Sparqling Nextprot Data
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

3De8d416-B844-4E9a-B3fb-A2e883a4083f 11119
Edinburgh Research Explorer Last rolls of the yoyo: Assessing the human canonical protein count Citation for published version: Southan, C 2017, 'Last rolls of the yoyo: Assessing the human canonical protein count', F1000Research, vol. 6, pp. 448+. https://doi.org/10.12688/f1000research.11119.1 Digital Object Identifier (DOI): 10.12688/f1000research.11119.1 Link: Link to publication record in Edinburgh Research Explorer Document Version: Publisher's PDF, also known as Version of record Published In: F1000Research Publisher Rights Statement: This is an open access article distributed under the terms of the Creative Commons Attribution Licence, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited. Data associated with the article are available under the terms of the Creative Commons Zero "No rights reserved" data waiver (CC0 1.0 Public domain dedication). General rights Copyright for the publications made accessible via the Edinburgh Research Explorer is retained by the author(s) and / or other copyright owners and it is a condition of accessing these publications that users recognise and abide by the legal requirements associated with these rights. Take down policy The University of Edinburgh has made every reasonable effort to ensure that Edinburgh Research Explorer content complies with UK legislation. If you believe that the public display of this file breaches copyright please contact [email protected] providing details, and we will remove access to the work immediately -

Study of Conditions for the Emergence of Cellular Communication Using Self-Adaptive Multi-Agent Systems Sebastien Maignan
Study of conditions for the emergence of cellular communication using self-adaptive multi-agent systems Sebastien Maignan To cite this version: Sebastien Maignan. Study of conditions for the emergence of cellular communication using self- adaptive multi-agent systems. Artificial Intelligence [cs.AI]. Université Paul Sabatier - Toulouse III, 2018. English. NNT : 2018TOU30094. tel-02094368 HAL Id: tel-02094368 https://tel.archives-ouvertes.fr/tel-02094368 Submitted on 9 Apr 2019 HAL is a multi-disciplinary open access L’archive ouverte pluridisciplinaire HAL, est archive for the deposit and dissemination of sci- destinée au dépôt et à la diffusion de documents entific research documents, whether they are pub- scientifiques de niveau recherche, publiés ou non, lished or not. The documents may come from émanant des établissements d’enseignement et de teaching and research institutions in France or recherche français ou étrangers, des laboratoires abroad, or from public or private research centers. publics ou privés. Délivré par l'Université Toulouse 3 Paul Sabatier (UT3 Paul Sabatier) Sébastien Maignan Le 30 août 2018 Study of conditions for the emergence of cellular communication using self-adaptive multi-agent systems École doctorale et discipline ou spécialité ED MITT : Domaine STIC : Intelligence Artificielle Unité de recherche Institut de Recherche en Informatique de Toulouse Directrice(s) ou Directeur(s) de Thèse Pierre Glize, Carole Bernon Jury Marie BEURTON-AIMAR Maître de Conférences, HdR, Université de Bordeaux Rapporteur Vincent -

The Nextprot Knowledgebase in 2020
D328–D334 Nucleic Acids Research, 2020, Vol. 48, Database issue Published online 14 November 2019 doi: 10.1093/nar/gkz995 The neXtProt knowledgebase in 2020: data, tools and usability improvements Monique Zahn-Zabal1, Pierre-Andre´ Michel1, Alain Gateau1,Fred´ eric´ Nikitin1, Mathieu Schaeffer1,2, Estelle Audot1, Pascale Gaudet 1, Paula D. Duek1, Daniel Teixeira1, Valentine Rech de Laval1,2,3, Kasun Samarasinghe1,2,AmosBairoch1,2 and Lydie Lane1,2,* 1CALIPHO group, SIB Swiss Institute of Bioinformatics, Geneva, Switzerland, 2Department of microbiology and molecular medicine, Faculty of Medicine, University of Geneva, Geneva, Switzerland and 3Haute ecole´ specialis´ ee´ de Suisse occidentale, Haute Ecole de Gestion de Geneve,` Carouge, Switzerland Received September 11, 2019; Revised October 10, 2019; Editorial Decision October 11, 2019; Accepted October 18, 2019 ABSTRACT post-translational modifications (PTMs), as well as peptide identified in mass spectrometry experiments and epitopes The neXtProt knowledgebase (https://www.nextprot. recognized by antibodies have been integrated from a num- org) is an integrative resource providing both data ber of resources. By doing so, neXtProt extends the contents on human protein and the tools to explore these. of UniProtKB/Swiss-Prot (2) to provide a more compre- In order to provide comprehensive and up-to-date hensive data set. data, we evaluate and add new data sets. We describe However, data alone is not sufficient for scientists to the incorporation of three new data sets that provide comprehend complex information rapidly. For this reason, expression, function, protein-protein binary interac- neXtProt organizes the information concerning an entry in tion, post-translational modifications (PTM) and vari- several views, with interactive viewers that allow the user to ant information. -

AP Biology: Chemistry B Mcgraw Hill AP Biology 2014-15 Contents
AP Biology: Chemistry B McGraw Hill AP Biology 2014-15 Contents 1 Carbohydrate 1 1.1 Structure .................................................. 1 1.2 Monosaccharides ............................................. 2 1.2.1 Classification of monosaccharides ................................ 2 1.2.2 Ring-straight chain isomerism .................................. 3 1.2.3 Use in living organisms ...................................... 3 1.3 Disaccharides ............................................... 3 1.4 Nutrition .................................................. 4 1.4.1 Classification ........................................... 5 1.5 Metabolism ................................................ 5 1.5.1 Catabolism ............................................ 5 1.6 Carbohydrate chemistry .......................................... 5 1.7 See also .................................................. 6 1.8 References ................................................. 6 1.9 External links ............................................... 7 2 Lipid 8 2.1 Categories of lipids ............................................ 8 2.1.1 Fatty acids ............................................. 8 2.1.2 Glycerolipids ........................................... 9 2.1.3 Glycerophospholipids ....................................... 9 2.1.4 Sphingolipids ........................................... 9 2.1.5 Sterol lipids ............................................ 10 2.1.6 Prenol lipids ............................................ 10 2.1.7 Saccharolipids .......................................... -

Converting Nextprot Into Linked Data and Nanopublications
Converting neXtProt into Linked Data and nanopublications Editor(s): Name Surname, University, Country Solicited review(s): Name Surname, University, Country Open review(s): Name Surname, University, Country Christine Chichester*a1, Oliver Karch*b, Pascale Gaudeta, Lydie Lanea, Barend Monsc, and Amos Bairocha a CALIPHO group, Swiss Institute of Bioinformatics, CMU 1 rue Michel Serve, 1211 Geneva 4, Switzerland b Biomarker Technologies, Discovery Bioinformatics, Merck Serono, Frankfurter Str. 250, 64271 Darmstadt, Germany c Netherlands Bioinformatics Centre, P.O. Box 9101, 6500 HB Nijmegen, The Netherlands Abstract. The development of Linked Data provides the opportunity for databases to supply extensive volumes of biological data, information, and knowledge in a machine interpretable format to make previously isolated data silos interoperable. To increase ease of use, often databases incorporate annotations from several different resources. Linked Data can overcome many formatting and identifier issues that prevent data interoperability, but the extensive cross incorporation of annotations between databases makes the tracking of provenance in open, decentralized systems especially important. With the diversity of published data, provenance information becomes critical to providing reliable and trustworthy services to scientists. The nanopublication system addresses many of these challenges. We have developed the neXtProt Linked Data by serializing in RDF/XML annotations specific to neXtProt and started employing the nanopublication model to give appropriate attribution to all data. Specifically, a use case demonstrates the handling of post-translational modification (PTM) data modeled as nanopublications to illustrate the how the different levels of provenance and data quality thresholds can be captured in this model. Keywords: biological database, linked data, semantic web, nanopublication, post-translation modification 1 Corresponding author. -

How Many Human Proteoforms Are There?
PERSPECTIVE PUBLISHED ONLINE: 14 FEBRUARY 2018 | DOI: 10.1038/NCHEMBIO.2576 How many human proteoforms are there? Ruedi Aebersold1, Jeffrey N Agar2, I Jonathan Amster3 , Mark S Baker4 , Carolyn R Bertozzi5, Emily S Boja6, Catherine E Costello7, Benjamin F Cravatt8 , Catherine Fenselau9, Benjamin A Garcia10, Ying Ge11,12, Jeremy Gunawardena13, Ronald C Hendrickson14, Paul J Hergenrother15, Christian G Huber16 , Alexander R Ivanov2, Ole N Jensen17, Michael C Jewett18, Neil L Kelleher19* , Laura L Kiessling20 , Nevan J Krogan21, Martin R Larsen17, Joseph A Loo22 , Rachel R Ogorzalek Loo22, Emma Lundberg23,24, Michael J MacCoss25, Parag Mallick5, Vamsi K Mootha13, Milan Mrksich18, Tom W Muir26, Steven M Patrie19, James J Pesavento27 , Sharon J Pitteri5 , Henry Rodriguez6, Alan Saghatelian28, Wendy Sandoval29, Hartmut Schlüter30 , Salvatore Sechi31, Sarah A Slavoff32, Lloyd M Smith12,33, Michael P Snyder24, Paul M Thomas19 , Mathias Uhlén34, Jennifer E Van Eyk35, Marc Vidal36, David R Walt37, Forest M White38, Evan R Williams39, Therese Wohlschlager16, Vicki H Wysocki40, Nathan A Yates41, Nicolas L Young42 & Bing Zhang42 Despite decades of accumulated knowledge about proteins and their post-translational modifications (PTMs), numerous ques- tions remain regarding their molecular composition and biological function. One of the most fundamental queries is the extent to which the combinations of DNA-, RNA- and PTM-level variations explode the complexity of the human proteome. Here, we outline what we know from current databases and measurement strategies including mass spectrometry–based proteomics. In doing so, we examine prevailing notions about the number of modifications displayed on human proteins and how they combine to generate the protein diversity underlying health and disease. -

Biomolecule and Bioentity Interaction Databases in Systems Biology: a Comprehensive Review
biomolecules Review Biomolecule and Bioentity Interaction Databases in Systems Biology: A Comprehensive Review Fotis A. Baltoumas 1,* , Sofia Zafeiropoulou 1, Evangelos Karatzas 1 , Mikaela Koutrouli 1,2, Foteini Thanati 1, Kleanthi Voutsadaki 1 , Maria Gkonta 1, Joana Hotova 1, Ioannis Kasionis 1, Pantelis Hatzis 1,3 and Georgios A. Pavlopoulos 1,3,* 1 Institute for Fundamental Biomedical Research, Biomedical Sciences Research Center “Alexander Fleming”, 16672 Vari, Greece; zafeiropoulou@fleming.gr (S.Z.); karatzas@fleming.gr (E.K.); [email protected] (M.K.); [email protected] (F.T.); voutsadaki@fleming.gr (K.V.); [email protected] (M.G.); hotova@fleming.gr (J.H.); [email protected] (I.K.); hatzis@fleming.gr (P.H.) 2 Novo Nordisk Foundation Center for Protein Research, University of Copenhagen, 2200 Copenhagen, Denmark 3 Center for New Biotechnologies and Precision Medicine, School of Medicine, National and Kapodistrian University of Athens, 11527 Athens, Greece * Correspondence: baltoumas@fleming.gr (F.A.B.); pavlopoulos@fleming.gr (G.A.P.); Tel.: +30-210-965-6310 (G.A.P.) Abstract: Technological advances in high-throughput techniques have resulted in tremendous growth Citation: Baltoumas, F.A.; of complex biological datasets providing evidence regarding various biomolecular interactions. Zafeiropoulou, S.; Karatzas, E.; To cope with this data flood, computational approaches, web services, and databases have been Koutrouli, M.; Thanati, F.; Voutsadaki, implemented to deal with issues such as data integration, visualization, exploration, organization, K.; Gkonta, M.; Hotova, J.; Kasionis, scalability, and complexity. Nevertheless, as the number of such sets increases, it is becoming more I.; Hatzis, P.; et al. Biomolecule and and more difficult for an end user to know what the scope and focus of each repository is and how Bioentity Interaction Databases in redundant the information between them is. -
Metrics for the Human Proteome Project 2016: Progress On
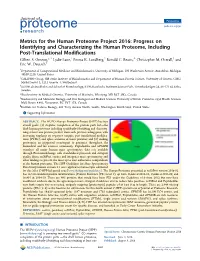
Perspective pubs.acs.org/jpr Metrics for the Human Proteome Project 2016: Progress on Identifying and Characterizing the Human Proteome, Including Post-Translational Modifications † ‡ § ⊥ ∥ Gilbert S. Omenn,*, Lydie Lane, Emma K. Lundberg, Ronald C. Beavis, Christopher M. Overall, and ¶ Eric W. Deutsch † Department of Computational Medicine and Bioinformatics, University of Michigan, 100 Washtenaw Avenue, Ann Arbor, Michigan 48109-2218, United States ‡ CALIPHO Group, SIB Swiss Institute of Bioinformatics and Department of Human Protein Science, University of Geneva, CMU, Michel-Servet 1, 1211 Geneva 4, Switzerland § SciLifeLab Stockholm and School of Biotechnology, KTH, Karolinska Institutet Science Park, Tomtebodavagen̈ 23, SE-171 65 Solna, Sweden ⊥ Biochemistry & Medical Genetics, University of Manitoba, Winnipeg, MB R3T 2N2, Canada ∥ Biochemistry and Molecular Biology, and Oral Biological and Medical Sciences University of British Columbia, 2350 Health Sciences Mall, Room 4.401, Vancouver, BC V6T 1Z3, Canada ¶ Institute for Systems Biology, 401 Terry Avenue North, Seattle, Washington 98109-5263, United States *S Supporting Information ABSTRACT: The HUPO Human Proteome Project (HPP) has two overall goals: (1) stepwise completion of the protein parts listthe draft human proteome including confidently identifying and character- izing at least one protein product from each protein-coding gene, with increasing emphasis on sequence variants, post-translational modifica- tions (PTMs), and splice isoforms of those proteins; and (2) making proteomics an integrated counterpart to genomics throughout the biomedical and life sciences community. PeptideAtlas and GPMDB reanalyze all major human mass spectrometry data sets available through ProteomeXchange with standardized protocols and stringent quality filters; neXtProt curates and integrates mass spectrometry and other findings to present the most up to date authorative compendium of the human proteome. -

Towards a Functional Definition of the Mitochondrial Human Proteome
EuPA Open Proteomics 10 (2016) 24–27 Contents lists available at ScienceDirect EuPA Open Proteomics journal homepage: www.elsevier.com/locate/euprot Towards a functional definition of the mitochondrial human proteome a, a b c d,e Mauro Fasano *, Tiziana Alberio , Mohan Babu , Emma Lundberg , Andrea Urbani a Division of Biomedical Research, Department of Theoretical and Applied Sciences, and Center of Neuroscience, University of Insubria, I-21052 Busto Arsizio, Italy b Department of Biochemistry, Research and Innovation Centre, University of Regina, Regina, SK, Canada c Science for Life Laboratory, KTH—Royal Institute of Technology, Stockholm, Sweden d Santa Lucia IRCCS Foundation, I-00143 Rome, Italy e Department of Experimental Medicine and Surgery, University of Rome “Tor Vergata”, I-00133 Rome, Italy A R T I C L E I N F O A B S T R A C T Article history: The mitochondrial human proteome project (mt-HPP) was initiated by the Italian HPP group as a part of Received 26 October 2015 both the chromosome-centric initiative (C-HPP) and the “biology and disease driven” initiative (B/D- Received in revised form 11 December 2015 HPP). In recent years several reports highlighted how mitochondrial biology and disease are regulated by Accepted 5 January 2016 specific interactions with non-mitochondrial proteins. Thus, it is of great relevance to extend our present Available online 7 January 2016 view of the mitochondrial proteome not only to those proteins that are encoded by or transported to mitochondria, but also to their interactors that take part in mitochondria functionality. Here, we propose a graphical representation of the functional mitochondrial proteome by retrieving mitochondrial proteins from the NeXtProt database and adding to the network their interactors as annotated in the IntAct database. -

2018 Metrics from the HUPO Human Proteome Project † ● ‡ § ∥ Gilbert S
Perspective Cite This: J. Proteome Res. 2018, 17, 4031−4041 pubs.acs.org/jpr Progress on Identifying and Characterizing the Human Proteome: 2018 Metrics from the HUPO Human Proteome Project † ● ‡ § ∥ Gilbert S. Omenn,*, , Lydie Lane, Christopher M. Overall, Fernando J. Corrales, ⊥ # ¶ ∇ ○ Jochen M. Schwenk, Young-Ki Paik, Jennifer E. Van Eyk, Siqi Liu, Michael Snyder, ⊗ ● Mark S. Baker, and Eric W. Deutsch † Department of Computational Medicine and Bioinformatics, University of Michigan, 100 Washtenaw Avenue, Ann Arbor, Michigan 48109-2218, United States ‡ CALIPHO Group, SIB Swiss Institute of Bioinformatics and Department of Microbiology and Molecular Medicine, Faculty of Medicine, University of Geneva, CMU, Michel-Servet 1, 1211 Geneva 4, Switzerland § Life Sciences Institute, Faculty of Dentistry, University of British Columbia, 2350 Health Sciences Mall, Room 4.401, Vancouver, BC V6T 1Z3, Canada ∥ Centro Nacional de Biotecnologia (CSIC), Darwin 3, 28049 Madrid, Spain ⊥ Science for Life Laboratory, KTH Royal Institute of Technology, Tomtebodavagen̈ 23A, 17165 Solna, Sweden # Yonsei Proteome Research Center, Yonsei University, Room 425, Building #114, 50 Yonsei-ro, Seodaemoon-ku, Seoul 120-749, Korea ¶ Advanced Clinical BioSystems Research Institute, Cedars Sinai Precision Biomarker Laboratories, Barbra Streisand Women’s Heart Center, Cedars-Sinai Medical Center, Los Angeles, California 90048, United States ∇ Department of Molecular Biology, University of Texas Southwestern Medical Center, Dallas, Texas 75390-9148, United States ○ Department -
![Comparative Rna-Seq Analysis of Phenotypically Different Sweet Potatoes (Ipomoea Batatas [L.] Lam)](https://docslib.b-cdn.net/cover/0936/comparative-rna-seq-analysis-of-phenotypically-different-sweet-potatoes-ipomoea-batatas-l-lam-3150936.webp)
Comparative Rna-Seq Analysis of Phenotypically Different Sweet Potatoes (Ipomoea Batatas [L.] Lam)
COMPARATIVE RNA-SEQ ANALYSIS OF PHENOTYPICALLY DIFFERENT SWEET POTATOES (IPOMOEA BATATAS [L.] LAM) by ELIZABETH FIEDLER A THESIS Submitted in partial fulfillment of the requirements for the degree of Master of Science in the Agricultural Sciences Graduate Program of Delaware State University DOVER, DELAWARE May 2019 This Thesis is approved by the following members of the Final Oral Review Committee: Dr. Venu (Kal) Kalavacharla, Committee Chairperson, Department of Agriculture and Natural Resources, Delaware State University Dr. Richard Barczewski, Committee Member, Department of Agriculture and Natural Resources, Delaware State University Dr. Marikis Alvarez, Committee Member, Department of Agriculture and Natural Resources, Delaware State University Dr. Muthusamy Manoharan, External Committee Member, Department of Agriculture, Fisheries and Human Sciences, University of Arkansas at Pine Bluff Comparative RNA-Seq Analysis of Phenotypically Different Sweet Potatoes (Ipomoea batatas [L.] Lam) Elizabeth Fiedler Faculty Advisor: Dr. Venu (Kal) Kalavacharla ABSTRACT Sweet potato is arguably one of Earth’s top ten most important crops. It is relatively low maintenance, packed with essential vitamins and nutrients, and in addition to serving as an effective food crop, it has been suggested for use as a material for synthesizing plastics and as a replacement for corn as a source for bioethanol production. Sweet potatoes are difficult to bring to seed, so most sweet potato plants are grown from slips, which are cuttings from sweet potato vines. This makes it very easy for sweet potato viruses to spread from generation to generation. Currently, virus disease complexes, which are infections of two or more viruses with a synergistic interaction, pose the biggest threat to sweet potato yields. -
![Arxiv:1705.08730V1 [Cs.DL] 24 May 2017 Addition, We Show That There Are Patterns of Reuse That Have Previously Been Shown to Be Associated with Percolation Errors](https://docslib.b-cdn.net/cover/8556/arxiv-1705-08730v1-cs-dl-24-may-2017-addition-we-show-that-there-are-patterns-of-reuse-that-have-previously-been-shown-to-be-associated-with-percolation-errors-3648556.webp)
Arxiv:1705.08730V1 [Cs.DL] 24 May 2017 Addition, We Show That There Are Patterns of Reuse That Have Previously Been Shown to Be Associated with Percolation Errors
On Patterns and Re-Use in Bioinformatics Databases Michael J Bell, and Phillip Lord, School of Computing Science Newcastle University November 5, 2018 Abstract 1 Motivation: As the quantity of data being depositing into biological databases con- tinues to increase, it becomes ever more vital to develop methods that enable us to understand this data and ensure that the knowledge is cor- rect. It is widely-held that data percolates between different databases, which causes particular concerns for data correctness; if this percolation occurs, incorrect data in one database may eventually affect many oth- ers while, conversely, corrections in one database may fail to percolate to others. In this paper, we test this widely-held belief by directly looking for sentence reuse both within and between databases. Further, we investi- gate patterns of how sentences are reused over time. Finally, we consider the limitations of this form of analysis and the implications that this may have for bioinformatics database design. 2 Results: We show that reuse of annotation is common within many different databases, and that also there is a detectable level of reuse between databases. In arXiv:1705.08730v1 [cs.DL] 24 May 2017 addition, we show that there are patterns of reuse that have previously been shown to be associated with percolation errors. 3 Introduction It is estimated that over 1,500 active databases are currently in existence [Fern´andez- Su´arezand Galperin, 2013]. While these are generally thought of as containing biological data, they often also contain collected and collated information about 1 the data they carry, which is described as annotation.