FcΓ Receptors: Structure, Function and Role As
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Gene Expression, Network Analysis, and Drug Discovery of Neurofibromatosis Type 2-Associated Vestibular Schwannomas Based on Bioinformatics Analysis
Hindawi Journal of Oncology Volume 2020, Article ID 5976465, 9 pages https://doi.org/10.1155/2020/5976465 Research Article Gene Expression, Network Analysis, and Drug Discovery of Neurofibromatosis Type 2-Associated Vestibular Schwannomas Based on Bioinformatics Analysis Qiao Huang , Si-Jia Zhai , Xing-Wei Liao , Yu-Chao Liu, and Shi-Hua Yin Department of Otolaryngology & Head and Neck Surgery, e Second Affiliated Hospital of Guangxi Medical University, Nanning 530007, China Correspondence should be addressed to Shi-Hua Yin; [email protected] Received 26 March 2020; Revised 27 May 2020; Accepted 1 June 2020; Published 15 July 2020 Academic Editor: Pierfrancesco Franco Copyright © 2020 Qiao Huang et al. ,is is an open access article distributed under the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited. Neurofibromatosis Type 2- (NF2-) associated vestibular schwannomas (VSs) are histologically benign tumors. ,is study aimed to determine disease-related genes, pathways, and potential therapeutic drugs associated with NF2-VSs using the bioinformatics method. Microarray data of GSE108524 were downloaded from the Gene Expression Omnibus (GEO) database, and differentially expressed genes (DEGs) were screened using GEO2R. ,e functional enrichment and pathway enrichment of DEGs were performed using Gene Ontology (GO) and Kyoto Encyclopedia of Genes Genomes (KEGG). Furthermore, the STRING and Cytoscape were used to analyze the protein-protein interaction (PPI) network of all differentially expressed genes and identify hub genes. Finally, the enriched gene sets belonging to the identified pathways were queried against the Drug-Gene Interaction database to find drug candidates for topical use in NF2-associated VSs. -

The Ligands for Human Igg and Their Effector Functions
antibodies Review The Ligands for Human IgG and Their Effector Functions Steven W. de Taeye 1,2,*, Theo Rispens 1 and Gestur Vidarsson 2 1 Sanquin Research, Dept Immunopathology and Landsteiner Laboratory, Amsterdam UMC, University of Amsterdam, 1066 CX Amsterdam, The Netherlands; [email protected] 2 Sanquin Research, Dept Experimental Immunohematology and Landsteiner Laboratory, Amsterdam UMC, University of Amsterdam, 1066 CX Amsterdam, The Netherlands; [email protected] * Correspondence: [email protected] Received: 26 March 2019; Accepted: 18 April 2019; Published: 25 April 2019 Abstract: Activation of the humoral immune system is initiated when antibodies recognize an antigen and trigger effector functions through the interaction with Fc engaging molecules. The most abundant immunoglobulin isotype in serum is Immunoglobulin G (IgG), which is involved in many humoral immune responses, strongly interacting with effector molecules. The IgG subclass, allotype, and glycosylation pattern, among other factors, determine the interaction strength of the IgG-Fc domain with these Fc engaging molecules, and thereby the potential strength of their effector potential. The molecules responsible for the effector phase include the classical IgG-Fc receptors (FcγR), the neonatal Fc-receptor (FcRn), the Tripartite motif-containing protein 21 (TRIM21), the first component of the classical complement cascade (C1), and possibly, the Fc-receptor-like receptors (FcRL4/5). Here we provide an overview of the interactions of IgG with effector molecules and discuss how natural variation on the antibody and effector molecule side shapes the biological activities of antibodies. The increasing knowledge on the Fc-mediated effector functions of antibodies drives the development of better therapeutic antibodies for cancer immunotherapy or treatment of autoimmune diseases. -

Associations Between FCGR3A Polymorphisms and Susceptibility to Rheumatoid Arthritis: a Metaanalysis YOUNG HO LEE, JONG DAE JI, and GWAN GYU SONG
Associations Between FCGR3A Polymorphisms and Susceptibility to Rheumatoid Arthritis: A Metaanalysis YOUNG HO LEE, JONG DAE JI, and GWAN GYU SONG ABSTRACT. Objective. To investigate whether the Fcγ receptor (FCGR) polymorphism confers susceptibility to rheumatoid arthritis (RA). Methods. We conducted metaanalyses on the associations between FCGR polymorphisms and RA susceptibility as determined using (1) allele contrast, (2) recessive models, (3) dominant models, and (4) contrast of homozygotes, using fixed or random effects models. Results. A total of 10 separate comparisons were considered, which comprised 6 European and 4 Asian population samples. Metaanalysis of FCGR3A polymorphism revealed a significant associa- tion between the VV genotype and the risk of developing RA relative to the VF+FF genotype (OR 1.256, 95% CI 1.045–1.510, p = 0.015), with no evidence of between-study heterogeneity (p = 0.167). In subjects of European descent, a stronger association was observed between the VV geno- type and RA than for the FF genotype (OR 1.374, 95% CI 1.101–1.714, p = 0.005). In Asians, no such association was found. Metaanalysis of the VV vs FF genotype revealed a significantly increased OR in Europeans (OR 1.399, 95% CI 1.107–1.769, p = 0.005), but not in Asians. No asso- ciation was found between RA and the FCGR2A and FCGR3B polymorphisms in all subjects and in European and Asian populations, except for the NA22 vs NA11 of FCGR3B in Europeans. Conclusion. No relation was found between the FCGR2A polymorphism and susceptibility to RA in Europeans or Asians. The FCGR3A polymorphism was found to be associated with RA in Europeans but not in Asians. -

FCGR2B) Is Associated with the Production of Anti-Cyclic Citrullinated Peptide Autoantibodies in Taiwanese RA
Genes and Immunity (2008) 9, 680–688 & 2008 Macmillan Publishers Limited All rights reserved 1466-4879/08 $32.00 www.nature.com/gene ORIGINAL ARTICLE A transmembrane polymorphism in FcgRIIb (FCGR2B) is associated with the production of anti-cyclic citrullinated peptide autoantibodies in Taiwanese RA J-Y Chen1, C-M Wang2, C-C Ma1, L-A Hsu3, H-H Ho1, Y-JJ Wu1, S-N Kuo1 and J Wu4 1Division of Allergy, Immunology and Rheumatology, Department of Medicine, Chang Gung Memorial Hospital, Chang Gung University College of Medicine, Tao-Yuan, Taiwan, Republic of China; 2Department of Rehabilitation, Chang Gung Memorial Hospital, Chang Gung University College of Medicine, Tao-Yuan, Taiwan, Republic of China; 3Department of Medicine, Division of First Cardiovascular, Chang Gung Memorial Hospital, Chang Gung University College of Medicine, Tao-Yuan, Taiwan, Republic of China and 4Division of Clinical Immunology and Rheumatology, Department of Medicine, University of Alabama at Birmingham, Birmingham, AL, USA The aim of the current study was to determine whether the FcgRIIb 187-Ile/Thr polymorphism is a predisposition factor for subtypes of RA defined by disease severity and production of autoantibodies against cyclic citrullinated peptides (anti-CCPs) in Taiwanese RA patients. Genotype distributions and allele frequencies of FcgRIIb 187-Ile/Thr were compared between 562 normal healthy controls and 640 RA patients as stratified by clinical parameters and autoantibodies. Significant enrichment of 187-Ile allele was observed in RA patients positive for anti-CCP antibodies as compared with the anti-CCP negative RA patients (P ¼ 0.001, OR 1.652 (95% CI 1.210–2.257)) or as compared with the normal controls (P ¼ 0.005, OR 1.348 (95% CI 1.092–1.664)). -

Supplementary Table 1: Adhesion Genes Data Set
Supplementary Table 1: Adhesion genes data set PROBE Entrez Gene ID Celera Gene ID Gene_Symbol Gene_Name 160832 1 hCG201364.3 A1BG alpha-1-B glycoprotein 223658 1 hCG201364.3 A1BG alpha-1-B glycoprotein 212988 102 hCG40040.3 ADAM10 ADAM metallopeptidase domain 10 133411 4185 hCG28232.2 ADAM11 ADAM metallopeptidase domain 11 110695 8038 hCG40937.4 ADAM12 ADAM metallopeptidase domain 12 (meltrin alpha) 195222 8038 hCG40937.4 ADAM12 ADAM metallopeptidase domain 12 (meltrin alpha) 165344 8751 hCG20021.3 ADAM15 ADAM metallopeptidase domain 15 (metargidin) 189065 6868 null ADAM17 ADAM metallopeptidase domain 17 (tumor necrosis factor, alpha, converting enzyme) 108119 8728 hCG15398.4 ADAM19 ADAM metallopeptidase domain 19 (meltrin beta) 117763 8748 hCG20675.3 ADAM20 ADAM metallopeptidase domain 20 126448 8747 hCG1785634.2 ADAM21 ADAM metallopeptidase domain 21 208981 8747 hCG1785634.2|hCG2042897 ADAM21 ADAM metallopeptidase domain 21 180903 53616 hCG17212.4 ADAM22 ADAM metallopeptidase domain 22 177272 8745 hCG1811623.1 ADAM23 ADAM metallopeptidase domain 23 102384 10863 hCG1818505.1 ADAM28 ADAM metallopeptidase domain 28 119968 11086 hCG1786734.2 ADAM29 ADAM metallopeptidase domain 29 205542 11085 hCG1997196.1 ADAM30 ADAM metallopeptidase domain 30 148417 80332 hCG39255.4 ADAM33 ADAM metallopeptidase domain 33 140492 8756 hCG1789002.2 ADAM7 ADAM metallopeptidase domain 7 122603 101 hCG1816947.1 ADAM8 ADAM metallopeptidase domain 8 183965 8754 hCG1996391 ADAM9 ADAM metallopeptidase domain 9 (meltrin gamma) 129974 27299 hCG15447.3 ADAMDEC1 ADAM-like, -

GP130 Cytokines in Breast Cancer and Bone
cancers Review GP130 Cytokines in Breast Cancer and Bone Tolu Omokehinde 1,2 and Rachelle W. Johnson 1,2,3,* 1 Program in Cancer Biology, Vanderbilt University, Nashville, TN 37232, USA; [email protected] 2 Vanderbilt Center for Bone Biology, Department of Medicine, Division of Clinical Pharmacology, Vanderbilt University Medical Center, Nashville, TN 37232, USA 3 Department of Medicine, Division of Clinical Pharmacology, Vanderbilt University Medical Center, Nashville, TN 37232, USA * Correspondence: [email protected]; Tel.: +1-615-875-8965 Received: 14 December 2019; Accepted: 29 January 2020; Published: 31 January 2020 Abstract: Breast cancer cells have a high predilection for skeletal homing, where they may either induce osteolytic bone destruction or enter a latency period in which they remain quiescent. Breast cancer cells produce and encounter autocrine and paracrine cytokine signals in the bone microenvironment, which can influence their behavior in multiple ways. For example, these signals can promote the survival and dormancy of bone-disseminated cancer cells or stimulate proliferation. The interleukin-6 (IL-6) cytokine family, defined by its use of the glycoprotein 130 (gp130) co-receptor, includes interleukin-11 (IL-11), leukemia inhibitory factor (LIF), oncostatin M (OSM), ciliary neurotrophic factor (CNTF), and cardiotrophin-1 (CT-1), among others. These cytokines are known to have overlapping pleiotropic functions in different cell types and are important for cross-talk between bone-resident cells. IL-6 cytokines have also been implicated in the progression and metastasis of breast, prostate, lung, and cervical cancer, highlighting the importance of these cytokines in the tumor–bone microenvironment. This review will describe the role of these cytokines in skeletal remodeling and cancer progression both within and outside of the bone microenvironment. -

Single‑Nucleotide Polymorphisms and Copy Number Variations of the FCGR2A and FCGR3A Genes in Healthy Japanese Subjects
BIOMEDICAL REPORTS 2: 265-269, 2014 Single‑nucleotide polymorphisms and copy number variations of the FCGR2A and FCGR3A genes in healthy Japanese subjects HIROYUKI MORIYA1, KATSUHIKO SAITO1,2, NUALA HELSBY3, NAOMI HAYASHI1, SHIGEKAZU SUGINO4, MICHIAKI YAMAKAGE4, TAKERU SAWAGUCHI5, MASAHIKO TAKASAKI5, MASATO TAKAHASHI6 and NAHOKO KUROSAWA1 1Department of Pharmacy, Hokkaido Pharmaceutical University School of Pharmacy, Otaru, Hokkaido 047-0264; 2Department of Pharmacy, Sapporo Hokuyu Hospital, Sapporo, Hokkaido 003-0006, Japan; 3Department of Molecular Medicine and Pathology, Faculty of Medical and Health Sciences, University of Auckland, Auckland 1142, New Zealand; 4Department of Anesthesiology, Sapporo Medical University School of Medicine, Sapporo, Hokkaido 060-8543; Departments of 5Pharmacy and 6Breast Surgery, National Hospital Organization Hokkaido Cancer Center, Sapporo, Hokkaido 003-0804, Japan Received October 24, 2013; Accepted November 18, 2013 DOI: 10.3892/br.2013.210 Abstract. FcγRII and FcγRIII are low-affinity Fcγ recep- previous findings regarding FCGR2A and FCGR3A alleles and tors that are encoded by the FCGR2A and FCGR3A genes, CNVs. These assays may provide a basis for the investigation of respectively. These genes contain functional single-nucleotide the role of these genes in the efficacy of antibody-based drugs, polymorphisms (SNPs), which alter the binding affinities of such as trastuzumab and rituximab, in Japanese subjects. these receptors for the γ chain of the Fc fragment of immu- noglobulin G. The known SNPs in FCGR2A and FCGR3A Introduction are rs1801274 (A>G; H131R) and rs396991 (T>G; F158V), respectively. It is also known that there are copy number varia- Fcγ receptors (FcγRs) bind specifically to the γ chain of the tions (CNVs) in the genetic locus (1q23) where FCGR2A and Fc fragment of immunoglobulin G (IgG) and are located on the FCGR3A are located. -

Mouse Model Recapitulating Human Fcγ Receptor Structural and Functional Diversity
Mouse model recapitulating human Fcγ receptor structural and functional diversity Patrick Smith1, David J. DiLillo1, Stylianos Bournazos, Fubin Li, and Jeffrey V. Ravetch2 Laboratory of Molecular Genetics and Immunology, The Rockefeller University, New York, NY 10021 Contributed by Jeffrey V. Ravetch, March 7, 2012 (sent for review March 2, 2012) The in vivo biological activities of IgG antibodies result from their a particular species, such that the absolute affinities of IgG sub- bifunctional nature, in which antigen recognition by the Fab is classes for their cognate FcγRs cannot be extrapolated between coupled to the effector and immunomodulatory diversity found in species, even for recently diverged human and primate species (1, the Fc domain. This diversity, resulting from both amino acid and 12). This situation is further complicated by the existence of poly- γ γ glycan heterogeneity, is translated into cellular responses through morphisms in the human population for Fc RIIA and Fc RIIIA γ γ that result in different affinities for huIgGs (13–16), as well as Fc receptors (Fc Rs), a structurally and functionally diverse family γ of cell surface receptors found throughout the immune system. polymorphisms in Fc RIIB regulating its level of expression or Although many of the overall features of this system are main- signaling (17). Attempts to model huIgG interactions with human FcγR-expressing cells in vitro fail to mirror the diversity of cellular tained throughout mammalian evolution, species diversity has pre- populations that may be required for an in vivo response. There- cluded direct analysis of human antibodies in animal species, and, fore, new systems to study the in vivo function of the huFcγRsystem thus, detailed investigations into the unique features of the human γ and the biological effects of engaging the activating and inhibitory IgG antibodies and their Fc Rs have been limited. -

Integrated Weighted Gene Co-Expression Network Analysis Identi Ed That TLR2 and CD40 Are Related to Coronary Artery Disease
Integrated weighted gene co-expression network analysis identied that TLR2 and CD40 are related to coronary artery disease Bin Qi Liuzhou People's Hospital Jian-Hong Chen Liuzhou People's Hospital Lin Tao Liuzhou People's Hospital Chuan-Meng Zhu Liuzhou People's Hospital Yong Wang Liuzhou People's Hospital Guo-Xiong Deng The First People's of Nanning Liu Miao ( [email protected] ) LiuZhou People's Hospital https://orcid.org/0000-0001-6642-7005 Research article Keywords: Gene Expression Omnibus, Integrated weighted gene co-expression network analysis (WGCNA), Functional enrichment, Functional validation and prognostic analysis. Posted Date: October 7th, 2020 DOI: https://doi.org/10.21203/rs.3.rs-86115/v1 License: This work is licensed under a Creative Commons Attribution 4.0 International License. Read Full License Page 1/16 Abstract Background: The current research attempted to identify possible hub genes and pathways of coronary artery disease (CAD) and to detect the possible mechanisms. Methods: Array data from GSE90074 were downloaded from the Gene Expression Omnibus (GEO) database. Integrated weighted gene co-expression network analysis (WGCNA) was performed to analyze the gene module and clinical characteristics. Gene Ontology annotation, Disease Ontology and the Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment analyses were performed by clusterProler and the DOSE package in R. A protein-protein interaction (PPI) network was established using Cytoscape software, and signicant modules were analyzed using Molecular Complex Detection to identify hub genes. Then, further functional validation of hub genes in other microarrays and population samples was performed, and survival analysis was performed to investigate the prognosis. -

First Infusion Reactions Are Mediated by Fcγriiib and Neutrophils
Pharm Res (2018) 35: 169 https://doi.org/10.1007/s11095-018-2448-8 RESEARCH PAPER First Infusion Reactions are Mediated by FcγRIIIb and Neutrophils Felix Weber 1 & Daniel Breustedt 1,2 & Sonja Schlicht 3 & Claas A. Meyer 3 & Jens Niewoehner 4 & Martin Ebeling 1 & Per-Ola Freskgard5 & Peter Bruenker6 & Thomas Singer1 & Michael Reth7 & Antonio Iglesias1 Received: 29 March 2018 /Accepted: 15 June 2018 /Published online: 27 June 2018 # The Author(s) 2018 ABSTRACT FIR. FcγRIIIb-mediated FIR was abolished by depleting neu- Purpose Administration of therapeutic monoclonal antibod- trophils or blocking FcγRIIIb with CD11b antibodies. ies (mAbs) is frequently accompanied by severe first infusion Conclusions Human FcγRIIIb and neutrophils are primarily reactions (FIR). The mechanism driving FIR is still unclear. responsible for triggering FIR. Clinical strategies to prevent This study aimed to investigate the cellular and molecular FIR in patients should focus on this pathway and may include mechanisms causing FIR in humanized mouse models and transient depletion of neutrophils or blocking FcγRIIIb with their potential for evaluating FIR risk in patients. specific mAbs. Methods Mice humanized for Fc gamma receptors (FcγRs) were generated by recombination-mediated genomic replace- KEY WORDS human FcγRIIIb . humanized mouse model . ment. Body temperature, cytokine release and reactive oxygen immunotoxicology . infusion reactions . neutrophils species (ROS) were measured to assess FIR to mAbs. Results Infusion of human mAb specific for mouse transferrin receptor (HamTfR) into FcγR-humanized mice, produced marked transient hypothermia accompanied by an increase ABBREVIATIONS in inflammatory cytokines KC and MIP-2, and ROS. FIR ADA Anti-drug antibodies were dependent on administration route and Fc-triggered ef- ADCC Antibody-dependent cellular cytotoxicity fector functions mediated by neutrophils. -
Type of the Paper (Article
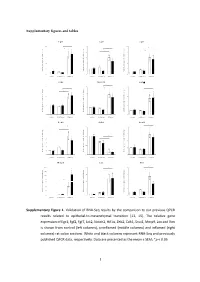
Supplementary figures and tables E g r 1 F g f2 F g f7 1 0 * 5 1 0 * * e e e * g g g * n n n * a a a 8 4 * 8 h h h * c c c d d d * l l l o o o * f f f * n n n o o o 6 3 6 i i i s s s s s s e e e r r r p p p x x x e e e 4 2 4 e e e n n n e e e g g g e e e v v v i i i t t t 2 1 2 a a a l l l e e e R R R 0 0 0 c o n tro l u n in fla m e d in fla m e d c o n tro l u n in fla m e d in fla m e d c o n tro l u n in fla m e d in fla m e d J a k 2 N o tc h 2 H if1 * 3 4 6 * * * e e e g g g n n n a a * * a * h h * h c c c 3 * d d * d l l l * o o o f f 2 f 4 n n n o o o i i i s s s s s s e e e r r 2 r p p p x x x e e e e e e n n n e e 1 e 2 g g g e e 1 e v v v i i i t t t a a a l l l e e e R R R 0 0 0 c o n tro l u n in fla m e d in fla m e d c o n tro l u n in fla m e d in fla m e d c o n tro l u n in fla m e d in fla m e d Z e b 2 C d h 1 S n a i1 * * 7 1 .5 4 * * e e e g g g 6 n n n * a a a * h h h c c c 3 * d d d l l l 5 o o o f f f 1 .0 * n n n * o o o i i i 4 * s s s s s s e e e r r r 2 p p p x x x 3 e e e e e e n n n e e e 0 .5 g g g 2 e e e 1 v v v i i i t t t a a a * l l l e e e 1 * R R R 0 0 .0 0 c o n tro l u n in fla m e d in fla m e d c o n tro l u n in fla m e d in fla m e d c o n tro l u n in fla m e d in fla m e d M m p 9 L o x V im 2 0 0 2 0 8 * * * e e e * g g g 1 5 0 * n n n * a a a * h h h * c c c 1 5 * 6 d d d l l l 1 0 0 o o o f f f n n n o o o i i i 5 0 s s s s s s * e e e r r r 1 0 4 3 0 p p p * x x x e e e * e e e n n n e e e 2 0 g g g e e e 5 2 v v v i i i t t t a a a l l l 1 0 e e e R R R 0 0 0 c o n tro l u n in fla m e d in fla m e d c o n tro l u n in fla m e d in fla m e d c o n tro l u n in fla m e d in fla m e d Supplementary Figure 1. -

Development and Validation of a Protein-Based Risk Score for Cardiovascular Outcomes Among Patients with Stable Coronary Heart Disease
Supplementary Online Content Ganz P, Heidecker B, Hveem K, et al. Development and validation of a protein-based risk score for cardiovascular outcomes among patients with stable coronary heart disease. JAMA. doi: 10.1001/jama.2016.5951 eTable 1. List of 1130 Proteins Measured by Somalogic’s Modified Aptamer-Based Proteomic Assay eTable 2. Coefficients for Weibull Recalibration Model Applied to 9-Protein Model eFigure 1. Median Protein Levels in Derivation and Validation Cohort eTable 3. Coefficients for the Recalibration Model Applied to Refit Framingham eFigure 2. Calibration Plots for the Refit Framingham Model eTable 4. List of 200 Proteins Associated With the Risk of MI, Stroke, Heart Failure, and Death eFigure 3. Hazard Ratios of Lasso Selected Proteins for Primary End Point of MI, Stroke, Heart Failure, and Death eFigure 4. 9-Protein Prognostic Model Hazard Ratios Adjusted for Framingham Variables eFigure 5. 9-Protein Risk Scores by Event Type This supplementary material has been provided by the authors to give readers additional information about their work. Downloaded From: https://jamanetwork.com/ on 10/02/2021 Supplemental Material Table of Contents 1 Study Design and Data Processing ......................................................................................................... 3 2 Table of 1130 Proteins Measured .......................................................................................................... 4 3 Variable Selection and Statistical Modeling ........................................................................................