Cotranscriptional Splicing Efficiency Differs Dramatically Between Drosophila and Mouse
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

A Family of Signal-Responsive E3 Ubiquitin Ligases Mediating TLR Signaling and T-Cell Tolerance
Cellular & Molecular Immunology (2012) 9, 113–122 ß 2012 CSI and USTC. All rights reserved 1672-7681/12 $32.00 www.nature.com/cmi REVIEW Peli: a family of signal-responsive E3 ubiquitin ligases mediating TLR signaling and T-cell tolerance Wei Jin1, Mikyoung Chang2 and Shao-Cong Sun2 E3 ubiquitin ligases play a crucial role in regulating immune receptor signaling and modulating immune homeostasis and activation. One emerging family of such E3s is the Pelle-interacting (Peli) proteins, characterized by the presence of a cryptic forkhead-associated domain involved in substrate binding and an atypical RING domain mediating formation of both lysine (K) 63- and K48-linked polyubiquitin chains. A well-recognized function of Peli family members is participation in the signal transduction mediated by Toll-like receptors (TLRs) and IL-1 receptor. Recent gene targeting studies have provided important insights into the in vivo functions of Peli1 in the regulation of TLR signaling and inflammation. These studies have also extended the biological functions of Peli1 to the regulation of T-cell tolerance. Consistent with its immunoregulatory functions, Peli1 responds to different immune stimuli for its gene expression and catalytic activation. In this review, we discuss the recent progress, as well as the historical perspectives in the regulation and biological functions of Peli. Cellular & Molecular Immunology (2012) 9, 113–122; doi:10.1038/cmi.2011.60; published online 6 February 2012 Keywords: Peli; Pellino; ubiquitination; NF-kB; TLR; T-cell tolerance -

The Human Gene Connectome As a Map of Short Cuts for Morbid Allele Discovery
The human gene connectome as a map of short cuts for morbid allele discovery Yuval Itana,1, Shen-Ying Zhanga,b, Guillaume Vogta,b, Avinash Abhyankara, Melina Hermana, Patrick Nitschkec, Dror Friedd, Lluis Quintana-Murcie, Laurent Abela,b, and Jean-Laurent Casanovaa,b,f aSt. Giles Laboratory of Human Genetics of Infectious Diseases, Rockefeller Branch, The Rockefeller University, New York, NY 10065; bLaboratory of Human Genetics of Infectious Diseases, Necker Branch, Paris Descartes University, Institut National de la Santé et de la Recherche Médicale U980, Necker Medical School, 75015 Paris, France; cPlateforme Bioinformatique, Université Paris Descartes, 75116 Paris, France; dDepartment of Computer Science, Ben-Gurion University of the Negev, Beer-Sheva 84105, Israel; eUnit of Human Evolutionary Genetics, Centre National de la Recherche Scientifique, Unité de Recherche Associée 3012, Institut Pasteur, F-75015 Paris, France; and fPediatric Immunology-Hematology Unit, Necker Hospital for Sick Children, 75015 Paris, France Edited* by Bruce Beutler, University of Texas Southwestern Medical Center, Dallas, TX, and approved February 15, 2013 (received for review October 19, 2012) High-throughput genomic data reveal thousands of gene variants to detect a single mutated gene, with the other polymorphic genes per patient, and it is often difficult to determine which of these being of less interest. This goes some way to explaining why, variants underlies disease in a given individual. However, at the despite the abundance of NGS data, the discovery of disease- population level, there may be some degree of phenotypic homo- causing alleles from such data remains somewhat limited. geneity, with alterations of specific physiological pathways under- We developed the human gene connectome (HGC) to over- come this problem. -
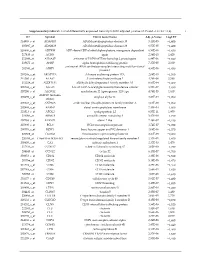
Supplementary Table S1. List of Differentially Expressed
Supplementary table S1. List of differentially expressed transcripts (FDR adjusted p‐value < 0.05 and −1.4 ≤ FC ≥1.4). 1 ID Symbol Entrez Gene Name Adj. p‐Value Log2 FC 214895_s_at ADAM10 ADAM metallopeptidase domain 10 3,11E‐05 −1,400 205997_at ADAM28 ADAM metallopeptidase domain 28 6,57E‐05 −1,400 220606_s_at ADPRM ADP‐ribose/CDP‐alcohol diphosphatase, manganese dependent 6,50E‐06 −1,430 217410_at AGRN agrin 2,34E‐10 1,420 212980_at AHSA2P activator of HSP90 ATPase homolog 2, pseudogene 6,44E‐06 −1,920 219672_at AHSP alpha hemoglobin stabilizing protein 7,27E‐05 2,330 aminoacyl tRNA synthetase complex interacting multifunctional 202541_at AIMP1 4,91E‐06 −1,830 protein 1 210269_s_at AKAP17A A‐kinase anchoring protein 17A 2,64E‐10 −1,560 211560_s_at ALAS2 5ʹ‐aminolevulinate synthase 2 4,28E‐06 3,560 212224_at ALDH1A1 aldehyde dehydrogenase 1 family member A1 8,93E‐04 −1,400 205583_s_at ALG13 ALG13 UDP‐N‐acetylglucosaminyltransferase subunit 9,50E‐07 −1,430 207206_s_at ALOX12 arachidonate 12‐lipoxygenase, 12S type 4,76E‐05 1,630 AMY1C (includes 208498_s_at amylase alpha 1C 3,83E‐05 −1,700 others) 201043_s_at ANP32A acidic nuclear phosphoprotein 32 family member A 5,61E‐09 −1,760 202888_s_at ANPEP alanyl aminopeptidase, membrane 7,40E‐04 −1,600 221013_s_at APOL2 apolipoprotein L2 6,57E‐11 1,600 219094_at ARMC8 armadillo repeat containing 8 3,47E‐08 −1,710 207798_s_at ATXN2L ataxin 2 like 2,16E‐07 −1,410 215990_s_at BCL6 BCL6 transcription repressor 1,74E‐07 −1,700 200776_s_at BZW1 basic leucine zipper and W2 domains 1 1,09E‐06 −1,570 222309_at -

Quantitative Trait Loci Mapping of Macrophage Atherogenic Phenotypes
QUANTITATIVE TRAIT LOCI MAPPING OF MACROPHAGE ATHEROGENIC PHENOTYPES BRIAN RITCHEY Bachelor of Science Biochemistry John Carroll University May 2009 submitted in partial fulfillment of requirements for the degree DOCTOR OF PHILOSOPHY IN CLINICAL AND BIOANALYTICAL CHEMISTRY at the CLEVELAND STATE UNIVERSITY December 2017 We hereby approve this thesis/dissertation for Brian Ritchey Candidate for the Doctor of Philosophy in Clinical-Bioanalytical Chemistry degree for the Department of Chemistry and the CLEVELAND STATE UNIVERSITY College of Graduate Studies by ______________________________ Date: _________ Dissertation Chairperson, Johnathan D. Smith, PhD Department of Cellular and Molecular Medicine, Cleveland Clinic ______________________________ Date: _________ Dissertation Committee member, David J. Anderson, PhD Department of Chemistry, Cleveland State University ______________________________ Date: _________ Dissertation Committee member, Baochuan Guo, PhD Department of Chemistry, Cleveland State University ______________________________ Date: _________ Dissertation Committee member, Stanley L. Hazen, MD PhD Department of Cellular and Molecular Medicine, Cleveland Clinic ______________________________ Date: _________ Dissertation Committee member, Renliang Zhang, MD PhD Department of Cellular and Molecular Medicine, Cleveland Clinic ______________________________ Date: _________ Dissertation Committee member, Aimin Zhou, PhD Department of Chemistry, Cleveland State University Date of Defense: October 23, 2017 DEDICATION I dedicate this work to my entire family. In particular, my brother Greg Ritchey, and most especially my father Dr. Michael Ritchey, without whose support none of this work would be possible. I am forever grateful to you for your devotion to me and our family. You are an eternal inspiration that will fuel me for the remainder of my life. I am extraordinarily lucky to have grown up in the family I did, which I will never forget. -

Content Based Search in Gene Expression Databases and a Meta-Analysis of Host Responses to Infection
Content Based Search in Gene Expression Databases and a Meta-analysis of Host Responses to Infection A Thesis Submitted to the Faculty of Drexel University by Francis X. Bell in partial fulfillment of the requirements for the degree of Doctor of Philosophy November 2015 c Copyright 2015 Francis X. Bell. All Rights Reserved. ii Acknowledgments I would like to acknowledge and thank my advisor, Dr. Ahmet Sacan. Without his advice, support, and patience I would not have been able to accomplish all that I have. I would also like to thank my committee members and the Biomed Faculty that have guided me. I would like to give a special thanks for the members of the bioinformatics lab, in particular the members of the Sacan lab: Rehman Qureshi, Daisy Heng Yang, April Chunyu Zhao, and Yiqian Zhou. Thank you for creating a pleasant and friendly environment in the lab. I give the members of my family my sincerest gratitude for all that they have done for me. I cannot begin to repay my parents for their sacrifices. I am eternally grateful for everything they have done. The support of my sisters and their encouragement gave me the strength to persevere to the end. iii Table of Contents LIST OF TABLES.......................................................................... vii LIST OF FIGURES ........................................................................ xiv ABSTRACT ................................................................................ xvii 1. A BRIEF INTRODUCTION TO GENE EXPRESSION............................. 1 1.1 Central Dogma of Molecular Biology........................................... 1 1.1.1 Basic Transfers .......................................................... 1 1.1.2 Uncommon Transfers ................................................... 3 1.2 Gene Expression ................................................................. 4 1.2.1 Estimating Gene Expression ............................................ 4 1.2.2 DNA Microarrays ...................................................... -

PP1-Associated Signaling and − B/AP-1 Κ Inhibition of NF- Tolerance
Downloaded from http://www.jimmunol.org/ by guest on October 3, 2021 is online at: average * and − B/AP-1 κ The Journal of Immunology published online 26 February 2014 from submission to initial decision 4 weeks from acceptance to publication http://www.jimmunol.org/content/early/2014/02/26/jimmun ol.1301610 Identification of Two Forms of TNF Tolerance in Human Monocytes: Differential Inhibition of NF- PP1-Associated Signaling Johannes Günther, Nico Vogt, Katharina Hampel, Rolf Bikker, Sharon Page, Benjamin Müller, Judith Kandemir, Michael Kracht, Oliver Dittrich-Breiholz, René Huber and Korbinian Brand J Immunol Submit online. Every submission reviewed by practicing scientists ? is published twice each month by Receive free email-alerts when new articles cite this article. Sign up at: http://jimmunol.org/alerts http://jimmunol.org/subscription Submit copyright permission requests at: http://www.aai.org/About/Publications/JI/copyright.html http://www.jimmunol.org/content/suppl/2014/02/26/jimmunol.130161 0.DCSupplemental Information about subscribing to The JI No Triage! Fast Publication! Rapid Reviews! 30 days* Why • • • Material Permissions Email Alerts Subscription Supplementary The Journal of Immunology The American Association of Immunologists, Inc., 1451 Rockville Pike, Suite 650, Rockville, MD 20852 Copyright © 2014 by The American Association of Immunologists, Inc. All rights reserved. Print ISSN: 0022-1767 Online ISSN: 1550-6606. This information is current as of October 3, 2021. Published February 26, 2014, doi:10.4049/jimmunol.1301610 The Journal of Immunology Identification of Two Forms of TNF Tolerance in Human Monocytes: Differential Inhibition of NF-kB/AP-1– and PP1-Associated Signaling Johannes Gunther,*€ ,1 Nico Vogt,*,1 Katharina Hampel,*,1 Rolf Bikker,* Sharon Page,* Benjamin Muller,*€ Judith Kandemir,* Michael Kracht,† Oliver Dittrich-Breiholz,‡ Rene´ Huber,* and Korbinian Brand* The molecular basis of TNF tolerance is poorly understood. -

Integrated Bioinformatics Analysis Reveals Novel Key Biomarkers and Potential Candidate Small Molecule Drugs in Gestational Diabetes Mellitus
bioRxiv preprint doi: https://doi.org/10.1101/2021.03.09.434569; this version posted March 10, 2021. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Integrated bioinformatics analysis reveals novel key biomarkers and potential candidate small molecule drugs in gestational diabetes mellitus Basavaraj Vastrad1, Chanabasayya Vastrad*2, Anandkumar Tengli3 1. Department of Biochemistry, Basaveshwar College of Pharmacy, Gadag, Karnataka 582103, India. 2. Biostatistics and Bioinformatics, Chanabasava Nilaya, Bharthinagar, Dharwad 580001, Karnataka, India. 3. Department of Pharmaceutical Chemistry, JSS College of Pharmacy, Mysuru and JSS Academy of Higher Education & Research, Mysuru, Karnataka, 570015, India * Chanabasayya Vastrad [email protected] Ph: +919480073398 Chanabasava Nilaya, Bharthinagar, Dharwad 580001 , Karanataka, India bioRxiv preprint doi: https://doi.org/10.1101/2021.03.09.434569; this version posted March 10, 2021. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Abstract Gestational diabetes mellitus (GDM) is one of the metabolic diseases during pregnancy. The identification of the central molecular mechanisms liable for the disease pathogenesis might lead to the advancement of new therapeutic options. The current investigation aimed to identify central differentially expressed genes (DEGs) in GDM. The transcription profiling by array data (E-MTAB-6418) was obtained from the ArrayExpress database. The DEGs between GDM samples and non GDM samples were analyzed with limma package. Gene ontology (GO) and REACTOME enrichment analysis were performed using ToppGene. Then we constructed the protein-protein interaction (PPI) network of DEGs by the Search Tool for the Retrieval of Interacting Genes database (STRING) and module analysis was performed. -

Epigenetic Silencing Mediated Through Activated PI3K/AKT Signaling in Breast Cancer
SUPPLEMENTARY: Epigenetic Silencing Mediated Through Activated PI3K/AKT Signaling in Breast Cancer Tao Zuo, Ta-Ming Liu, Xun Lan, Yu-I Weng, Rulong Shen, Fei Gu, Yi-Wen Huang, Sandya Liyanarachchi, Daniel E. Deatherage, Pei-Yin Hsu, Cenny Taslim, Bhuvaneswari Ramaswamy, Charles L. Shapiro, Huey-Jen L. Lin, Alfred S. L. Cheng, Victor X. Jin, and Tim H.-M. Huang 1 Table of Content: 1. Fig. S1. Elevated AKT1 kinase activity in myrAKT1- transfected MCF10A cells. ---------------------------------------------------------------------------------------------------- page 4 2. Fig. S2. Genome-wide view of H3K27me3 in mammary epithelial cells. ---------------------------------------------------------------------------------------------------- page 5 3. Fig. S3. Biological functions of PI3K/AKT- downstream genes. ----------------- page 6 4. Fig. S4. Genomic maps of 40 genes. ------------------------------------------------- page 8 5. Fig. S5. Comparing DNA methylation levels of four CpG islands measured by meDIP-qPCR and MassARRAY.------------------------------------------------------- page 11 6. Fig. S6. Validation of DNA methylation by MBDCap-seq.----------------------- page 13 7. Table S1. Summary of H3K27me3 ChIP-seq data ------------------------------ page 14 8. Table S2. Distribution of H3K27me3 in MCF10A genome ---------------------- page 14 9. Table S3. Genes (n=488) enriched with H3K27me3 and downregulated by PI3K/AKT- signaling ---------------------------------------------------------------------- page 15 10. Table S4. Genes (n=1005) enriched -

Homozygosity Mapping Reveals Null Mutations in FAM161A As a Cause of Autosomal-Recessive Retinitis Pigmentosa
REPORT Homozygosity Mapping Reveals Null Mutations in FAM161A as a Cause of Autosomal-Recessive Retinitis Pigmentosa Dikla Bandah-Rozenfeld,1 Liliana Mizrahi-Meissonnier,1 Chen Farhy,2 Alexey Obolensky,1 Itay Chowers,1 Jacob Pe’er,1 Saul Merin,3 Tamar Ben-Yosef,4 Ruth Ashery-Padan,2 Eyal Banin,1,* and Dror Sharon1,* Retinitis pigmentosa (RP) is a heterogeneous group of inherited retinal degenerations caused by mutations in at least 45 genes. Using homozygosity mapping, we identified a ~4 Mb homozygous region on chromosome 2p15 in patients with autosomal-recessive RP (arRP). This region partially overlaps with RP28, a previously identified arRP locus. Sequence analysis of 12 candidate genes revealed three null mutations in FAM161A in 20 families. RT-PCR analysis in 21 human tissues revealed high levels of FAM161A expression in the retina and lower levels in the brain and testis. In the human retina, we identified two alternatively spliced transcripts with an intact open reading frame, the major one lacking a highly conserved exon. During mouse embryonic development, low levels of Fam161a tran- scripts were detected throughout the optic cup. After birth, Fam161a expression was elevated and confined to the photoreceptor layer. FAM161A encodes a protein of unknown function that is moderately conserved in mammals. Clinical manifestations of patients with FAM161A mutations varied but were largely within the spectrum associated with arRP. On funduscopy, pallor of the optic discs and attenuation of blood vessels were common, but bone-spicule-like pigmentation was often mild or lacking. Most patients had nonrecord- able electroretinographic responses and constriction of visual fields upon diagnosis. -

Epigenetic Study of Plasma Circulating DNA in Prostate Cancer
Epigenetic Study of Plasma Circulating DNA in Prostate Cancer by Andrew Kwan A thesis submitted in conformity with the requirements for the degree of Master of Science Graduate Department of Pharmacology and Toxicology University of Toronto © Copyright by Andrew Kwan (2015) Epigenetic Study of Plasma Circulating DNA in Prostate Cancer Andrew Kwan Master of Science Graduate Department of Pharmacology and Toxicology University of Toronto 2015 Abstract Malignant cells exhibit epigenetic differences compared to normal cells, which can also be detected in the cell-free circulating DNA (cirDNA) of bodily fluids. DNA modification studies in cirDNA may lead to the development of non-invasive cancer biomarkers. This thesis investigates plasma cirDNA modifications of prostate cancer patients, benign prostate hyperplasia patients, and men without any known prostate disease. A modification-sensitive restriction enzyme approach was used to enrich the modified DNA fractions, which were interrogated on CpG island- and Affymetrix tiling microarrays. Our initial discovery screen identified a number of differentially modified genes and non-coding loci between PCa patients and controls; a subset of candidate loci were verified using bisulfite pyrosequencing. We applied machine-learning techniques to develop a multi-locus biomarker set, which exhibited promising predictive performance in distinguishing cancer patients from unaffected controls. Our results suggest that epigenomics of cirDNA is a promising source for diagnostic non-invasive biomarkers of solid tumors. ii Acknowledgments I would like to start by expressing my gratitude to my supervisor, Dr. Arturas Petronis, for giving me the opportunity to work under his mentorship and guidance for the past few years. It is under his tutelage that I have grown both as a person and a scientist. -

The Glomerular Transcriptome and a Predicted Protein–Protein Interaction Network
BASIC RESEARCH www.jasn.org The Glomerular Transcriptome and a Predicted Protein–Protein Interaction Network Liqun He,* Ying Sun,* Minoru Takemoto,* Jenny Norlin,* Karl Tryggvason,* Tore Samuelsson,† and Christer Betsholtz*‡ *Division of Matrix Biology, Department of Medical Biochemistry and Biophysics, and ‡Department of Medicine, Karolinska Institutet, Stockholm, and †Department of Medical Biochemistry, Go¨teborg University, Go¨teborg, Sweden ABSTRACT To increase our understanding of the molecular composition of the kidney glomerulus, we performed a meta-analysis of available glomerular transcriptional profiles made from mouse and man using five different methodologies. We generated a combined catalogue of glomerulus-enriched genes that emerged from these different sources and then used this to construct a predicted protein–protein interaction network in the glomerulus (GlomNet). The combined glomerulus-enriched gene catalogue provides the most comprehensive picture of the molecular composition of the glomerulus currently available, and GlomNet contributes an integrative systems biology approach to the understanding of glomerular signaling networks that operate during development, function, and disease. J Am Soc Nephrol 19: 260–268, 2008. doi: 10.1681/ASN.2007050588 Many kidney diseases and, importantly, approxi- nins have been shown to be mutated in Alport syn- mately two thirds of all cases of ESRD originate with drome and Pierson congenital nephrotic syndromes, glomerular disease. Most cases of glomerular disease respectively.11,12 Genetic studies in mice have further are caused by systemic disorders (e.g., diabetes, hyper- revealed genes and proteins of importance for glomer- tension, lupus, obesity) for which the molecular ulus development and function, such as podoca- pathogeneses of the glomerular complications are un- lyxin,13 CD2AP,14 NEPH1,15 FAT1,16 forkhead box known. -

Gene Expression Profiles in Peripheral Lymphocytes by Arsenic Exposure and Skin Lesion Status in a Bangladeshi Population
1367 Gene Expression Profiles in Peripheral Lymphocytes by Arsenic Exposure and Skin Lesion Status in a Bangladeshi Population Maria Argos,1 Muhammad G. Kibriya,1 Faruque Parvez,2 Farzana Jasmine,1 Muhammad Rakibuz-Zaman,4 and Habibul Ahsan1,3 Departments of 1Epidemiology and 2Environmental Health Sciences, Mailman School of Public Health, and 3Herbert Irving Comprehensive Cancer Center, Columbia University, New York, New York and 4Columbia University Arsenic Project in Bangladesh, Bangladesh Abstract Millions of individuals worldwide are chronically exposed hundred sixty-eight genes were differentially expressed to arsenic through their drinking water. In this study, the between these two groups, from which 312 differentially effect of arsenic exposure and arsenical skin lesion status expressed genes were identified by restricting the analysis on genome-wide gene expression patterns was evaluated to female never-smokers. We also explored possible using RNA from peripheral blood lymphocytes of individ- differential gene expression by arsenic exposure levels uals selected from the Health Effects of Arsenic Longitudi- among individuals without manifest arsenical skin lesions; nal Study. Affymetrix HG-U133A GeneChip (Affymetrix, however, no differentially expressed genes could be Santa Clara, CA) arrays were used to measure the identified from this comparison. Our findings show that expression of f22,000 transcripts. Our primary statistical microarray-based gene expression analysis is a powerful analysis involved identifying differentially expressed genes method to characterize the molecular profile of arsenic between participants with and without arsenical skin exposure and arsenic-induced diseases. Genes identified lesions based on the significance analysis of microarrays from this analysis may provide insights into the underlying statistic with an a priori defined 1% false discovery rate to processes of arsenic-induced disease and represent potential minimize false positives.