 2 Parameter Estimation for Wrapped Distributions
ABSTRACT RAVINDRAN, PALANIKUMAR. Bayesian Analysis of Circular Data Using Wrapped Dis- tributions. (Under the direction of Associate Professor Sujit K. Ghosh). Circular data arise in a number of different areas such as geological, meteorolog- ical, biological and industrial sciences. We cannot use standard statistical techniques to model circular data, due to the circular geometry of the sample space. One of the com- mon methods used to analyze such data is the wrapping approach. Using the wrapping approach, we assume that, by wrapping a probability distribution from the real line onto the circle, we obtain the probability distribution for circular data. This approach creates a vast class of probability distributions that are flexible to account for different features of circular data. However, the likelihood-based inference for such distributions can be very complicated and computationally intensive. The EM algorithm used to compute the MLE is feasible, but is computationally unsatisfactory. Instead, we use Markov Chain Monte Carlo (MCMC) methods with a data augmentation step, to overcome such computational difficul- ties. Given a probability distribution on the circle, we assume that the original distribution was distributed on the real line, and then wrapped onto the circle. If we can unwrap the distribution off the circle and obtain a distribution on the real line, then the standard sta- tistical techniques for data on the real line can be used. Our proposed methods are flexible and computationally efficient to fit a wide class of wrapped distributions. Furthermore, we can easily compute the usual summary statistics. We present extensive simulation studies to validate the performance of our method. -

A Compendium of Conjugate Priors
A Compendium of Conjugate Priors Daniel Fink Environmental Statistics Group Department of Biology Montana State Univeristy Bozeman, MT 59717 May 1997 Abstract This report reviews conjugate priors and priors closed under sampling for a variety of data generating processes where the prior distributions are univariate, bivariate, and multivariate. The effects of transformations on conjugate prior relationships are considered and cases where conjugate prior relationships can be applied under transformations are identified. Univariate and bivariate prior relationships are verified using Monte Carlo methods. Contents 1 Introduction Experimenters are often in the position of having had collected some data from which they desire to make inferences about the process that produced that data. Bayes' theorem provides an appealing approach to solving such inference problems. Bayes theorem, π(θ) L(θ x ; : : : ; x ) g(θ x ; : : : ; x ) = j 1 n (1) j 1 n π(θ) L(θ x ; : : : ; x )dθ j 1 n is commonly interpreted in the following wayR. We want to make some sort of inference on the unknown parameter(s), θ, based on our prior knowledge of θ and the data collected, x1; : : : ; xn . Our prior knowledge is encapsulated by the probability distribution on θ; π(θ). The data that has been collected is combined with our prior through the likelihood function, L(θ x ; : : : ; x ) . The j 1 n normalized product of these two components yields a probability distribution of θ conditional on the data. This distribution, g(θ x ; : : : ; x ) , is known as the posterior distribution of θ. Bayes' j 1 n theorem is easily extended to cases where is θ multivariate, a vector of parameters. -

Recursive Bayesian Filtering in Circular State Spaces
Recursive Bayesian Filtering in Circular State Spaces Gerhard Kurza, Igor Gilitschenskia, Uwe D. Hanebecka aIntelligent Sensor-Actuator-Systems Laboratory (ISAS) Institute for Anthropomatics and Robotics Karlsruhe Institute of Technology (KIT), Germany Abstract For recursive circular filtering based on circular statistics, we introduce a general framework for estimation of a circular state based on different circular distributions, specifically the wrapped normal distribution and the von Mises distribution. We propose an estimation method for circular systems with nonlinear system and measurement functions. This is achieved by relying on efficient deterministic sampling techniques. Furthermore, we show how the calculations can be simplified in a variety of important special cases, such as systems with additive noise as well as identity system or measurement functions. We introduce several novel key components, particularly a distribution-free prediction algorithm, a new and superior formula for the multiplication of wrapped normal densities, and the ability to deal with non-additive system noise. All proposed methods are thoroughly evaluated and compared to several state-of-the-art solutions. 1. Introduction Estimation of circular quantities is an omnipresent issue, be it the wind direction, the angle of a robotic revolute joint, the orientation of a turntable, or the direction a vehicle is facing. Circular estimation is not limited to applications involving angles, however, and can be applied to a variety of periodic phenomena. For example phase estimation is a common issue in signal processing, and tracking objects that periodically move along a certain trajectory is also of interest. Standard approaches to circular estimation are typically based on estimation techniques designed for linear scenarios that are tweaked to deal with some of the issues arising in the presence of circular quantities. -
Efficient Evaluation of the Probability Density Function
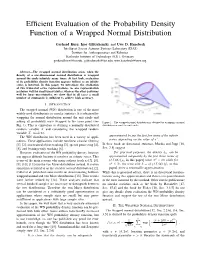
Efficient Evaluation of the Probability Density Function of a Wrapped Normal Distribution Gerhard Kurz, Igor Gilitschenski, and Uwe D. Hanebeck Intelligent Sensor-Actuator-Systems Laboratory (ISAS) Institute for Anthropomatics and Robotics Karlsruhe Institute of Technology (KIT), Germany [email protected], [email protected], [email protected] Abstract—The wrapped normal distribution arises when the density of a one-dimensional normal distribution is wrapped around the circle infinitely many times. At first look, evaluation of its probability density function appears tedious as an infinite series is involved. In this paper, we investigate the evaluation of two truncated series representations. As one representation performs well for small uncertainties, whereas the other performs well for large uncertainties, we show that in all cases a small number of summands is sufficient to achieve high accuracy. I. INTRODUCTION The wrapped normal (WN) distribution is one of the most widely used distributions in circular statistics. It is obtained by wrapping the normal distribution around the unit circle and adding all probability mass wrapped to the same point (see Figure 1. The wrapped normal distribution is obtained by wrapping a normal Fig. 1). This is equivalent to defining a normally distributed distribution around the unit circle. random variable X and considering the wrapped random variable X mod 2π. approximated by just the first few terms of the infinite The WN distribution has been used in a variety of appli- series, depending on the value of σ2. cations. These applications include nonlinear circular filtering [1], [2], constrained object tracking [3], speech processing [4], In their book on directional statistics, Mardia and Jupp [16, [5], and bearings-only tracking [6]. -

Inferences Based on a Bivariate Distribution with Von Mises Marginals
Ann. Inst. Statist. Math. Vol. 57, No. 4, 789-802 (2005) (~)2005 The Institute of Statistical Mathematics INFERENCES BASED ON A BIVARIATE DISTRIBUTION WITH VON MISES MARGINALS GRACE S. SHIEH I AND RICHARD A. JOHNSON2 l Institute of Statistical Science, Academia Sinica, Taipei 115, Taiwan, R. O. C., e-mail: gshieh@stat .sinica.edu.tw 2Department of Statistics, University of Wisconsin-Madison, WI 53706, U.S.A., e-mail: [email protected] (Received April 21, 2004; revised August 24, 2004) Abstract. There is very little literature concerning modeling the correlation be- tween paired angular observations. We propose a bivariate model with von Mises marginal distributions. An algorithm for generating bivariate angles from this von Mises distribution is given. Maximum likelihood estimation is then addressed. We also develop a likelihood ratio test for independence in paired circular data. Applica- tion of the procedures to paired wind directions is illustrated. Employing simulation, using the proposed model, we compare the power of the likelihood ratio test with six existing tests of independence. Key words and phrases: Angular observations, maximum likelihood estimation, models of dependence, power, testing. 1. Introduction Testing for independence in paired circular (bivariate angular) data has been ex- plored by many statisticians; see Fisher (1995) and Mardia and Jupp (2000) for thorough accounts. Paired circular data are either both angular or both cyclic in nature. For in- stance, wind directions at 6 a.m. and at noon recorded at a weather station for 21 consecutive days (Johnson and Wehrly (1977)), estimated peak times for two successive measurements of diastolic blood pressure (Downs (1974)), among others. -

MML Mixture Modelling of Rnulti-State, Poisson, Von Mises Circular and Gaussian Distributions
MML mixture modelling of rnulti-state, Poisson, von Mises circular and Gaussian distributions Chris S. Wallace and David L. Dowe Department of Computer Science, Monash University, Clayton, Victoria 3168, Australia e-mail: {csw, dld} @cs.monash.edu.au Abstract Bayes's Theorem. Since D and Pr(D) are given and we wish to infer H , we can regard the problem of maximis Minimum Message Length (MML) is an invariant ing the posterior probability, Pr(H!D), as one of choos Bayesian point estimation technique which is also con ing H so as to maximise Pr(H). Pr(DIH) . sistent and efficient. We provide a brief overview of An information-theoretic interpretation of MML is MML inductive inference (Wallace and Boulton (1968) , that elementary coding theory tells us that an event of Wallace and Freeman (1987)) , and how it has both probability p can be coded (e.g. by a Huffman code) by an information-theoretic and a Bayesian interpretation. a message of length l = - log2 p bits. (Negligible or no We then outline how MML is used for statistical pa harm is done by ignoring effects of rounding up to the rameter estimation, and how the MML mixture mod next positive integer.) elling program, Snob (Wallace and Boulton (1968), Wal So, since - log2 (Pr(H). Pr(D!H)) = - log2 (Pr(H)) - lace (1986) , Wallace and Dowe(1994)) uses the message log2 (Pr(DjH)), maximising the posterior probability, lengths from various parameter estimates to enable it to Pr(HID), is equivalent to minimising combine parameter estimation with selection of the num ber of components. -

Comparing Measures of Fit for Circular Distributions
Comparing measures of fit for circular distributions by Zheng Sun B.Sc., Simon Fraser University, 2006 A Thesis Submitted in Partial Fulfillment of the Requirements for the Degree of MASTER OF SCIENCE in the Department of Mathematics and Statistics © Zheng Sun, 2009 University of Victoria All rights reserved. This dissertation may not be reproduced in whole or in part, by photocopying or other means, without the permission of the author. ii Comparing measures of fit for circular distributions by Zheng Sun B.Sc., Simon Fraser University, 2006 Supervisory Committee Dr. William J. Reed, Supervisor (Department of Mathematics and Statistics) Dr. Laura Cowen, Departmental Member (Department of Mathematics and Statistics) Dr. Mary Lesperance, Departmental Member (Department of Mathematics and Statistics) iii Supervisory Committee Dr. William J. Reed, Supervisor (Department of Mathematics and Statistics) Dr. Laura Cowen, Departmental Member (Department of Mathematics and Statistics) Dr. Mary Lesperance, Departmental Member (Department of Mathematics and Statistics) ABSTRACT This thesis shows how to test the fit of a data set to a number of different models, using Watson's U 2 statistic for both grouped and continuous data. While Watson's U 2 statistic was introduced for continuous data, in recent work, the statistic has been adapted for grouped data. However, when using Watson's U 2 for continuous data, the asymptotic distribution is difficult to obtain, particularly, for some skewed circular distributions that contain four or five parameters. Until now, U 2 asymptotic points are worked out only for uniform distribution and the von Mises distribution among all circular distributions. We give U 2 asymptotic points for the wrapped exponential distributions, and we show that U 2 asymptotic points when data are grouped is usually easier to obtain for other more advanced circular distributions. -

Regularized Multivariate Von Mises Distribution
Regularized Multivariate von Mises Distribution Luis Rodriguez-Lujan, Concha Bielza, and Pedro Larra~naga Departamento de Inteligencia Artificial Universidad Polit´ecnicade Madrid Campus de Montegancedo, 28660 Boadilla del Monte, Madrid, Spain [email protected], [email protected], [email protected] http://cig.fi.upm.es Abstract. Regularization is necessary to avoid overfitting when the number of data samples is low compared to the number of parameters of the model. In this paper, we introduce a flexible L1 regularization for the multivariate von Mises distribution. We also propose a circular distance that can be used to estimate the Kullback-Leibler divergence between two circular distributions by means of sampling, and also serves as goodness-of-fit measure. We compare the models on synthetic data and real morphological data from human neurons and show that the regularized model achieves better results than non regularized von Mises model. Keywords: Von Mises distribution, directional distributions, circular distance, machine learning 1 Introduction Directional data is ubiquitous in science, from the direction of the wind to the branching angles of the trees. However, directional data has been traditionally treated as regular linear data despite of its different nature. Directional statistics [4] provides specific tools for modelling directional data. If the normal distribu- tion is the most famous distribution for linear data, the von Mises distribution [6] is its analogue for directional data. If we extend the von Mises as a multivariate distribution [3], we face the problem that no closed formulation is known for the normalization term when the number of variables is greater than two, and, therefore it cannot be easily fitted nor compared to other distributions. -

Circular Data Analysis
NCSS Statistical Software NCSS.com Chapter 230 Circular Data Analysis Introduction This procedure computes summary statistics, generates rose plots and circular histograms, computes hypothesis tests appropriate for one, two, and several groups, and computes the circular correlation coefficient for circular data. Angular data, recorded in degrees or radians, is generated in a wide variety of scientific research areas. Examples of angular (and cyclical) data include daily wind directions, ocean current directions, departure directions of animals, direction of bone-fracture plane, and orientation of bees in a beehive after stimuli. The usual summary statistics, such as the sample mean and standard deviation, cannot be used with angular values. For example, consider the average of the angular values 1 and 359. The simple average is 180. But with a little thought, we might conclude that 0 is a better answer. Because of this and other problems, a special set of techniques have been developed for analyzing angular data. This procedure implements many of those techniques. 230-1 © NCSS, LLC. All Rights Reserved. NCSS Statistical Software NCSS.com Circular Data Analysis Technical Details Suppose a sample of n angles a1,a2 ,...,an is to be summarized. It is assumed that these angles are in degrees. Fisher (1993) and Mardia & Jupp (2000) contain definitions of various summary statistics that are used for angular data. These results will be presented next. Let n n Cp S p Cp = ∑ cos( pai ) , Cp = , S p = ∑sin( pai ) , S p = , i=1 n i=1 n R R = C2 + S 2 , R = p p p p p n S −1 p > > tan Cp 0, S p 0 Cp S −1 p + π < Tp = tan Cp 0 Cp S −1 p + π < > tan 2 S p 0, Cp 0 Cp To interpret these quantities it may be useful to imagine that each angle represents a vector of length one in the direction of the angle. -

Package 'Directional'
Package ‘Directional’ May 26, 2021 Type Package Title A Collection of R Functions for Directional Data Analysis Version 5.0 URL Date 2021-05-26 Author Michail Tsagris, Giorgos Athineou, Anamul Sajib, Eli Amson, Micah J. Waldstein Maintainer Michail Tsagris <[email protected]> Description A collection of functions for directional data (including massive data, with millions of ob- servations) analysis. Hypothesis testing, discriminant and regression analysis, MLE of distribu- tions and more are included. The standard textbook for such data is the ``Directional Statis- tics'' by Mardia, K. V. and Jupp, P. E. (2000). Other references include a) Phillip J. Paine, Si- mon P. Preston Michail Tsagris and Andrew T. A. Wood (2018). An elliptically symmetric angu- lar Gaussian distribution. Statistics and Computing 28(3): 689-697. <doi:10.1007/s11222-017- 9756-4>. b) Tsagris M. and Alenazi A. (2019). Comparison of discriminant analysis meth- ods on the sphere. Communications in Statistics: Case Studies, Data Analysis and Applica- tions 5(4):467--491. <doi:10.1080/23737484.2019.1684854>. c) P. J. Paine, S. P. Pre- ston, M. Tsagris and Andrew T. A. Wood (2020). Spherical regression models with general co- variates and anisotropic errors. Statistics and Computing 30(1): 153--165. <doi:10.1007/s11222- 019-09872-2>. License GPL-2 Imports bigstatsr, parallel, doParallel, foreach, RANN, Rfast, Rfast2, rgl RoxygenNote 6.1.1 NeedsCompilation no Repository CRAN Date/Publication 2021-05-26 15:20:02 UTC R topics documented: Directional-package . .4 A test for testing the equality of the concentration parameters for ciruclar data . .5 1 2 R topics documented: Angular central Gaussian random values simulation .