Protein Folds in the Worm Genome
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Ankrd9 Is a Metabolically-Controlled Regulator of Impdh2 Abundance and Macro-Assembly
ANKRD9 IS A METABOLICALLY-CONTROLLED REGULATOR OF IMPDH2 ABUNDANCE AND MACRO-ASSEMBLY by Dawn Hayward A dissertation submitted to The Johns Hopkins University in conformity with the requirements of the degree of Doctor of Philosophy Baltimore, Maryland April 2019 ABSTRACT Members of a large family of Ankyrin Repeat Domains proteins (ANKRD) regulate numerous cellular processes by binding and changing properties of specific protein targets. We show that interactions with a target protein and the functional outcomes can be markedly altered by cells’ metabolic state. ANKRD9 facilitates degradation of inosine monophosphate dehydrogenase 2 (IMPDH2), the rate-limiting enzyme in GTP biosynthesis. Under basal conditions ANKRD9 is largely segregated from the cytosolic IMPDH2 by binding to vesicles. Upon nutrient limitation, ANKRD9 loses association with vesicles and assembles with IMPDH2 into rod-like structures, in which IMPDH2 is stable. Inhibition of IMPDH2 with Ribavirin favors ANKRD9 binding to rods. The IMPDH2/ANKRD9 assembly is reversed by guanosine, which restores association of ANKRD9 with vesicles. The conserved Cys109Cys110 motif in ANKRD9 is required for the vesicles-to-rods transition as well as binding and regulation of IMPDH2. ANKRD9 knockdown increases IMPDH2 levels and prevents formation of IMPDH2 rods upon nutrient limitation. Thus, the status of guanosine pools affects the mode of ANKRD9 action towards IMPDH2. Advisor: Dr. Svetlana Lutsenko, Department of Physiology, Johns Hopkins University School of Medicine Second reader: -

Repeat Proteins Challenge the Concept of Structural Domains
844 Biochemical Society Transactions (2015) Volume 43, part 5 Repeat proteins challenge the concept of structural domains Rocıo´ Espada*1, R. Gonzalo Parra*1, Manfred J. Sippl†, Thierry Mora‡, Aleksandra M. Walczak§ and Diego U. Ferreiro*2 *Protein Physiology Lab, Dep de Qu´ımica Biologica, ´ Facultad de Ciencias Exactas y Naturales, UBA-CONICET-IQUIBICEN, Buenos Aires, C1430EGA, Argentina †Center of Applied Molecular Engineering, Division of Bioinformatics, Department of Molecular Biology, University of Salzburg, 5020 Salzburg, Austria ‡Laboratoire de physique statistique, CNRS, UPMC and Ecole normale superieure, ´ 24 rue Lhomond, 75005 Paris, France §Laboratoire de physique theorique, ´ CNRS, UPMC and Ecole normale superieure, ´ 24 rue Lhomond, 75005 Paris, France Abstract Structural domains are believed to be modules within proteins that can fold and function independently. Some proteins show tandem repetitions of apparent modular structure that do not fold independently, but rather co-operate in stabilizing structural forms that comprise several repeat-units. For many natural repeat-proteins, it has been shown that weak energetic links between repeats lead to the breakdown of co-operativity and the appearance of folding sub-domains within an apparently regular repeat array. The quasi-1D architecture of repeat-proteins is crucial in detailing how the local energetic balances can modulate the folding dynamics of these proteins, which can be related to the physiological behaviour of these ubiquitous biological systems. Introduction and between repeats, challenging the concept of structural It was early on noted that many natural proteins typically domain. collapse stretches of amino acid chains into compact units, defining structural domains [1]. These domains typically correlate with biological activities and many modern proteins can be described as composed by novel ‘domain arrange- Definition of the repeat-units ments’ [2]. -

A Structural Guide to Proteins of the NF-Kb Signaling Module
Downloaded from http://cshperspectives.cshlp.org/ on September 26, 2021 - Published by Cold Spring Harbor Laboratory Press A Structural Guide to Proteins of the NF-kB Signaling Module Tom Huxford1 and Gourisankar Ghosh2 1Department of Chemistry and Biochemistry, San Diego State University, 5500 Campanile Drive, San Diego, California 92182-1030 2Department of Chemistry and Biochemistry, University of California, San Diego, 9500 Gilman Drive, La Jolla, California 92093-0375 Correspondence: [email protected] The prosurvival transcription factor NF-kB specifically binds promoter DNA to activate target gene expression. NF-kB is regulated through interactions with IkB inhibitor proteins. Active proteolysis of these IkB proteins is, in turn, under the control of the IkB kinase complex (IKK). Together, these three molecules form the NF-kB signaling module. Studies aimed at charac- terizing the molecular mechanisms of NF-kB, IkB, and IKK in terms of their three-dimen- sional structures have lead to a greater understanding of this vital transcription factor system. F-kB is a master transcription factor that from the perspective of their three-dimensional Nresponds to diverse cell stimuli by activat- structures. ing the expression of stress response genes. Multiple signals, including cytokines, growth factors, engagement of the T-cell receptor, and NF-kB bacterial and viral products, induce NF-kB Introduction to NF-kB transcriptional activity (Hayden and Ghosh 2008). A point of convergence for the myriad NF-kB was discovered in the laboratory of of NF-kB inducing signals is the IkB kinase David Baltimore as a nuclear activity with bind- complex (IKK). Active IKK in turn controls ing specificity toward a ten-base-pair DNA transcription factor NF-kB by regulating pro- sequence 50-GGGACTTTCC-30 present within teolysis of the IkB inhibitor protein (Fig. -
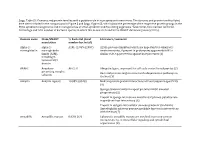
Supp. Table S2: Domains and Protein Families with a Putative Role in Host-Symbiont Interactions
Supp. Table S2: Domains and protein families with a putative role in host-symbiont interactions. The domains and protein families listed here were included in the comparisons in Figure 5 and Supp. Figure S5, which show the percentage of the respective protein groups in the Riftia symbiont metagenome and in metagenomes of other symbiotic and free-living organisms. % bacterial, total number bacterial: Percentage and total number of bacterial species in which this domain is found in the SMART database (January 2019). Domain name Pfam/SMART % bacterial (total Literature/comment annotation number bacterial) Alpha-2- alpha-2- A2M: 42.05% (2057) A2Ms: protease inhibitors which are important for eukaryotic macroglobulin macroglobulin innate immunity, if present in prokaryotes apparently fulfill a family (A2M), similar role, e.g. protection against host proteases (1) including N- terminal MG1 domain ANAPC Anaphase- APC2: 0 Ubiquitin ligase, important for cell cycle control in eukaryotes (2) promoting complex Bacterial proteins might interact with ubiquitination pathways in subunits the host (3) Ankyrin Ankyrin repeats 10.88% (8348) Mediate protein-protein interactions without sequence specificity (4) Sponge symbiont ankyrin-repeat proteins inhibit amoebal phagocytosis (5) Present in sponge microbiome metatranscriptomes, putative role in symbiont-host interactions (6) Present in obligate intracellular amoeba symbiont Candidatus Amoebophilus asiaticus genome, probable function in interactions with the host (7) Armadillo Armadillo repeats 0.83% (67) -

Appendix 2. Significantly Differentially Regulated Genes in Term Compared with Second Trimester Amniotic Fluid Supernatant
Appendix 2. Significantly Differentially Regulated Genes in Term Compared With Second Trimester Amniotic Fluid Supernatant Fold Change in term vs second trimester Amniotic Affymetrix Duplicate Fluid Probe ID probes Symbol Entrez Gene Name 1019.9 217059_at D MUC7 mucin 7, secreted 424.5 211735_x_at D SFTPC surfactant protein C 416.2 206835_at STATH statherin 363.4 214387_x_at D SFTPC surfactant protein C 295.5 205982_x_at D SFTPC surfactant protein C 288.7 1553454_at RPTN repetin solute carrier family 34 (sodium 251.3 204124_at SLC34A2 phosphate), member 2 238.9 206786_at HTN3 histatin 3 161.5 220191_at GKN1 gastrokine 1 152.7 223678_s_at D SFTPA2 surfactant protein A2 130.9 207430_s_at D MSMB microseminoprotein, beta- 99.0 214199_at SFTPD surfactant protein D major histocompatibility complex, class II, 96.5 210982_s_at D HLA-DRA DR alpha 96.5 221133_s_at D CLDN18 claudin 18 94.4 238222_at GKN2 gastrokine 2 93.7 1557961_s_at D LOC100127983 uncharacterized LOC100127983 93.1 229584_at LRRK2 leucine-rich repeat kinase 2 HOXD cluster antisense RNA 1 (non- 88.6 242042_s_at D HOXD-AS1 protein coding) 86.0 205569_at LAMP3 lysosomal-associated membrane protein 3 85.4 232698_at BPIFB2 BPI fold containing family B, member 2 84.4 205979_at SCGB2A1 secretoglobin, family 2A, member 1 84.3 230469_at RTKN2 rhotekin 2 82.2 204130_at HSD11B2 hydroxysteroid (11-beta) dehydrogenase 2 81.9 222242_s_at KLK5 kallikrein-related peptidase 5 77.0 237281_at AKAP14 A kinase (PRKA) anchor protein 14 76.7 1553602_at MUCL1 mucin-like 1 76.3 216359_at D MUC7 mucin 7, -

WSB1: from Homeostasis to Hypoxia Moinul Haque1,2,3, Joseph Keith Kendal1,2,3, Ryan Matthew Macisaac1,2,3 and Douglas James Demetrick1,2,3,4*
Haque et al. Journal of Biomedical Science (2016) 23:61 DOI 10.1186/s12929-016-0270-3 REVIEW Open Access WSB1: from homeostasis to hypoxia Moinul Haque1,2,3, Joseph Keith Kendal1,2,3, Ryan Matthew MacIsaac1,2,3 and Douglas James Demetrick1,2,3,4* Abstract The wsb1 gene has been identified to be important in developmental biology and cancer. A complex transcriptional regulation of wsb1 yields at least three functional transcripts. The major expressed isoform, WSB1 protein, is a substrate recognition protein within an E3 ubiquitin ligase, with the capability to bind diverse targets and mediate ubiquitinylation and proteolytic degradation. Recent data suggests a new role for WSB1 as a component of a neuroprotective pathway which results in modification and aggregation of neurotoxic proteins such as LRRK2 in Parkinson’s Disease, via an unusual mode of protein ubiquitinylation. WSB1 is also involved in thyroid hormone homeostasis, immune regulation and cellular metabolism, particularly glucose metabolism and hypoxia. In hypoxia, wsb1 is a HIF-1 target, and is a regulator of the degradation of diverse proteins associated with the cellular response to hypoxia, including HIPK2, RhoGDI2 and VHL. Major roles are to both protect HIF-1 function through degradation of VHL, and decrease apoptosis through degradation of HIPK2. These activities suggest a role for wsb1 in cancer cell proliferation and metastasis. As well, recent work has identified a role for WSB1 in glucose metabolism, and perhaps in mediating the Warburg effect in cancer cells by maintaining the function of HIF1. Furthermore, studies of cancer specimens have identified dysregulation of wsb1 associated with several types of cancer, suggesting a biologically relevant role in cancer development and/or progression. -

Folding-TIM Barrel
Protein Folding Practical September 2011 Folding up the TIM barrel Preliminary Examine the parallel beta barrel that you constructed, noting the stagger of the strands that was needed to connect the ends of the 8-stranded parallel beta sheet into the 8-stranded beta barrel. Notice that the stagger dictates which side of the sheet is on the inside and which is on the outside. This will be key information in folding the complete TIM linear peptide into the TIM barrel. Assembling the full linear peptide 1. Make sure the white beta strands are extended correctly, and the 8 yellow helices (with the green loops at each end) are correctly folded into an alpha helix (right handed with H-bonds to the 4th ahead in the chain). 2. starting with a beta strand connect an alpha helix and green loop to make the blue-red connecting peptide bond. Making sure that you connect the carbonyl (red) end of the beta strand to the amino (blue) end of the loop-helix-loop. Secure the just connected peptide bond bond with a twist-tie as shown. 3. complete step 2 for all beta strand/loop-helix-loop pairs, working in parallel with your partners 4. As pairs are completed attach the carboxy end of the strand- loop-helix-loop to the amino end of the next strand-loop-helix-loop module and secure the new peptide bond with a twist-tie as before. Repeat until the full linear TIM polypeptide chain is assembled. Make sure all strands and helices are still in the correct conformations. -

Supplemental Table 7. Every Significant Association
Supplemental Table 7. Every significant association between an individual covariate and functional group (assigned to the KO level) as determined by CPGLM regression analysis. Variable Unit RelationshipLabel See also CBCL Aggressive Behavior K05914 + CBCL Emotionally Reactive K05914 + CBCL Externalizing Behavior K05914 + K15665 K15658 CBCL Total K05914 + K15660 K16130 KO: E1.13.12.7; photinus-luciferin 4-monooxygenase (ATP-hydrolysing) [EC:1.13.12.7] :: PFAMS: AMP-binding enzyme; CBQ Inhibitory Control K05914 - K12239 K16120 Condensation domain; Methyltransferase domain; Thioesterase domain; AMP-binding enzyme C-terminal domain LEC Family Separation/Social Services K05914 + K16129 K16416 LEC Poverty Related Events K05914 + K16124 LEC Total K05914 + LEC Turmoil K05914 + CBCL Aggressive Behavior K15665 + CBCL Anxious Depressed K15665 + CBCL Emotionally Reactive K15665 + K05914 K15658 CBCL Externalizing Behavior K15665 + K15660 K16130 KO: K15665, ppsB, fenD; fengycin family lipopeptide synthetase B :: PFAMS: Condensation domain; AMP-binding enzyme; CBCL Total K15665 + K12239 K16120 Phosphopantetheine attachment site; AMP-binding enzyme C-terminal domain; Transferase family CBQ Inhibitory Control K15665 - K16129 K16416 LEC Poverty Related Events K15665 + K16124 LEC Total K15665 + LEC Turmoil K15665 + CBCL Aggressive Behavior K11903 + CBCL Anxiety Problems K11903 + CBCL Anxious Depressed K11903 + CBCL Depressive Problems K11903 + LEC Turmoil K11903 + MODS: Type VI secretion system K01220 K01058 CBCL Anxiety Problems K11906 + CBCL Depressive -

Tertiary Structure
Comments Structural motif v sequence motif polyproline (“PXXP”) motif for SH3 binding “RGD” motif for integrin binding “GXXXG” motif within the TM domain of membrane protein Most common type I’ beta turn sequences: X – (N/D/G)G – X Most common type II’ beta turn sequences: X – G(S/T) – X 1 Putting it together Alpha helices and beta sheets are not proteins—only marginally stable by themselves … Extremely small “proteins” can’t do much 2 Tertiary structure • Concerns with how the secondary structure units within a single polypeptide chain associate with each other to give a three- dimensional structure • Secondary structure, super secondary structure, and loops come together to form “domains”, the smallest tertiary structural unit • Structural domains (“domains”) usually contain 100 – 200 amino acids and fold stably. • Domains may be considered to be connected units which are to varying extents independent in terms of their structure, function and folding behavior. Each domain can be described by its fold, i.e. how the secondary structural elements are arranged. • Tertiary structure also includes the way domains fit together 3 Domains are modular •Because they are self-stabilizing, domains can be swapped both in nature and in the laboratory PI3 kinase beta-barrel GFP Branden & Tooze 4 fluorescence localization experiment Chimeras Recombinant proteins are often expressed and purified as fusion proteins (“chimeras”) with – glutathione S-transferase – maltose binding protein – or peptide tags, e.g. hexa-histidine, FLAG epitope helps with solubility, stability, and purification 5 Structural Classification All classifications are done at the domain level In many cases, structural similarity implies a common evolutionary origin – structural similarity without evolutionary relationship is possible – but no structural similarity means no evolutionary relationship Each domain has its corresponding “fold”, i.e. -

BMC Structural Biology Biomed Central
BMC Structural Biology BioMed Central Research article Open Access Natural history of S-adenosylmethionine-binding proteins Piotr Z Kozbial*1 and Arcady R Mushegian1,2 Address: 1Stowers Institute for Medical Research, 1000 E. 50th St., Kansas City, MO 64110, USA and 2Department of Microbiology, Molecular Genetics, and Immunology, University of Kansas Medical Center, Kansas City, Kansas 66160, USA Email: Piotr Z Kozbial* - [email protected]; Arcady R Mushegian - [email protected] * Corresponding author Published: 14 October 2005 Received: 21 July 2005 Accepted: 14 October 2005 BMC Structural Biology 2005, 5:19 doi:10.1186/1472-6807-5-19 This article is available from: http://www.biomedcentral.com/1472-6807/5/19 © 2005 Kozbial and Mushegian; licensee BioMed Central Ltd. This is an Open Access article distributed under the terms of the Creative Commons Attribution License (http://creativecommons.org/licenses/by/2.0), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited. Abstract Background: S-adenosylmethionine is a source of diverse chemical groups used in biosynthesis and modification of virtually every class of biomolecules. The most notable reaction requiring S- adenosylmethionine, transfer of methyl group, is performed by a large class of enzymes, S- adenosylmethionine-dependent methyltransferases, which have been the focus of considerable structure-function studies. Evolutionary trajectories of these enzymes, and especially of other classes of S-adenosylmethionine-binding proteins, nevertheless, remain poorly understood. We addressed this issue by computational comparison of sequences and structures of various S- adenosylmethionine-binding proteins. Results: Two widespread folds, Rossmann fold and TIM barrel, have been repeatedly used in evolution for diverse types of S-adenosylmethionine conversion. -

Molecular Modeling 2021 Lecture 3 -- Tues Feb 2
Molecular Modeling 2021 lecture 3 -- Tues Feb 2 Protein classification SCOP TOPS Contact maps domains Domains To a cell biologist a domain is a sequential unit within a gene, usually with a specific function. To a structural biologist a domain is a compact globular unit within a protein, classified by its 3D structure. 2 A domain is... • ... an autonomously-folding substructure of a protein. • ... > 30 residues, but typically < 200. May be bigger. • ...usually has a single hydrophobic core • ... usually composed of one chain (occasionally composed of multiple chains) • ...is usually composed on one contiguous segment (occasionally made of discontiguous segments of the same chain) 3 SARS-CoV-2 spike protein — a multi domain protein 4 SCOPe -- classification of domains !http://scop.berkeley.edu similar secondary structure (1) class content (2) fold vague structural homology (3) superfamily Clear structural homology (4) family increasing structural similarity structural increasing (5) protein Clear sequence homology (6) species nearly identical sequences individual structures SCOPe -- class 1. all α (289) classes of domains 2. all β (178) 3. α/β (148) 4. α+β (388) 5. multidomain (71) 6. membrane (60) 7. small (98) Not true classes of globular 8. coiled coil (7) protein domains 9. low-resolution (25) 10. peptides (148) 11. designed proteins (44) 12. artifacts (1) Proteins of the same class conserve secondary structure content SCOPe -- fold level within α/β proteins -- Mainly parallel beta sheets (beta-alpha-beta units) TIM-barrel (22) swivelling beta/beta/alpha domain (5) Many folds have historical spoIIaa-like (2) names. “TIM” barrel was flavodoxin-like (10) first seen in TIM. -

8-Barrel Enzymes John a Gerlt� and Frank M Raushely
252 Evolution of function in (b/a)8-barrel enzymes John A Gerltà and Frank M Raushely The (b/a)8-barrel is the most common fold in structurally closed, parallel b-sheet structure of the (b/a)8-barrel is characterized enzymes. Whether the functionally diverse formed from eight parallel (b/a)-units linked by hydrogen enzymes that share this fold are the products of either divergent bonds that form a cylindrical core. Despite its eightfold or convergent evolution (or both) is an unresolved question that pseudosymmetry, the packing within the interior of the will probably be answered as the sequence databases continue barrel is better described as four (b/a)2-subdomains in to expand. Recent work has examined natural, designed, and which distinct hydrophobic cores are located between the directed evolution of function in several superfamilies of (b/a)8- b-sheets and flanking a-helices [2]. barrel containing enzymes. The active sites are located at the C-terminal ends of the Addresses b-strands. So placed, the functional groups surround the ÃDepartments of Biochemistry and Chemistry, University of Illinois at active site and are structurally independent: the ‘old’ and Urbana-Champaign, 600 South Mathews Avenue, Urbana, ‘new’ enzymes retain functional groups at the ends of Illinois 61801, USA e-mail: [email protected] some b-strands, but others are altered to allow the ‘new’ yDepartment of Chemistry, Texas A&M University, PO Box 30012, activity [3]. With this blueprint, the (b/a)8-barrel is opti- College Station, Texas 77842-3012, USA mized for evolution of new functions.