Associations Between FCGR3A Polymorphisms and Susceptibility to Rheumatoid Arthritis: a Metaanalysis YOUNG HO LEE, JONG DAE JI, and GWAN GYU SONG
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Gene Expression, Network Analysis, and Drug Discovery of Neurofibromatosis Type 2-Associated Vestibular Schwannomas Based on Bioinformatics Analysis
Hindawi Journal of Oncology Volume 2020, Article ID 5976465, 9 pages https://doi.org/10.1155/2020/5976465 Research Article Gene Expression, Network Analysis, and Drug Discovery of Neurofibromatosis Type 2-Associated Vestibular Schwannomas Based on Bioinformatics Analysis Qiao Huang , Si-Jia Zhai , Xing-Wei Liao , Yu-Chao Liu, and Shi-Hua Yin Department of Otolaryngology & Head and Neck Surgery, e Second Affiliated Hospital of Guangxi Medical University, Nanning 530007, China Correspondence should be addressed to Shi-Hua Yin; [email protected] Received 26 March 2020; Revised 27 May 2020; Accepted 1 June 2020; Published 15 July 2020 Academic Editor: Pierfrancesco Franco Copyright © 2020 Qiao Huang et al. ,is is an open access article distributed under the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited. Neurofibromatosis Type 2- (NF2-) associated vestibular schwannomas (VSs) are histologically benign tumors. ,is study aimed to determine disease-related genes, pathways, and potential therapeutic drugs associated with NF2-VSs using the bioinformatics method. Microarray data of GSE108524 were downloaded from the Gene Expression Omnibus (GEO) database, and differentially expressed genes (DEGs) were screened using GEO2R. ,e functional enrichment and pathway enrichment of DEGs were performed using Gene Ontology (GO) and Kyoto Encyclopedia of Genes Genomes (KEGG). Furthermore, the STRING and Cytoscape were used to analyze the protein-protein interaction (PPI) network of all differentially expressed genes and identify hub genes. Finally, the enriched gene sets belonging to the identified pathways were queried against the Drug-Gene Interaction database to find drug candidates for topical use in NF2-associated VSs. -

CD226 T Cells Expressing the Receptors TIGIT and Divergent Phenotypes of Human Regulatory
The Journal of Immunology Divergent Phenotypes of Human Regulatory T Cells Expressing the Receptors TIGIT and CD226 Christopher A. Fuhrman,*,1 Wen-I Yeh,*,1 Howard R. Seay,* Priya Saikumar Lakshmi,* Gaurav Chopra,† Lin Zhang,* Daniel J. Perry,* Stephanie A. McClymont,† Mahesh Yadav,† Maria-Cecilia Lopez,‡ Henry V. Baker,‡ Ying Zhang,x Yizheng Li,{ Maryann Whitley,{ David von Schack,x Mark A. Atkinson,* Jeffrey A. Bluestone,‡ and Todd M. Brusko* Regulatory T cells (Tregs) play a central role in counteracting inflammation and autoimmunity. A more complete understanding of cellular heterogeneity and the potential for lineage plasticity in human Treg subsets may identify markers of disease pathogenesis and facilitate the development of optimized cellular therapeutics. To better elucidate human Treg subsets, we conducted direct transcriptional profiling of CD4+FOXP3+Helios+ thymic-derived Tregs and CD4+FOXP3+Helios2 T cells, followed by comparison with CD4+FOXP32Helios2 T conventional cells. These analyses revealed that the coinhibitory receptor T cell Ig and ITIM domain (TIGIT) was highly expressed on thymic-derived Tregs. TIGIT and the costimulatory factor CD226 bind the common ligand CD155. Thus, we analyzed the cellular distribution and suppressive activity of isolated subsets of CD4+CD25+CD127lo/2 T cells expressing CD226 and/or TIGIT. We observed TIGIT is highly expressed and upregulated on Tregs after activation and in vitro expansion, and is associated with lineage stability and suppressive capacity. Conversely, the CD226+TIGIT2 population was associated with reduced Treg purity and suppressive capacity after expansion, along with a marked increase in IL-10 and effector cytokine production. These studies provide additional markers to delineate functionally distinct Treg subsets that may help direct cellular therapies and provide important phenotypic markers for assessing the role of Tregs in health and disease. -

Single-Cell RNA Sequencing Demonstrates the Molecular and Cellular Reprogramming of Metastatic Lung Adenocarcinoma
ARTICLE https://doi.org/10.1038/s41467-020-16164-1 OPEN Single-cell RNA sequencing demonstrates the molecular and cellular reprogramming of metastatic lung adenocarcinoma Nayoung Kim 1,2,3,13, Hong Kwan Kim4,13, Kyungjong Lee 5,13, Yourae Hong 1,6, Jong Ho Cho4, Jung Won Choi7, Jung-Il Lee7, Yeon-Lim Suh8,BoMiKu9, Hye Hyeon Eum 1,2,3, Soyean Choi 1, Yoon-La Choi6,10,11, Je-Gun Joung1, Woong-Yang Park 1,2,6, Hyun Ae Jung12, Jong-Mu Sun12, Se-Hoon Lee12, ✉ ✉ Jin Seok Ahn12, Keunchil Park12, Myung-Ju Ahn 12 & Hae-Ock Lee 1,2,3,6 1234567890():,; Advanced metastatic cancer poses utmost clinical challenges and may present molecular and cellular features distinct from an early-stage cancer. Herein, we present single-cell tran- scriptome profiling of metastatic lung adenocarcinoma, the most prevalent histological lung cancer type diagnosed at stage IV in over 40% of all cases. From 208,506 cells populating the normal tissues or early to metastatic stage cancer in 44 patients, we identify a cancer cell subtype deviating from the normal differentiation trajectory and dominating the metastatic stage. In all stages, the stromal and immune cell dynamics reveal ontological and functional changes that create a pro-tumoral and immunosuppressive microenvironment. Normal resident myeloid cell populations are gradually replaced with monocyte-derived macrophages and dendritic cells, along with T-cell exhaustion. This extensive single-cell analysis enhances our understanding of molecular and cellular dynamics in metastatic lung cancer and reveals potential diagnostic and therapeutic targets in cancer-microenvironment interactions. 1 Samsung Genome Institute, Samsung Medical Center, Seoul 06351, Korea. -

Associations Between FCGR3A Polymorphisms and Susceptibility to Rheumatoid Arthritis: a Metaanalysis YOUNG HO LEE, JONG DAE JI, and GWAN GYU SONG
Associations Between FCGR3A Polymorphisms and Susceptibility to Rheumatoid Arthritis: A Metaanalysis YOUNG HO LEE, JONG DAE JI, and GWAN GYU SONG ABSTRACT. Objective. To investigate whether the Fcγ receptor (FCGR) polymorphism confers susceptibility to rheumatoid arthritis (RA). Methods. We conducted metaanalyses on the associations between FCGR polymorphisms and RA susceptibility as determined using (1) allele contrast, (2) recessive models, (3) dominant models, and (4) contrast of homozygotes, using fixed or random effects models. Results. A total of 10 separate comparisons were considered, which comprised 6 European and 4 Asian population samples. Metaanalysis of FCGR3A polymorphism revealed a significant associa- tion between the VV genotype and the risk of developing RA relative to the VF+FF genotype (OR 1.256, 95% CI 1.045–1.510, p = 0.015), with no evidence of between-study heterogeneity (p = 0.167). In subjects of European descent, a stronger association was observed between the VV geno- type and RA than for the FF genotype (OR 1.374, 95% CI 1.101–1.714, p = 0.005). In Asians, no such association was found. Metaanalysis of the VV vs FF genotype revealed a significantly increased OR in Europeans (OR 1.399, 95% CI 1.107–1.769, p = 0.005), but not in Asians. No asso- ciation was found between RA and the FCGR2A and FCGR3B polymorphisms in all subjects and in European and Asian populations, except for the NA22 vs NA11 of FCGR3B in Europeans. Conclusion. No relation was found between the FCGR2A polymorphism and susceptibility to RA in Europeans or Asians. The FCGR3A polymorphism was found to be associated with RA in Europeans but not in Asians. -

Single-Cell RNA Sequencing of Tocilizumab-Treated Peripheral Blood Mononuclear Cells As an in Vitro Model of Inflammation
UCSF UC San Francisco Previously Published Works Title Single-Cell RNA Sequencing of Tocilizumab-Treated Peripheral Blood Mononuclear Cells as an in vitro Model of Inflammation. Permalink https://escholarship.org/uc/item/8q84n3jb Authors Zarinsefat, Arya Hartoularos, George Rychkov, Dmitry et al. Publication Date 2020 DOI 10.3389/fgene.2020.610682 Peer reviewed eScholarship.org Powered by the California Digital Library University of California fgene-11-610682 December 19, 2020 Time: 19:44 # 1 BRIEF RESEARCH REPORT published: 05 January 2021 doi: 10.3389/fgene.2020.610682 Single-Cell RNA Sequencing of Tocilizumab-Treated Peripheral Blood Mononuclear Cells as an in vitro Model of Inflammation Arya Zarinsefat1*, George Hartoularos2, Dmitry Rychkov1, Priyanka Rashmi1, Sindhu Chandran3, Flavio Vincenti1, Chun J. Yee2 and Minnie M. Sarwal1 1 Department of Surgery, University of California, San Francisco, San Francisco, CA, United States, 2 Department of Bioengineering and Therapeutic Sciences, University of California, San Francisco, San Francisco, CA, United States, 3 Department of Medicine, University of California, San Francisco, San Francisco, CA, United States COVID-19 has posed a significant threat to global health. Early data has revealed that IL-6, a key regulatory cytokine, plays an important role in the cytokine storm of COVID-19. Multiple trials are therefore looking at the effects of Tocilizumab, an IL-6 receptor antibody that inhibits IL-6 activity, on treatment of COVID-19, with promising findings. As part of a clinical trial looking at the effects of Tocilizumab treatment on Edited by: kidney transplant recipients with subclinical rejection, we performed single-cell RNA Gabriele Bucci, sequencing of comparing stimulated PBMCs before and after Tocilizumab treatment. -

CD29 Identifies IFN-Γ–Producing Human CD8+ T Cells With
+ CD29 identifies IFN-γ–producing human CD8 T cells with an increased cytotoxic potential Benoît P. Nicoleta,b, Aurélie Guislaina,b, Floris P. J. van Alphenc, Raquel Gomez-Eerlandd, Ton N. M. Schumacherd, Maartje van den Biggelaarc,e, and Monika C. Wolkersa,b,1 aDepartment of Hematopoiesis, Sanquin Research, 1066 CX Amsterdam, The Netherlands; bLandsteiner Laboratory, Oncode Institute, Amsterdam University Medical Center, University of Amsterdam, 1105 AZ Amsterdam, The Netherlands; cDepartment of Research Facilities, Sanquin Research, 1066 CX Amsterdam, The Netherlands; dDivision of Molecular Oncology and Immunology, Oncode Institute, The Netherlands Cancer Institute, 1066 CX Amsterdam, The Netherlands; and eDepartment of Molecular and Cellular Haemostasis, Sanquin Research, 1066 CX Amsterdam, The Netherlands Edited by Anjana Rao, La Jolla Institute for Allergy and Immunology, La Jolla, CA, and approved February 12, 2020 (received for review August 12, 2019) Cytotoxic CD8+ T cells can effectively kill target cells by producing therefore developed a protocol that allowed for efficient iso- cytokines, chemokines, and granzymes. Expression of these effector lation of RNA and protein from fluorescence-activated cell molecules is however highly divergent, and tools that identify and sorting (FACS)-sorted fixed T cells after intracellular cytokine + preselect CD8 T cells with a cytotoxic expression profile are lacking. staining. With this top-down approach, we performed an un- + Human CD8 T cells can be divided into IFN-γ– and IL-2–producing biased RNA-sequencing (RNA-seq) and mass spectrometry cells. Unbiased transcriptomics and proteomics analysis on cytokine- γ– – + + (MS) analyses on IFN- and IL-2 producing primary human producing fixed CD8 T cells revealed that IL-2 cells produce helper + + + CD8 Tcells. -

FCGR Polymorphisms Influence Response to IL2 in Metastatic Renal Cell Carcinoma
Published OnlineFirst October 14, 2016; DOI: 10.1158/1078-0432.CCR-16-1874 Cancer Therapy: Clinical Clinical Cancer Research FCGR Polymorphisms Influence Response to IL2 in Metastatic Renal Cell Carcinoma Amy K. Erbe1,WeiWang1, Jacob Goldberg1, Mikayla Gallenberger1, KyungMann Kim2, Lakeesha Carmichael2,DustinHess1,EneidaA.Mendonca2,3, Yiqiang Song2, Jacquelyn A. Hank1,Su-ChunCheng4, Sabina Signoretti5, Michael Atkins6,7, Alexander Carlson8,JamesW.Mier6,8, David J. Panka8, David F. McDermott6,8,and Paul M. Sondel1,3 Abstract Purpose: Fc-gamma receptors (FCGRs) are expressed on Results: When higher-affinity genotypes for FCGR2A, FCGR3A, immune cells, bind to antibodies, and trigger antibody-induced and FCGR2C were considered together, they were associated with cell-mediated antitumor responses when tumor-reactive antibo- significantly increased tumor shrinkage and prolonged survival in dies are present. The affinity of the FCGR/antibody interaction is response to HD-IL2. variable and dependent upon FCGR polymorphisms. Prior stud- Conclusions: Although associations of higher-affinity FCGR ies of patients with cancer treated with immunotherapy indicate genotype with clinical outcome have been demonstrated that FCGR polymorphisms can influence antitumor response for with mAb therapy and with idiotype vaccines, to our knowl- certain immunotherapies that act via therapeutically administered edge,thisisthefirst study to show associations of FCGR mAbs or via endogenous tumor-reactive antibodies induced from genotypes with outcome following HD-IL2 treatment. We tumor antigen vaccines. The previously published "SELECT" trial hypothesize that endogenous antitumor antibodies may of high-dose aldesleukin (HD-IL2) for metastatic renal cell carci- engage immune cells through their FCGRs, and HD-IL2 may noma resulted in an objective response rate of 25%. -

Single‑Nucleotide Polymorphisms and Copy Number Variations of the FCGR2A and FCGR3A Genes in Healthy Japanese Subjects
BIOMEDICAL REPORTS 2: 265-269, 2014 Single‑nucleotide polymorphisms and copy number variations of the FCGR2A and FCGR3A genes in healthy Japanese subjects HIROYUKI MORIYA1, KATSUHIKO SAITO1,2, NUALA HELSBY3, NAOMI HAYASHI1, SHIGEKAZU SUGINO4, MICHIAKI YAMAKAGE4, TAKERU SAWAGUCHI5, MASAHIKO TAKASAKI5, MASATO TAKAHASHI6 and NAHOKO KUROSAWA1 1Department of Pharmacy, Hokkaido Pharmaceutical University School of Pharmacy, Otaru, Hokkaido 047-0264; 2Department of Pharmacy, Sapporo Hokuyu Hospital, Sapporo, Hokkaido 003-0006, Japan; 3Department of Molecular Medicine and Pathology, Faculty of Medical and Health Sciences, University of Auckland, Auckland 1142, New Zealand; 4Department of Anesthesiology, Sapporo Medical University School of Medicine, Sapporo, Hokkaido 060-8543; Departments of 5Pharmacy and 6Breast Surgery, National Hospital Organization Hokkaido Cancer Center, Sapporo, Hokkaido 003-0804, Japan Received October 24, 2013; Accepted November 18, 2013 DOI: 10.3892/br.2013.210 Abstract. FcγRII and FcγRIII are low-affinity Fcγ recep- previous findings regarding FCGR2A and FCGR3A alleles and tors that are encoded by the FCGR2A and FCGR3A genes, CNVs. These assays may provide a basis for the investigation of respectively. These genes contain functional single-nucleotide the role of these genes in the efficacy of antibody-based drugs, polymorphisms (SNPs), which alter the binding affinities of such as trastuzumab and rituximab, in Japanese subjects. these receptors for the γ chain of the Fc fragment of immu- noglobulin G. The known SNPs in FCGR2A and FCGR3A Introduction are rs1801274 (A>G; H131R) and rs396991 (T>G; F158V), respectively. It is also known that there are copy number varia- Fcγ receptors (FcγRs) bind specifically to the γ chain of the tions (CNVs) in the genetic locus (1q23) where FCGR2A and Fc fragment of immunoglobulin G (IgG) and are located on the FCGR3A are located. -

Integrated Weighted Gene Co-Expression Network Analysis Identi Ed That TLR2 and CD40 Are Related to Coronary Artery Disease
Integrated weighted gene co-expression network analysis identied that TLR2 and CD40 are related to coronary artery disease Bin Qi Liuzhou People's Hospital Jian-Hong Chen Liuzhou People's Hospital Lin Tao Liuzhou People's Hospital Chuan-Meng Zhu Liuzhou People's Hospital Yong Wang Liuzhou People's Hospital Guo-Xiong Deng The First People's of Nanning Liu Miao ( [email protected] ) LiuZhou People's Hospital https://orcid.org/0000-0001-6642-7005 Research article Keywords: Gene Expression Omnibus, Integrated weighted gene co-expression network analysis (WGCNA), Functional enrichment, Functional validation and prognostic analysis. Posted Date: October 7th, 2020 DOI: https://doi.org/10.21203/rs.3.rs-86115/v1 License: This work is licensed under a Creative Commons Attribution 4.0 International License. Read Full License Page 1/16 Abstract Background: The current research attempted to identify possible hub genes and pathways of coronary artery disease (CAD) and to detect the possible mechanisms. Methods: Array data from GSE90074 were downloaded from the Gene Expression Omnibus (GEO) database. Integrated weighted gene co-expression network analysis (WGCNA) was performed to analyze the gene module and clinical characteristics. Gene Ontology annotation, Disease Ontology and the Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment analyses were performed by clusterProler and the DOSE package in R. A protein-protein interaction (PPI) network was established using Cytoscape software, and signicant modules were analyzed using Molecular Complex Detection to identify hub genes. Then, further functional validation of hub genes in other microarrays and population samples was performed, and survival analysis was performed to investigate the prognosis. -

First Infusion Reactions Are Mediated by Fcγriiib and Neutrophils
Pharm Res (2018) 35: 169 https://doi.org/10.1007/s11095-018-2448-8 RESEARCH PAPER First Infusion Reactions are Mediated by FcγRIIIb and Neutrophils Felix Weber 1 & Daniel Breustedt 1,2 & Sonja Schlicht 3 & Claas A. Meyer 3 & Jens Niewoehner 4 & Martin Ebeling 1 & Per-Ola Freskgard5 & Peter Bruenker6 & Thomas Singer1 & Michael Reth7 & Antonio Iglesias1 Received: 29 March 2018 /Accepted: 15 June 2018 /Published online: 27 June 2018 # The Author(s) 2018 ABSTRACT FIR. FcγRIIIb-mediated FIR was abolished by depleting neu- Purpose Administration of therapeutic monoclonal antibod- trophils or blocking FcγRIIIb with CD11b antibodies. ies (mAbs) is frequently accompanied by severe first infusion Conclusions Human FcγRIIIb and neutrophils are primarily reactions (FIR). The mechanism driving FIR is still unclear. responsible for triggering FIR. Clinical strategies to prevent This study aimed to investigate the cellular and molecular FIR in patients should focus on this pathway and may include mechanisms causing FIR in humanized mouse models and transient depletion of neutrophils or blocking FcγRIIIb with their potential for evaluating FIR risk in patients. specific mAbs. Methods Mice humanized for Fc gamma receptors (FcγRs) were generated by recombination-mediated genomic replace- KEY WORDS human FcγRIIIb . humanized mouse model . ment. Body temperature, cytokine release and reactive oxygen immunotoxicology . infusion reactions . neutrophils species (ROS) were measured to assess FIR to mAbs. Results Infusion of human mAb specific for mouse transferrin receptor (HamTfR) into FcγR-humanized mice, produced marked transient hypothermia accompanied by an increase ABBREVIATIONS in inflammatory cytokines KC and MIP-2, and ROS. FIR ADA Anti-drug antibodies were dependent on administration route and Fc-triggered ef- ADCC Antibody-dependent cellular cytotoxicity fector functions mediated by neutrophils. -
Type of the Paper (Article
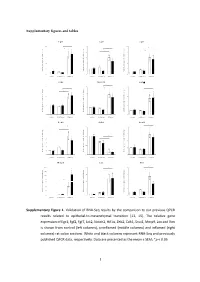
Supplementary figures and tables E g r 1 F g f2 F g f7 1 0 * 5 1 0 * * e e e * g g g * n n n * a a a 8 4 * 8 h h h * c c c d d d * l l l o o o * f f f * n n n o o o 6 3 6 i i i s s s s s s e e e r r r p p p x x x e e e 4 2 4 e e e n n n e e e g g g e e e v v v i i i t t t 2 1 2 a a a l l l e e e R R R 0 0 0 c o n tro l u n in fla m e d in fla m e d c o n tro l u n in fla m e d in fla m e d c o n tro l u n in fla m e d in fla m e d J a k 2 N o tc h 2 H if1 * 3 4 6 * * * e e e g g g n n n a a * * a * h h * h c c c 3 * d d * d l l l * o o o f f 2 f 4 n n n o o o i i i s s s s s s e e e r r 2 r p p p x x x e e e e e e n n n e e 1 e 2 g g g e e 1 e v v v i i i t t t a a a l l l e e e R R R 0 0 0 c o n tro l u n in fla m e d in fla m e d c o n tro l u n in fla m e d in fla m e d c o n tro l u n in fla m e d in fla m e d Z e b 2 C d h 1 S n a i1 * * 7 1 .5 4 * * e e e g g g 6 n n n * a a a * h h h c c c 3 * d d d l l l 5 o o o f f f 1 .0 * n n n * o o o i i i 4 * s s s s s s e e e r r r 2 p p p x x x 3 e e e e e e n n n e e e 0 .5 g g g 2 e e e 1 v v v i i i t t t a a a * l l l e e e 1 * R R R 0 0 .0 0 c o n tro l u n in fla m e d in fla m e d c o n tro l u n in fla m e d in fla m e d c o n tro l u n in fla m e d in fla m e d M m p 9 L o x V im 2 0 0 2 0 8 * * * e e e * g g g 1 5 0 * n n n * a a a * h h h * c c c 1 5 * 6 d d d l l l 1 0 0 o o o f f f n n n o o o i i i 5 0 s s s s s s * e e e r r r 1 0 4 3 0 p p p * x x x e e e * e e e n n n e e e 2 0 g g g e e e 5 2 v v v i i i t t t a a a l l l 1 0 e e e R R R 0 0 0 c o n tro l u n in fla m e d in fla m e d c o n tro l u n in fla m e d in fla m e d c o n tro l u n in fla m e d in fla m e d Supplementary Figure 1. -

Development and Validation of a Protein-Based Risk Score for Cardiovascular Outcomes Among Patients with Stable Coronary Heart Disease
Supplementary Online Content Ganz P, Heidecker B, Hveem K, et al. Development and validation of a protein-based risk score for cardiovascular outcomes among patients with stable coronary heart disease. JAMA. doi: 10.1001/jama.2016.5951 eTable 1. List of 1130 Proteins Measured by Somalogic’s Modified Aptamer-Based Proteomic Assay eTable 2. Coefficients for Weibull Recalibration Model Applied to 9-Protein Model eFigure 1. Median Protein Levels in Derivation and Validation Cohort eTable 3. Coefficients for the Recalibration Model Applied to Refit Framingham eFigure 2. Calibration Plots for the Refit Framingham Model eTable 4. List of 200 Proteins Associated With the Risk of MI, Stroke, Heart Failure, and Death eFigure 3. Hazard Ratios of Lasso Selected Proteins for Primary End Point of MI, Stroke, Heart Failure, and Death eFigure 4. 9-Protein Prognostic Model Hazard Ratios Adjusted for Framingham Variables eFigure 5. 9-Protein Risk Scores by Event Type This supplementary material has been provided by the authors to give readers additional information about their work. Downloaded From: https://jamanetwork.com/ on 10/02/2021 Supplemental Material Table of Contents 1 Study Design and Data Processing ......................................................................................................... 3 2 Table of 1130 Proteins Measured .......................................................................................................... 4 3 Variable Selection and Statistical Modeling ........................................................................................