Identification of Candidate Biomarkers and Pathways Associated with Type 1 Diabetes Mellitus Using Bioinformatics Analysis
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Analysis of Trans Esnps Infers Regulatory Network Architecture
Analysis of trans eSNPs infers regulatory network architecture Anat Kreimer Submitted in partial fulfillment of the requirements for the degree of Doctor of Philosophy in the Graduate School of Arts and Sciences COLUMBIA UNIVERSITY 2014 © 2014 Anat Kreimer All rights reserved ABSTRACT Analysis of trans eSNPs infers regulatory network architecture Anat Kreimer eSNPs are genetic variants associated with transcript expression levels. The characteristics of such variants highlight their importance and present a unique opportunity for studying gene regulation. eSNPs affect most genes and their cell type specificity can shed light on different processes that are activated in each cell. They can identify functional variants by connecting SNPs that are implicated in disease to a molecular mechanism. Examining eSNPs that are associated with distal genes can provide insights regarding the inference of regulatory networks but also presents challenges due to the high statistical burden of multiple testing. Such association studies allow: simultaneous investigation of many gene expression phenotypes without assuming any prior knowledge and identification of unknown regulators of gene expression while uncovering directionality. This thesis will focus on such distal eSNPs to map regulatory interactions between different loci and expose the architecture of the regulatory network defined by such interactions. We develop novel computational approaches and apply them to genetics-genomics data in human. We go beyond pairwise interactions to define network motifs, including regulatory modules and bi-fan structures, showing them to be prevalent in real data and exposing distinct attributes of such arrangements. We project eSNP associations onto a protein-protein interaction network to expose topological properties of eSNPs and their targets and highlight different modes of distal regulation. -

Edinburgh Research Explorer
Edinburgh Research Explorer International Union of Basic and Clinical Pharmacology. LXXXVIII. G protein-coupled receptor list Citation for published version: Davenport, AP, Alexander, SPH, Sharman, JL, Pawson, AJ, Benson, HE, Monaghan, AE, Liew, WC, Mpamhanga, CP, Bonner, TI, Neubig, RR, Pin, JP, Spedding, M & Harmar, AJ 2013, 'International Union of Basic and Clinical Pharmacology. LXXXVIII. G protein-coupled receptor list: recommendations for new pairings with cognate ligands', Pharmacological reviews, vol. 65, no. 3, pp. 967-86. https://doi.org/10.1124/pr.112.007179 Digital Object Identifier (DOI): 10.1124/pr.112.007179 Link: Link to publication record in Edinburgh Research Explorer Document Version: Publisher's PDF, also known as Version of record Published In: Pharmacological reviews Publisher Rights Statement: U.S. Government work not protected by U.S. copyright General rights Copyright for the publications made accessible via the Edinburgh Research Explorer is retained by the author(s) and / or other copyright owners and it is a condition of accessing these publications that users recognise and abide by the legal requirements associated with these rights. Take down policy The University of Edinburgh has made every reasonable effort to ensure that Edinburgh Research Explorer content complies with UK legislation. If you believe that the public display of this file breaches copyright please contact [email protected] providing details, and we will remove access to the work immediately and investigate your claim. Download date: 02. Oct. 2021 1521-0081/65/3/967–986$25.00 http://dx.doi.org/10.1124/pr.112.007179 PHARMACOLOGICAL REVIEWS Pharmacol Rev 65:967–986, July 2013 U.S. -

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

Supplemental Materials ZNF281 Enhances Cardiac Reprogramming
Supplemental Materials ZNF281 enhances cardiac reprogramming by modulating cardiac and inflammatory gene expression Huanyu Zhou, Maria Gabriela Morales, Hisayuki Hashimoto, Matthew E. Dickson, Kunhua Song, Wenduo Ye, Min S. Kim, Hanspeter Niederstrasser, Zhaoning Wang, Beibei Chen, Bruce A. Posner, Rhonda Bassel-Duby and Eric N. Olson Supplemental Table 1; related to Figure 1. Supplemental Table 2; related to Figure 1. Supplemental Table 3; related to the “quantitative mRNA measurement” in Materials and Methods section. Supplemental Table 4; related to the “ChIP-seq, gene ontology and pathway analysis” and “RNA-seq” and gene ontology analysis” in Materials and Methods section. Supplemental Figure S1; related to Figure 1. Supplemental Figure S2; related to Figure 2. Supplemental Figure S3; related to Figure 3. Supplemental Figure S4; related to Figure 4. Supplemental Figure S5; related to Figure 6. Supplemental Table S1. Genes included in human retroviral ORF cDNA library. Gene Gene Gene Gene Gene Gene Gene Gene Symbol Symbol Symbol Symbol Symbol Symbol Symbol Symbol AATF BMP8A CEBPE CTNNB1 ESR2 GDF3 HOXA5 IL17D ADIPOQ BRPF1 CEBPG CUX1 ESRRA GDF6 HOXA6 IL17F ADNP BRPF3 CERS1 CX3CL1 ETS1 GIN1 HOXA7 IL18 AEBP1 BUD31 CERS2 CXCL10 ETS2 GLIS3 HOXB1 IL19 AFF4 C17ORF77 CERS4 CXCL11 ETV3 GMEB1 HOXB13 IL1A AHR C1QTNF4 CFL2 CXCL12 ETV7 GPBP1 HOXB5 IL1B AIMP1 C21ORF66 CHIA CXCL13 FAM3B GPER HOXB6 IL1F3 ALS2CR8 CBFA2T2 CIR1 CXCL14 FAM3D GPI HOXB7 IL1F5 ALX1 CBFA2T3 CITED1 CXCL16 FASLG GREM1 HOXB9 IL1F6 ARGFX CBFB CITED2 CXCL3 FBLN1 GREM2 HOXC4 IL1F7 -

Patient-Based Cross-Platform Comparison of Oligonucleotide Microarray Expression Profiles
Laboratory Investigation (2005) 85, 1024–1039 & 2005 USCAP, Inc All rights reserved 0023-6837/05 $30.00 www.laboratoryinvestigation.org Patient-based cross-platform comparison of oligonucleotide microarray expression profiles Joerg Schlingemann1,*, Negusse Habtemichael2,*, Carina Ittrich3, Grischa Toedt1, Heidi Kramer1, Markus Hambek4, Rainald Knecht4, Peter Lichter1, Roland Stauber2 and Meinhard Hahn1 1Division of Molecular Genetics, Deutsches Krebsforschungszentrum, Heidelberg, Germany; 2Chemotherapeutisches Forschungsinstitut Georg-Speyer-Haus, Frankfurt am Main, Germany; 3Central Unit Biostatistics, Deutsches Krebsforschungszentrum, Heidelberg, Germany and 4Department of Otorhinolaryngology, Universita¨tsklinik, Johann-Wolfgang-Goethe-Universita¨t Frankfurt, Frankfurt, Germany The comparison of gene expression measurements obtained with different technical approaches is of substantial interest in order to clarify whether interplatform differences may conceal biologically significant information. To address this concern, we analyzed gene expression in a set of head and neck squamous cell carcinoma patients, using both spotted oligonucleotide microarrays made from a large collection of 70-mer probes and commercial arrays produced by in situ synthesis of sets of multiple 25-mer oligonucleotides per gene. Expression measurements were compared for 4425 genes represented on both platforms, which revealed strong correlations between the corresponding data sets. Of note, a global tendency towards smaller absolute ratios was observed when -

Angiotensin-II and Arginine Vasopressin Regulate Body Fluid
View metadata, citation and similar papers at core.ac.uk brought to you by CORE provided by Apollo 1 Angiotensin II type 1 receptor dependent GLP-1 and PYY secretion in mice and 2 humans 3 4 Ramona Pais, Juraj Rievaj, Pierre Larraufie, Fiona Gribble and Frank Reimann 5 6 Wellcome Trust-MRC Institute of Metabolic Science, Metabolic Research Laboratories, 7 University of Cambridge, Addenbrooke’s Hospital, Box 289, Hills Road, Cambridge, CB2 8 0QQ, United Kingdom 9 10 Abbreviated Title: Control of L-cells by Angiotensin II 11 12 Key terms: Renin-Angiotensin System (RAS); Angiotensin II; glucagon-like peptide-1 (GLP- 13 1); L-cells; Peptide YY (PYY) 14 15 Word count: 4652 16 17 No of figures and tables: 6 figures, 0 tables 18 19 Correspondence and reprint requests should be addressed to: 20 Dr. Frank Reimann and Prof. Fiona Gribble 21 Wellcome Trust-MRC Institute of Metabolic Science, Metabolic Research Laboratories 22 Addenbrooke’s Hospital, Box 289 23 University of Cambridge 24 Hills Road, Cambridge, CB2 0QQ, United Kingdom 25 Phone: +44 (0)1223 336746 26 Fax: + 44 (0)1223 330598 27 Email: [email protected], [email protected] 28 29 Disclosure statement: The authors have nothing to disclose. 30 Abstract 31 Angiotensin II (Ang II) is the key hormone mediator of the renin angiotensin system which 32 regulates blood pressure and fluid and electrolyte balance in the body. Here we report that in 33 colonic epithelium the Ang II type 1 receptor (AT1R) is highly and exclusively expressed in 34 enteroendocrine L-cells which produce the gut hormones glucagon-like peptide-1 (GLP-1) 35 and peptide YY (PYY). -

Supplementary Table 1: Adhesion Genes Data Set
Supplementary Table 1: Adhesion genes data set PROBE Entrez Gene ID Celera Gene ID Gene_Symbol Gene_Name 160832 1 hCG201364.3 A1BG alpha-1-B glycoprotein 223658 1 hCG201364.3 A1BG alpha-1-B glycoprotein 212988 102 hCG40040.3 ADAM10 ADAM metallopeptidase domain 10 133411 4185 hCG28232.2 ADAM11 ADAM metallopeptidase domain 11 110695 8038 hCG40937.4 ADAM12 ADAM metallopeptidase domain 12 (meltrin alpha) 195222 8038 hCG40937.4 ADAM12 ADAM metallopeptidase domain 12 (meltrin alpha) 165344 8751 hCG20021.3 ADAM15 ADAM metallopeptidase domain 15 (metargidin) 189065 6868 null ADAM17 ADAM metallopeptidase domain 17 (tumor necrosis factor, alpha, converting enzyme) 108119 8728 hCG15398.4 ADAM19 ADAM metallopeptidase domain 19 (meltrin beta) 117763 8748 hCG20675.3 ADAM20 ADAM metallopeptidase domain 20 126448 8747 hCG1785634.2 ADAM21 ADAM metallopeptidase domain 21 208981 8747 hCG1785634.2|hCG2042897 ADAM21 ADAM metallopeptidase domain 21 180903 53616 hCG17212.4 ADAM22 ADAM metallopeptidase domain 22 177272 8745 hCG1811623.1 ADAM23 ADAM metallopeptidase domain 23 102384 10863 hCG1818505.1 ADAM28 ADAM metallopeptidase domain 28 119968 11086 hCG1786734.2 ADAM29 ADAM metallopeptidase domain 29 205542 11085 hCG1997196.1 ADAM30 ADAM metallopeptidase domain 30 148417 80332 hCG39255.4 ADAM33 ADAM metallopeptidase domain 33 140492 8756 hCG1789002.2 ADAM7 ADAM metallopeptidase domain 7 122603 101 hCG1816947.1 ADAM8 ADAM metallopeptidase domain 8 183965 8754 hCG1996391 ADAM9 ADAM metallopeptidase domain 9 (meltrin gamma) 129974 27299 hCG15447.3 ADAMDEC1 ADAM-like, -

Cdk15 Igfals Lingo4 Gjb3 Tpbg Lrrc38 Serpinf1 Apod Trp73 Lama4 Chrnd Col9a1col11a1col5a2 Fgl2 Pitx2 Col2a1 Col3a1 Lamb3 Col24a1
Bnc2 Wdr72 Ptchd1 Abtb2 Spag5 Zfp385a Trim17 Ier2 Il1rapl1 Tpd52l1 Fam20a Car8 Syt5 Plxnc1 Sema3e Ndrg4 Snph St6galnac5 Mcpt2 B3galt2 Sphkap Arhgap24 Prss34 Lhfpl2 Ermap Rnf165 Shroom1 Grm4 Mobp Dock2 Tmem9b Slc35d3 Otud7b Serpinb3a Sh3d19 Syt6 Zan Trim67 Clec18a Mcoln1 Tob1 Slc45a2 Pcdhb9 Pcdh17 Plscr1 Gpr143 Cela1 Frem1 Sema3f Lgi2 Igsf9 Fjx1 Cpne4 Adgb Depdc7 Gzmm C1qtnf5 Capn11 Sema3c H2-T22 Unc5c Sytl4 Galnt5 Sytl2 Arhgap11a Pcdha1 Cdh20 Slc35f2 Trim29 B3gnt5 Dock5 Trim9 Padi4 Pcdh19 Abi2 Cldn11 Slitrk1 Fam13a Nrgn Cpa4 Clmp Il1rap Trpm1 Fat4 Nexn Pmel Mmp15 Fat3 H2-M5 Prss38 Wdr41 Prtg Mlana Mettl22 Tnrc6b Cdh6 Sema3b Ptgfrn Cldn1 Cntn4 Bcl2a1b Capn6 Capn5 Pcdhb19 Tcf15 Bmf Rgs8 Tecrl Tyrp1 Rhot1 Rnf123 Cldn6 Adam9 Hlx Rilpl1 Disp1 Atcay Vwc2 Fat2 Srpx2 Cldn3 Unc13c Creb3l1 Rab39b Robo3 Gpnmb Bves Orai2 Slc22a2 Prss8 Cdh10 Scg3 Adam33 Nyx Dchs1 Chmp4c Syt9 Ap1m2 Megf10 Cthrc1 Penk Igsf9b Akap2 Ltbp3 Dnmbp Tff2 Pnoc Vldlr Cpa3 Snx18 Capn3 Btla Htr1b Gm17231 Pcdh9Rab27a Grm8 Cnih2 Scube2 Id2 Reep1 Cpeb3 Mmp16 Slc18b1 Snx33 Clcn5 Cckbr Pkp2 Drp2 Mapk8ip1 Lrrc3b Cxcl14 Zfhx3 Esrp1 Prx Dock3 Sec14l1 Prokr1 Pstpip2 Usp2 Cpvl Syn2 Ntn1 Ptger1 Rxfp3 Tyr Snap91 Htr1d Mtnr1a Gadd45g Mlph Drd4 Foxc2 Cldn4 Birc7 Cdh17 Twist2 Scnn1b Abcc4 Pkp1 Dlk2 Rab3b Amph Mreg Il33 Slit2 Hpse Micu1 Creb3l2 Dsp Lifr S1pr5 Krt15 Svep1 Ahnak Kcnh1 Sphk1 Vwce Clcf1 Ptch2 Pmp22 Sfrp1 Sema6a Lfng Hs3st5 Efcab1 Tlr5 Muc5acKalrn Vwa2 Fzd8 Lpar6 Bmp5 Slc16a9 Cacng4 Arvcf Igfbp2 Mrvi1 Dusp15 Krt5 Atp13a5 Dsg1a Kcnj14 Edn3Memo1 Ngef Prickle2 Cma1 Alx4 Bmp3 Blnk GastAgtr2 -

Cellular and Molecular Signatures in the Disease Tissue of Early
Cellular and Molecular Signatures in the Disease Tissue of Early Rheumatoid Arthritis Stratify Clinical Response to csDMARD-Therapy and Predict Radiographic Progression Frances Humby1,* Myles Lewis1,* Nandhini Ramamoorthi2, Jason Hackney3, Michael Barnes1, Michele Bombardieri1, Francesca Setiadi2, Stephen Kelly1, Fabiola Bene1, Maria di Cicco1, Sudeh Riahi1, Vidalba Rocher-Ros1, Nora Ng1, Ilias Lazorou1, Rebecca E. Hands1, Desiree van der Heijde4, Robert Landewé5, Annette van der Helm-van Mil4, Alberto Cauli6, Iain B. McInnes7, Christopher D. Buckley8, Ernest Choy9, Peter Taylor10, Michael J. Townsend2 & Costantino Pitzalis1 1Centre for Experimental Medicine and Rheumatology, William Harvey Research Institute, Barts and The London School of Medicine and Dentistry, Queen Mary University of London, Charterhouse Square, London EC1M 6BQ, UK. Departments of 2Biomarker Discovery OMNI, 3Bioinformatics and Computational Biology, Genentech Research and Early Development, South San Francisco, California 94080 USA 4Department of Rheumatology, Leiden University Medical Center, The Netherlands 5Department of Clinical Immunology & Rheumatology, Amsterdam Rheumatology & Immunology Center, Amsterdam, The Netherlands 6Rheumatology Unit, Department of Medical Sciences, Policlinico of the University of Cagliari, Cagliari, Italy 7Institute of Infection, Immunity and Inflammation, University of Glasgow, Glasgow G12 8TA, UK 8Rheumatology Research Group, Institute of Inflammation and Ageing (IIA), University of Birmingham, Birmingham B15 2WB, UK 9Institute of -

Mucin Family Genes Are Downregulated in Colorectal
ooggeenneessii iinn ss && rrcc aa MM CC uu tt ff aa Journal ofJournal of oo gg Aziz et al., J Carcinogene Mutagene 2014, S10 ll ee ee aa aa nn nn nn nn ee ee rr rr ss ss uu uu ii ii ss ss oo oo DOI: 4172/2157-2518.S10-009 JJ JJ ISSN: 2157-2518 CarCarcinogenesiscinogenesis & Mutagenesis Research Article Article OpenOpen Access Access Mucin Family Genes are Downregulated in Colorectal Cancer Patients Mohammad Azhar Aziz*, Majed AlOtaibi, Abdulkareem AlAbdulrahman, Mohammed AlDrees and Ibrahim AlAbdulkarim Department of Medical Genomics, KIng Abdullah Intl. Med. Res. Ctr., King Saud Bin Abdul Aziz University for Health Sciences, Riyadh, Saudi Arabia Abstract Mucins are very well known to be associated with different types of cancer. Their role in colorectal cancer has been extensively studied without direct correlation with their change in expression levels. In the present study we employed the human exon array from Affymetrix to provide evidence that mucin family genes are downregulated in colorectal cancer tumor samples. We analyzed 92 samples taken from normal and tumor tissues. All mucin family genes except MUCL1 were downregulated with the fold change value ranging from -3.53 to 1.78 as calculated using AltAnalyze software. Maximum drop in RNA transcripts were observed for MUC2 with a fold change of -3.53. Further, we carried out Integromics analysis to analyze mucin genes using hierarchical clustering. MUC1 and MUC4 were found to be potential biomarkers for human colorectal cancer. Top upstream regulators were identified for mucin genes. Network analyses were carried out to further our understanding about potential mechanisms by which mucins can be involved in causing colorectal cancer. -
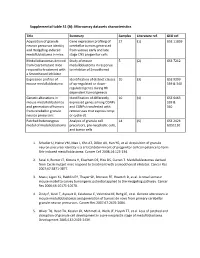
Supplemental Table S1 (A): Microarray Datasets Characteristics
Supplemental table S1 (A): Microarray datasets characteristics Title Summary Samples Literature ref. GEO ref. Acquisition of granule Gene expression profiling of 27 (1) GSE 11859 neuron precursor identity cerebellar tumors generated and Hedgehog‐induced from various early and late medulloblastoma in mice. stage CNS progenitor cells Medulloblastomas derived Study of mouse 5 (2) GSE 7212 from Cxcr6 mutant mice medulloblastoma in response respond to treatment with to inhibitor of Smoothened a Smoothened inhibitor Expression profiles of Identification of distinct classes 10 (3) GSE 9299 mouse medulloblastoma of up‐regulated or down‐ 339 & 340 regulated genes during Hh dependent tumorigenesis Genetic alterations in Identification of differently 10 (4) GSE 6463 mouse medulloblastomas expressed genes among CGNPs 339 & and generation of tumors and CGNPs transfected with 340 from cerebellar granule retroviruses that express nmyc neuron precursors or cyclin‐d1 Patched heterozygous Analysis of granule cell 14 (5) GSE 2426 model of medulloblastoma precursors, pre‐neoplastic cells, GDS1110 and tumor cells 1. Schuller U, Heine VM, Mao J, Kho AT, Dillon AK, Han YG, et al. Acquisition of granule neuron precursor identity is a critical determinant of progenitor cell competence to form Shh‐induced medulloblastoma. Cancer Cell 2008;14:123‐134. 2. Sasai K, Romer JT, Kimura H, Eberhart DE, Rice DS, Curran T. Medulloblastomas derived from Cxcr6 mutant mice respond to treatment with a smoothened inhibitor. Cancer Res 2007;67:3871‐3877. 3. Mao J, Ligon KL, Rakhlin EY, Thayer SP, Bronson RT, Rowitch D, et al. A novel somatic mouse model to survey tumorigenic potential applied to the Hedgehog pathway. Cancer Res 2006;66:10171‐10178. -

Novel and Highly Recurrent Chromosomal Alterations in Se´Zary Syndrome
Research Article Novel and Highly Recurrent Chromosomal Alterations in Se´zary Syndrome Maarten H. Vermeer,1 Remco van Doorn,1 Remco Dijkman,1 Xin Mao,3 Sean Whittaker,3 Pieter C. van Voorst Vader,4 Marie-Jeanne P. Gerritsen,5 Marie-Louise Geerts,6 Sylke Gellrich,7 Ola So¨derberg,8 Karl-Johan Leuchowius,8 Ulf Landegren,8 Jacoba J. Out-Luiting,1 Jeroen Knijnenburg,2 Marije IJszenga,2 Karoly Szuhai,2 Rein Willemze,1 and Cornelis P. Tensen1 Departments of 1Dermatology and 2Molecular Cell Biology, Leiden University Medical Center, Leiden, the Netherlands; 3Department of Dermatology, St Thomas’ Hospital, King’s College, London, United Kingdom; 4Department of Dermatology, University Medical Center Groningen, Groningen, the Netherlands; 5Department of Dermatology, Radboud University Nijmegen Medical Center, Nijmegen, the Netherlands; 6Department of Dermatology, Gent University Hospital, Gent, Belgium; 7Department of Dermatology, Charite, Berlin, Germany; and 8Department of Genetics and Pathology, Rudbeck Laboratory, University of Uppsala, Uppsala, Sweden Abstract Introduction This study was designed to identify highly recurrent genetic Se´zary syndrome (Sz) is an aggressive type of cutaneous T-cell alterations typical of Se´zary syndrome (Sz), an aggressive lymphoma/leukemia of skin-homing, CD4+ memory T cells and is cutaneous T-cell lymphoma/leukemia, possibly revealing characterized by erythroderma, generalized lymphadenopathy, and pathogenetic mechanisms and novel therapeutic targets. the presence of neoplastic T cells (Se´zary cells) in the skin, lymph High-resolution array-based comparative genomic hybridiza- nodes, and peripheral blood (1). Sz has a poor prognosis, with a tion was done on malignant T cells from 20 patients. disease-specific 5-year survival of f24% (1).