Differential Physiological Role of BIN1 Isoforms in Skeletal Muscle Development, Function and Regeneration
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Dynamin Functions and Ligands: Classical Mechanisms Behind
1521-0111/91/2/123–134$25.00 http://dx.doi.org/10.1124/mol.116.105064 MOLECULAR PHARMACOLOGY Mol Pharmacol 91:123–134, February 2017 Copyright ª 2017 by The American Society for Pharmacology and Experimental Therapeutics MINIREVIEW Dynamin Functions and Ligands: Classical Mechanisms Behind Mahaveer Singh, Hemant R. Jadhav, and Tanya Bhatt Department of Pharmacy, Birla Institute of Technology and Sciences Pilani, Pilani Campus, Rajasthan, India Received May 5, 2016; accepted November 17, 2016 Downloaded from ABSTRACT Dynamin is a GTPase that plays a vital role in clathrin-dependent pathophysiology of various disorders, such as Alzheimer’s disease, endocytosis and other vesicular trafficking processes by acting Parkinson’s disease, Huntington’s disease, Charcot-Marie-Tooth as a pair of molecular scissors for newly formed vesicles originating disease, heart failure, schizophrenia, epilepsy, cancer, dominant ’ from the plasma membrane. Dynamins and related proteins are optic atrophy, osteoporosis, and Down s syndrome. This review is molpharm.aspetjournals.org important components for the cleavage of clathrin-coated vesicles, an attempt to illustrate the dynamin-related mechanisms involved phagosomes, and mitochondria. These proteins help in organelle in the above-mentioned disorders and to help medicinal chemists division, viral resistance, and mitochondrial fusion/fission. Dys- to design novel dynamin ligands, which could be useful in the function and mutations in dynamin have been implicated in the treatment of dynamin-related disorders. Introduction GTP hydrolysis–dependent conformational change of GTPase dynamin assists in membrane fission, leading to the generation Dynamins were originally discovered in the brain and identi- of endocytic vesicles (Praefcke and McMahon, 2004; Ferguson at ASPET Journals on September 23, 2021 fied as microtubule binding partners. -

Solitary and Repetitive Binding Motifs for the Ap2 Complex Α-Appendage in Amphiphysin and Other Accessory Proteins
SOLITARY AND REPETITIVE BINDING MOTIFS FOR THE AP2 COMPLEX α-APPENDAGE IN AMPHIPHYSIN AND OTHER ACCESSORY PROTEINS Lene E. Olesen*, Eva M. Schmid*, Marijn G. J. Ford#*, Yvonne Vallis, M. Madan Babu, Peter Li, Ian G. Mills∑, Harvey T. McMahon§ and Gerrit J.K. Praefcke♣§ Laboratory of Molecular Biology, Medical Research Council, Neurobiology Division, Hills Road, Cambridge CB2 2QH, UK. ♣ Center for Molecular Medicine Cologne (CMMC), Institute for Genetics, Zülpicher Straße 47, 50674 Köln, Germany. Running Title: Classification of AP2 α-appendage-binding FxDxF and DxF/W motifs § Address correspondence to: Harvey T. McMahon, Email: [email protected]; Tel.: +44(0)1223-402311; Fax: +44(0)1223-402310 or Gerrit J.K. Praefcke, Email [email protected]; Tel.: +49(0)221-470-1561; Fax: +49(0)221-470-6749 Adaptor protein (AP) complexes bind to coated pits, for the platform subdomain of the transmembrane proteins destined for α-appendage. The motif domain of internalisation and to membrane lipids, so amphiphysin1 contains one copy of each of a linking cargo to the accessory internalisation DxF/W and FxDxF motif. We find that the machinery. This machinery interacts with the FxDxF motif is the main determinant for the appendage domains of APs, which have high affinity interaction with the α-adaptin platform and β-sandwich subdomains, appendage. We describe the optimal sequence forming the binding surfaces for interacting of the FxDxF motif using thermodynamic and proteins. Proteins which interact with the structural data and show how sequence subdomains do so via short motifs, usually variation controls the affinities of these motifs found in regions of low structural complexity for the α-appendage. -

BIN1/M-Amphiphysin2 Induces Clustering of Phosphoinositides to Recruit Its Downstream Partner Dynamin
ARTICLE Received 19 May 2014 | Accepted 22 Oct 2014 | Published 9 Dec 2014 DOI: 10.1038/ncomms6647 BIN1/M-Amphiphysin2 induces clustering of phosphoinositides to recruit its downstream partner dynamin Laura Picas1, Julien Viaud2, Kristine Schauer1, Stefano Vanni3, Karim Hnia4, Vincent Fraisier5, Aure´lien Roux6, Patricia Bassereau7,Fre´de´rique Gaits-Iacovoni2, Bernard Payrastre2, Jocelyn Laporte4, Jean-Baptiste Manneville1 & Bruno Goud1 Phosphoinositides play a central role in many physiological processes by assisting the recruitment of proteins to membranes through specific phosphoinositide-binding motifs. How this recruitment is coordinated in space and time is not well understood. Here we show that BIN1/M-Amphiphysin2, a protein involved in T-tubule biogenesis in muscle cells and fre- quently mutated in centronuclear myopathies, clusters PtdIns(4,5)P2 to recruit its down- stream partner dynamin. By using several mutants associated with centronuclear myopathies, we find that the N-BAR and the SH3 domains of BIN1 control the kinetics and the accumu- lation of dynamin on membranes, respectively. We show that phosphoinositide clustering is a mechanism shared by other proteins that interact with PtdIns(4,5)P2, but do not contain a BAR domain. Our numerical simulations point out that clustering is a diffusion-driven process in which phosphoinositide molecules are not sequestered. We propose that this mechanism plays a key role in the recruitment of downstream phosphoinositide-binding proteins. 1 Institut Curie and CNRS UMR 144, 26 rue d’Ulm, 75005 Paris, France. 2 INSERM, UMR1048, Universite´ Toulouse III, Institut des Maladies Me´taboliques et Cardiovasculaires, 1 avenue Jean Poulhe`s, 31432 Toulouse, France. -

Murine Megakaryopoiesis Is Critical for P21 SCL-Mediated Regulation Of
From bloodjournal.hematologylibrary.org at UNIVERSITY OF BIRMINGHAM on March 1, 2012. For personal use only. 2011 118: 723-735 Prepublished online May 19, 2011; doi:10.1182/blood-2011-01-328765 SCL-mediated regulation of the cell-cycle regulator p21 is critical for murine megakaryopoiesis Hedia Chagraoui, Mira Kassouf, Sreemoti Banerjee, Nicolas Goardon, Kevin Clark, Ann Atzberger, Andrew C. Pearce, Radek C. Skoda, David J. P. Ferguson, Steve P. Watson, Paresh Vyas and Catherine Porcher Updated information and services can be found at: http://bloodjournal.hematologylibrary.org/content/118/3/723.full.html Articles on similar topics can be found in the following Blood collections Platelets and Thrombopoiesis (260 articles) Information about reproducing this article in parts or in its entirety may be found online at: http://bloodjournal.hematologylibrary.org/site/misc/rights.xhtml#repub_requests Information about ordering reprints may be found online at: http://bloodjournal.hematologylibrary.org/site/misc/rights.xhtml#reprints Information about subscriptions and ASH membership may be found online at: http://bloodjournal.hematologylibrary.org/site/subscriptions/index.xhtml Blood (print ISSN 0006-4971, online ISSN 1528-0020), is published weekly by the American Society of Hematology, 2021 L St, NW, Suite 900, Washington DC 20036. Copyright 2011 by The American Society of Hematology; all rights reserved. From bloodjournal.hematologylibrary.org at UNIVERSITY OF BIRMINGHAM on March 1, 2012. For personal use only. PLATELETS AND THROMBOPOIESIS SCL-mediated regulation of the cell-cycle regulator p21 is critical for murine megakaryopoiesis Hedia Chagraoui,1 *Mira Kassouf,1 *Sreemoti Banerjee,1 Nicolas Goardon,1 Kevin Clark,1 Ann Atzberger,1 Andrew C. -

Mechanisms of Synaptic Plasticity Mediated by Clathrin Adaptor-Protein Complexes 1 and 2 in Mice
Mechanisms of synaptic plasticity mediated by Clathrin Adaptor-protein complexes 1 and 2 in mice Dissertation for the award of the degree “Doctor rerum naturalium” at the Georg-August-University Göttingen within the doctoral program “Molecular Biology of Cells” of the Georg-August University School of Science (GAUSS) Submitted by Ratnakar Mishra Born in Birpur, Bihar, India Göttingen, Germany 2019 1 Members of the Thesis Committee Prof. Dr. Peter Schu Institute for Cellular Biochemistry, (Supervisor and first referee) University Medical Center Göttingen, Germany Dr. Hans Dieter Schmitt Neurobiology, Max Planck Institute (Second referee) for Biophysical Chemistry, Göttingen, Germany Prof. Dr. med. Thomas A. Bayer Division of Molecular Psychiatry, University Medical Center, Göttingen, Germany Additional Members of the Examination Board Prof. Dr. Silvio O. Rizzoli Department of Neuro-and Sensory Physiology, University Medical Center Göttingen, Germany Dr. Roland Dosch Institute of Developmental Biochemistry, University Medical Center Göttingen, Germany Prof. Dr. med. Martin Oppermann Institute of Cellular and Molecular Immunology, University Medical Center, Göttingen, Germany Date of oral examination: 14th may 2019 2 Table of Contents List of abbreviations ................................................................................. 5 Abstract ................................................................................................... 7 Chapter 1: Introduction ............................................................................ -

The Alzheimer's Disease Genetic Risk Factor BIN1 Induces Isoform
bioRxiv preprint doi: https://doi.org/10.1101/2021.04.02.438184; this version posted April 4, 2021. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. The Alzheimer’s disease genetic risk factor BIN1 induces isoform-dependent neurotoxicity through early endosome defects Erwan Lambert1,*, Orthis Saha1,*, Bruna Soares Landeira1, Ana Raquel Melo de Farias1, Xavier Hermant1, Arnaud Carrier2, Alexandre Pelletier2, Lindsay Davoine1, Cloé Dupont1, Philippe Amouyel1, Frank Lafont3, Farida Adelfettah1, Patrik Verstreken4,5, Julien Chapuis1, Nicolas Barois3,6, Fabien Delahaye2, Bart Dermaut7, Jean-Charles Lambert1, Marcos R. Costa1,**,#, Pierre Dourlen1,**,# *co-first ** co-last # corresponding author 1. Univ. Lille, Inserm, CHU Lille, Institut Pasteur de Lille, U1167-RID-AGE facteurs de risque et déterminants moléculaires des maladies liées au vieillissement, DISTALZ, Lille, France 2. Univ. Lille, Inserm, CHU Lille, Institut Pasteur de Lille, U1283-UMR 8199 EGID, Lille, France 3. Univ. Lille, CNRS, Inserm, CHU Lille, Institut Pasteur de Lille, U1019-UMR 9017-CIIL-Center for Infection and Immunity of Lille, Lille, France. 4. VIB Center for Brain & Disease Research, KU Leuven, Leuven, Belgium. 5. Department of Neurosciences, Leuven Brain Institute, KU Leuven, Leuven, Belgium. 6. Univ. Lille, CNRS, Inserm, CHU Lille, Institut Pasteur de Lille, US41-UMS2014-PLBS, Lille, France. 7. Centre for Medical Genetics, Ghent University Hospital, Ghent, Belgium Correspondence should be addressed to: Marcos Costa, MD, PhD INSERM UMR1167 Institut Pasteur de Lille 1rue du Pr. -

Sorting Nexin 4 and Amphiphysin 2, a New Partnership Between Endocytosis and Intracellular Trafficking
Research Article 1937 Sorting nexin 4 and amphiphysin 2, a new partnership between endocytosis and intracellular trafficking Corinne Leprince1,*, Erwan Le Scolan1, Brigitte Meunier1, Vincent Fraisier2, Nathalie Brandon1, Jean De Gunzburg1 and Jacques Camonis1 1INSERM U528, 2CNRS UMR144, Institut Curie Section de Recherche, 26 rue d’Ulm, 75248 Paris Cedex 05, France *Author for correspondence (e-mail: [email protected]) Accepted 30 January 2003 Journal of Cell Science 116, 1937-1948 © 2003 The Company of Biologists Ltd doi:10.1242/jcs.00403 Summary Endocytosis is a regulated physiological process by which terminal or full-length SNX4 was able to inhibit transferrin membrane receptors and their extracellular ligands are receptor endocytosis as efficiently as the SH3 domain of internalized. After internalization, they enter the amphiphysin 2. At lower levels of expression, SNX4 endosomal trafficking pathway for sorting and processing. colocalized with transferrin-containing vesicles, some of Amphiphysins consist of a family of proteins conserved which were also positive for amphiphysin 2. These results throughout evolution that are crucial elements of the indicate that SNX4 may be part of the endocytic machinery endocytosis machinery in mammalian cells. They act as or, alternatively, that SNX4 may associate with key adaptors for a series of proteins important for the endocytic elements of endocytosis such as amphiphysin 2 and process, such as dynamin. In order to improve our sequester them when overexpressed. The presence of knowledge of amphiphysin function, we performed a two- amphiphysin 2 on intracellular vesicles and its interplay hybrid screen with the N-terminal part of murine with SNX4, which is likely to take part in intracellular amphiphysin 2 (residues 1-304). -

Regulate Macrophage Phagocytosis Phospholipases D1 and D2
Phospholipases D1 and D2 Coordinately Regulate Macrophage Phagocytosis Shankar S. Iyer, James A. Barton, Sylvain Bourgoin and David J. Kusner This information is current as of September 26, 2021. J Immunol 2004; 173:2615-2623; ; doi: 10.4049/jimmunol.173.4.2615 http://www.jimmunol.org/content/173/4/2615 Downloaded from References This article cites 79 articles, 55 of which you can access for free at: http://www.jimmunol.org/content/173/4/2615.full#ref-list-1 Why The JI? Submit online. http://www.jimmunol.org/ • Rapid Reviews! 30 days* from submission to initial decision • No Triage! Every submission reviewed by practicing scientists • Fast Publication! 4 weeks from acceptance to publication *average by guest on September 26, 2021 Subscription Information about subscribing to The Journal of Immunology is online at: http://jimmunol.org/subscription Permissions Submit copyright permission requests at: http://www.aai.org/About/Publications/JI/copyright.html Email Alerts Receive free email-alerts when new articles cite this article. Sign up at: http://jimmunol.org/alerts The Journal of Immunology is published twice each month by The American Association of Immunologists, Inc., 1451 Rockville Pike, Suite 650, Rockville, MD 20852 Copyright © 2004 by The American Association of Immunologists All rights reserved. Print ISSN: 0022-1767 Online ISSN: 1550-6606. The Journal of Immunology Phospholipases D1 and D2 Coordinately Regulate Macrophage Phagocytosis1 Shankar S. Iyer,* James A. Barton,* Sylvain Bourgoin,§ and David J. Kusner,2*†‡ Phagocytosis is a fundamental feature of the innate immune system, required for antimicrobial defense, resolution of inflamma- tion, and tissue remodeling. Furthermore, phagocytosis is coupled to a diverse range of cytotoxic effector mechanisms, including the respiratory burst, secretion of inflammatory mediators and Ag presentation. -

Integrating Protein Copy Numbers with Interaction Networks to Quantify Stoichiometry in Mammalian Endocytosis
bioRxiv preprint doi: https://doi.org/10.1101/2020.10.29.361196; this version posted October 29, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-ND 4.0 International license. Integrating protein copy numbers with interaction networks to quantify stoichiometry in mammalian endocytosis Daisy Duan1, Meretta Hanson1, David O. Holland2, Margaret E Johnson1* 1TC Jenkins Department of Biophysics, Johns Hopkins University, 3400 N Charles St, Baltimore, MD 21218. 2NIH, Bethesda, MD, 20892. *Corresponding Author: [email protected] bioRxiv preprint doi: https://doi.org/10.1101/2020.10.29.361196; this version posted October 29, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-ND 4.0 International license. Abstract Proteins that drive processes like clathrin-mediated endocytosis (CME) are expressed at various copy numbers within a cell, from hundreds (e.g. auxilin) to millions (e.g. clathrin). Between cell types with identical genomes, copy numbers further vary significantly both in absolute and relative abundance. These variations contain essential information about each protein’s function, but how significant are these variations and how can they be quantified to infer useful functional behavior? Here, we address this by quantifying the stoichiometry of proteins involved in the CME network. We find robust trends across three cell types in proteins that are sub- vs super-stoichiometric in terms of protein function, network topology (e.g. -

Figure S1. HAEC ROS Production and ML090 NOX5-Inhibition
Figure S1. HAEC ROS production and ML090 NOX5-inhibition. (a) Extracellular H2O2 production in HAEC treated with ML090 at different concentrations and 24 h after being infected with GFP and NOX5-β adenoviruses (MOI 100). **p< 0.01, and ****p< 0.0001 vs control NOX5-β-infected cells (ML090, 0 nM). Results expressed as mean ± SEM. Fold increase vs GFP-infected cells with 0 nM of ML090. n= 6. (b) NOX5-β overexpression and DHE oxidation in HAEC. Representative images from three experiments are shown. Intracellular superoxide anion production of HAEC 24 h after infection with GFP and NOX5-β adenoviruses at different MOIs treated or not with ML090 (10 nM). MOI: Multiplicity of infection. Figure S2. Ontology analysis of HAEC infected with NOX5-β. Ontology analysis shows that the response to unfolded protein is the most relevant. Figure S3. UPR mRNA expression in heart of infarcted transgenic mice. n= 12-13. Results expressed as mean ± SEM. Table S1: Altered gene expression due to NOX5-β expression at 12 h (bold, highlighted in yellow). N12hvsG12h N18hvsG18h N24hvsG24h GeneName GeneDescription TranscriptID logFC p-value logFC p-value logFC p-value family with sequence similarity NM_052966 1.45 1.20E-17 2.44 3.27E-19 2.96 6.24E-21 FAM129A 129. member A DnaJ (Hsp40) homolog. NM_001130182 2.19 9.83E-20 2.94 2.90E-19 3.01 1.68E-19 DNAJA4 subfamily A. member 4 phorbol-12-myristate-13-acetate- NM_021127 0.93 1.84E-12 2.41 1.32E-17 2.69 1.43E-18 PMAIP1 induced protein 1 E2F7 E2F transcription factor 7 NM_203394 0.71 8.35E-11 2.20 2.21E-17 2.48 1.84E-18 DnaJ (Hsp40) homolog. -
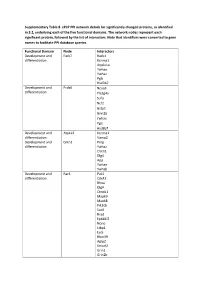
Supplementary Table 8. Cpcp PPI Network Details for Significantly Changed Proteins, As Identified in 3.2, Underlying Each of the Five Functional Domains
Supplementary Table 8. cPCP PPI network details for significantly changed proteins, as identified in 3.2, underlying each of the five functional domains. The network nodes represent each significant protein, followed by the list of interactors. Note that identifiers were converted to gene names to facilitate PPI database queries. Functional Domain Node Interactors Development and Park7 Rack1 differentiation Kcnma1 Atp6v1a Ywhae Ywhaz Pgls Hsd3b7 Development and Prdx6 Ncoa3 differentiation Pla2g4a Sufu Ncf2 Gstp1 Grin2b Ywhae Pgls Hsd3b7 Development and Atp1a2 Kcnma1 differentiation Vamp2 Development and Cntn1 Prnp differentiation Ywhaz Clstn1 Dlg4 App Ywhae Ywhab Development and Rac1 Pak1 differentiation Cdc42 Rhoa Dlg4 Ctnnb1 Mapk9 Mapk8 Pik3cb Sod1 Rrad Epb41l2 Nono Ltbp1 Evi5 Rbm39 Aplp2 Smurf2 Grin1 Grin2b Xiap Chn2 Cav1 Cybb Pgls Ywhae Development and Hbb-b1 Atp5b differentiation Hba Kcnma1 Got1 Aldoa Ywhaz Pgls Hsd3b4 Hsd3b7 Ywhae Development and Myh6 Mybpc3 differentiation Prkce Ywhae Development and Amph Capn2 differentiation Ap2a2 Dnm1 Dnm3 Dnm2 Atp6v1a Ywhab Development and Dnm3 Bin1 differentiation Amph Pacsin1 Grb2 Ywhae Bsn Development and Eef2 Ywhaz differentiation Rpgrip1l Atp6v1a Nphp1 Iqcb1 Ezh2 Ywhae Ywhab Pgls Hsd3b7 Hsd3b4 Development and Gnai1 Dlg4 differentiation Development and Gnao1 Dlg4 differentiation Vamp2 App Ywhae Ywhab Development and Psmd3 Rpgrip1l differentiation Psmd4 Hmga2 Development and Thy1 Syp differentiation Atp6v1a App Ywhae Ywhaz Ywhab Hsd3b7 Hsd3b4 Development and Tubb2a Ywhaz differentiation Nphp4 -

BIN1 Gene Bridging Integrator 1
BIN1 gene bridging integrator 1 Normal Function The BIN1 gene provides instructions for making a protein that is found in tissues throughout the body, where it interacts with a variety of other proteins. The BIN1 protein is thought to be involved in the transportation of materials from the cell surface into the cell (endocytosis) and the self-destruction of cells (apoptosis). The BIN1 protein may act as a tumor suppressor protein, which means it prevents cells from growing and dividing too rapidly or in an uncontrolled way. Several different versions (isoforms) of the BIN1 protein are produced from the BIN1 gene. These isoforms vary by size and are active in different tissues. The BIN1 protein isoform that is expressed in muscle cells is thought to be involved in the formation of structures called transverse tubules or T tubules. These structures are found within the membrane of muscle cells, where they play a role in muscle tensing (contraction) and relaxation. Health Conditions Related to Genetic Changes Centronuclear myopathy At least 10 mutations in the BIN1 gene have been found to cause centronuclear myopathy, a condition that is characterized by muscle weakness (myopathy) in the skeletal muscles, which are the muscles used for movement. Most of these mutations change single protein building blocks (amino acids) in the BIN1 protein. BIN1 gene mutations result in the production of a protein that cannot form T tubules. A shortage of T tubules in muscle fibers alters their structure, which prevents them from contracting and relaxing normally. The abnormal muscle fibers underlie the muscle weakness characteristic of centronuclear myopathy.