Multiple Ways for Medical Data Visualization Using 3D Slicer
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Management of Large Sets of Image Data Capture, Databases, Image Processing, Storage, Visualization Karol Kozak
Management of large sets of image data Capture, Databases, Image Processing, Storage, Visualization Karol Kozak Download free books at Karol Kozak Management of large sets of image data Capture, Databases, Image Processing, Storage, Visualization Download free eBooks at bookboon.com 2 Management of large sets of image data: Capture, Databases, Image Processing, Storage, Visualization 1st edition © 2014 Karol Kozak & bookboon.com ISBN 978-87-403-0726-9 Download free eBooks at bookboon.com 3 Management of large sets of image data Contents Contents 1 Digital image 6 2 History of digital imaging 10 3 Amount of produced images – is it danger? 18 4 Digital image and privacy 20 5 Digital cameras 27 5.1 Methods of image capture 31 6 Image formats 33 7 Image Metadata – data about data 39 8 Interactive visualization (IV) 44 9 Basic of image processing 49 Download free eBooks at bookboon.com 4 Click on the ad to read more Management of large sets of image data Contents 10 Image Processing software 62 11 Image management and image databases 79 12 Operating system (os) and images 97 13 Graphics processing unit (GPU) 100 14 Storage and archive 101 15 Images in different disciplines 109 15.1 Microscopy 109 360° 15.2 Medical imaging 114 15.3 Astronomical images 117 15.4 Industrial imaging 360° 118 thinking. 16 Selection of best digital images 120 References: thinking. 124 360° thinking . 360° thinking. Discover the truth at www.deloitte.ca/careers Discover the truth at www.deloitte.ca/careers © Deloitte & Touche LLP and affiliated entities. Discover the truth at www.deloitte.ca/careers © Deloitte & Touche LLP and affiliated entities. -

A 3D Interactive Multi-Object Segmentation Tool Using Local Robust Statistics Driven Active Contours
A 3D interactive multi-object segmentation tool using local robust statistics driven active contours The Harvard community has made this article openly available. Please share how this access benefits you. Your story matters Citation Gao, Yi, Ron Kikinis, Sylvain Bouix, Martha Shenton, and Allen Tannenbaum. 2012. A 3D Interactive Multi-Object Segmentation Tool Using Local Robust Statistics Driven Active Contours. Medical Image Analysis 16, no. 6: 1216–1227. doi:10.1016/j.media.2012.06.002. Published Version doi:10.1016/j.media.2012.06.002 Citable link http://nrs.harvard.edu/urn-3:HUL.InstRepos:28548930 Terms of Use This article was downloaded from Harvard University’s DASH repository, and is made available under the terms and conditions applicable to Other Posted Material, as set forth at http:// nrs.harvard.edu/urn-3:HUL.InstRepos:dash.current.terms-of- use#LAA NIH Public Access Author Manuscript Med Image Anal. Author manuscript; available in PMC 2013 August 01. NIH-PA Author ManuscriptPublished NIH-PA Author Manuscript in final edited NIH-PA Author Manuscript form as: Med Image Anal. 2012 August ; 16(6): 1216–1227. doi:10.1016/j.media.2012.06.002. A 3D Interactive Multi-object Segmentation Tool using Local Robust Statistics Driven Active Contours Yi Gaoa,*, Ron Kikinisb, Sylvain Bouixa, Martha Shentona, and Allen Tannenbaumc aPsychiatry Neuroimaging Laboratory, Brigham & Women's Hospital, Harvard Medical School, Boston, MA 02115 bSurgical Planning Laboratory, Brigham & Women's Hospital, Harvard Medical School, Boston, MA 02115 cDepartments of Electrical and Computer Engineering and Biomedical Engineering, Boston University, Boston, MA 02115 Abstract Extracting anatomical and functional significant structures renders one of the important tasks for both the theoretical study of the medical image analysis, and the clinical and practical community. -
![Downloaded from the Cellprofiler Site [31] to Provide a Starting Point for New Analyses](https://docslib.b-cdn.net/cover/6758/downloaded-from-the-cellprofiler-site-31-to-provide-a-starting-point-for-new-analyses-626758.webp)
Downloaded from the Cellprofiler Site [31] to Provide a Starting Point for New Analyses
Open Access Software2006CarpenteretVolume al. 7, Issue 10, Article R100 CellProfiler: image analysis software for identifying and quantifying comment cell phenotypes Anne E Carpenter*, Thouis R Jones*†, Michael R Lamprecht*, Colin Clarke*†, In Han Kang†, Ola Friman‡, David A Guertin*, Joo Han Chang*, Robert A Lindquist*, Jason Moffat*, Polina Golland† and David M Sabatini*§ reviews Addresses: *Whitehead Institute for Biomedical Research, Cambridge, MA 02142, USA. †Computer Sciences and Artificial Intelligence Laboratory, Massachusetts Institute of Technology, Cambridge, MA 02142, USA. ‡Department of Radiology, Brigham and Women's Hospital, Boston, MA 02115, USA. §Department of Biology, Massachusetts Institute of Technology, Cambridge, MA 02142, USA. Correspondence: David M Sabatini. Email: [email protected] Published: 31 October 2006 Received: 15 September 2006 Accepted: 31 October 2006 reports Genome Biology 2006, 7:R100 (doi:10.1186/gb-2006-7-10-r100) The electronic version of this article is the complete one and can be found online at http://genomebiology.com/2006/7/10/R100 © 2006 Carpenter et al.; licensee BioMed Central Ltd. This is an open access article distributed under the terms of the Creative Commons Attribution License (http://creativecommons.org/licenses/by/2.0), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited. deposited research Cell<p>CellProfiler, image analysis the software first free, open-source system for flexible and high-throughput cell image analysis is described.</p> Abstract Biologists can now prepare and image thousands of samples per day using automation, enabling chemical screens and functional genomics (for example, using RNA interference). Here we describe the first free, open-source system designed for flexible, high-throughput cell image analysis, research refereed CellProfiler. -

Survey of Databases Used in Image Processing and Their Applications
International Journal of Scientific & Engineering Research Volume 2, Issue 10, Oct-2011 1 ISSN 2229-5518 Survey of Databases Used in Image Processing and Their Applications Shubhpreet Kaur, Gagandeep Jindal Abstract- This paper gives review of Medical image database (MIDB) systems which have been developed in the past few years for research for medical fraternity and students. In this paper, I have surveyed all available medical image databases relevant for research and their use. Keywords: Image database, Medical Image Database System. —————————— —————————— 1. INTRODUCTION Measurement and recording techniques, such as electroencephalography, magnetoencephalography Medical imaging is the technique and process used to (MEG), Electrocardiography (EKG) and others, can create images of the human for clinical purposes be seen as forms of medical imaging. Image Analysis (medical procedures seeking to reveal, diagnose or is done to ensure database consistency and reliable examine disease) or medical science. As a discipline, image processing. it is part of biological imaging and incorporates radiology, nuclear medicine, investigative Open source software for medical image analysis radiological sciences, endoscopy, (medical) Several open source software packages are available thermography, medical photography and for performing analysis of medical images: microscopy. ImageJ 3D Slicer ITK Shubhpreet Kaur is currently pursuing masters degree OsiriX program in Computer Science and engineering in GemIdent Chandigarh Engineering College, Mohali, India. E-mail: MicroDicom [email protected] FreeSurfer Gagandeep Jindal is currently assistant processor in 1.1 Images used in Medical Research department Computer Science and Engineering in Here is the description of various modalities that are Chandigarh Engineering College, Mohali, India. E-mail: used for the purpose of research by medical and [email protected] engineering students as well as doctors. -

An Open-Source Research Platform for Image-Guided Therapy
Int J CARS DOI 10.1007/s11548-015-1292-0 ORIGINAL ARTICLE CustusX: an open-source research platform for image-guided therapy Christian Askeland1,3 · Ole Vegard Solberg1 · Janne Beate Lervik Bakeng1 · Ingerid Reinertsen1 · Geir Arne Tangen1 · Erlend Fagertun Hofstad1 · Daniel Høyer Iversen1,2,3 · Cecilie Våpenstad1,2 · Tormod Selbekk1,3 · Thomas Langø1,3 · Toril A. Nagelhus Hernes2,3 · Håkon Olav Leira2,3 · Geirmund Unsgård2,3 · Frank Lindseth1,2,3 Received: 3 July 2015 / Accepted: 31 August 2015 © The Author(s) 2015. This article is published with open access at Springerlink.com Abstract Results The validation experiments show a navigation sys- Purpose CustusX is an image-guided therapy (IGT) research tem accuracy of <1.1mm, a frame rate of 20 fps, and latency platform dedicated to intraoperative navigation and ultra- of 285ms for a typical setup. The current platform is exten- sound imaging. In this paper, we present CustusX as a robust, sible, user-friendly and has a streamlined architecture and accurate, and extensible platform with full access to data and quality process. CustusX has successfully been used for algorithms and show examples of application in technologi- IGT research in neurosurgery, laparoscopic surgery, vascular cal and clinical IGT research. surgery, and bronchoscopy. Methods CustusX has been developed continuously for Conclusions CustusX is now a mature research platform more than 15years based on requirements from clinical for intraoperative navigation and ultrasound imaging and is and technological researchers within the framework of a ready for use by the IGT research community. CustusX is well-defined software quality process. The platform was open-source and freely available at http://www.custusx.org. -

Medical Image Processing Software
Wohlers Report 2018 Medical Image Processing Software Medical image Patient-specific medical devices and anatomical models are almost always produced using radiological imaging data. Medical image processing processing software is used to translate between radiology file formats and various software AM file formats. Theoretically, any volumetric radiological imaging dataset by Andy Christensen could be used to create these devices and models. However, without high- and Nicole Wake quality medical image data, the output from AM can be less than ideal. In this field, the old adage of “garbage in, garbage out” definitely applies. Due to the relative ease of image post-processing, computed tomography (CT) is the usual method for imaging bone structures and contrast- enhanced vasculature. In the dental field and for oral- and maxillofacial surgery, in-office cone-beam computed tomography (CBCT) has become popular. Another popular imaging technique that can be used to create anatomical models is magnetic resonance imaging (MRI). MRI is less useful for bone imaging, but its excellent soft tissue contrast makes it useful for soft tissue structures, solid organs, and cancerous lesions. Computed tomography: CT uses many X-ray projections through a subject to computationally reconstruct a cross-sectional image. As with traditional 2D X-ray imaging, a narrow X-ray beam is directed to pass through the subject and project onto an opposing detector. To create a cross-sectional image, the X-ray source and detector rotate around a stationary subject and acquire images at a number of angles. An image of the cross-section is then computed from these projections in a post-processing step. -

Imagej Basics (Version 1.38)
ImageJ Basics (Version 1.38) ImageJ is a powerful image analysis program that was created at the National Institutes of Health. It is in the public domain, runs on a variety of operating systems and is updated frequently. You may download this program from the source (http://rsb.info.nih.gov/ij/) or copy the ImageJ folder from the C drive of your lab computer. The ImageJ website has instructions for use of the program and links to useful resources. Installing ImageJ on your PC (Windows operating system): Copy the ImageJ folder and transfer it to the C drive of your personal computer. Open the ImageJ folder in the C drive and copy the shortcut (microscope with arrow) to your computer’s desktop. Double click on this desktop shortcut to run ImageJ. See the ImageJ website for Macintosh instructions. ImageJ Window: The ImageJ window will appear on the desktop; do not enlarge this window. Note that this window has a Menu Bar, a Tool Bar and a Status Bar. Menu Bar → Tool Bar → Status Bar → Graphics are from the ImageJ website (http://rsb.info.nih.gov/ij/). Adjusting Memory Allocation: Use the Edit → Options → Memory command to adjust the default memory allocation. Setting the maximum memory value to more than about 75% of real RAM may result in poor performance due to virtual memory "thrashing". Opening an Image File: Select File → Open from the menu bar to open a stored image file. Tool Bar: The various buttons on the tool bar allow you measure, draw, label, fill, etc. A right- click or a double left-click may expand your options with some of the tool buttons. -

Paleoanthropology Society Meeting Abstracts, St. Louis, Mo, 13-14 April 2010
PALEOANTHROPOLOGY SOCIETY MEETING ABSTRACTS, ST. LOUIS, MO, 13-14 APRIL 2010 New Data on the Transition from the Gravettian to the Solutrean in Portuguese Estremadura Francisco Almeida , DIED DEPA, Igespar, IP, PORTUGAL Henrique Matias, Department of Geology, Faculdade de Ciências da Universidade de Lisboa, PORTUGAL Rui Carvalho, Department of Geology, Faculdade de Ciências da Universidade de Lisboa, PORTUGAL Telmo Pereira, FCHS - Departamento de História, Arqueologia e Património, Universidade do Algarve, PORTUGAL Adelaide Pinto, Crivarque. Lda., PORTUGAL From an anthropological perspective, the passage from the Gravettian to the Solutrean is one of the most interesting transition peri- ods in Old World Prehistory. Between 22 kyr BP and 21 kyr BP, during the beginning stages of the Last Glacial Maximum, Iberia and Southwest France witness a process of substitution of a Pan-European Technocomplex—the Gravettian—to one of the first examples of regionalism by Anatomically Modern Humans in the European continent—the Solutrean. While the question of the origins of the Solutrean is almost as old as its first definition, the process under which it substituted the Gravettian started to be readdressed, both in Portugal and in France, after the mid 1990’s. Two chronological models for the transition have been advanced, but until very recently the lack of new archaeological contexts of the period, and the fact that the many of the sequences have been drastically affected by post depositional disturbances during the Lascaux event, prevented their systematic evaluation. Between 2007 and 2009, and in the scope of mitigation projects, archaeological fieldwork has been carried in three open air sites—Terra do Manuel (Rio Maior), Portela 2 (Leiria), and Calvaria 2 (Porto de Mós) whose stratigraphic sequences date precisely to the beginning stages of the LGM. -

Anastasia Tyurina [email protected]
1 Anastasia Tyurina [email protected] Summary A specialist in applying or creating mathematical methods to solving problems of developing technologies. A rare expert in solving problems starting from the stage of a stated “word problem” to proof of concept and production software development. Such successful uses of an educational background in mathematics, intellectual courage, and tenacious character include: • developed a unique method of statistical analysis of spectral composition in 1D and 2D stochastic processes for quality control in ultra-precision mirror polishing • developed novel methods of detection, tracking and classification of small moving targets for aerial IR and EO sensors. Used SIFT, and SIRF features, and developed innovative feature-signatures of motion of interest. • developed image processing software for bioinformatics, point source (diffraction objects) detection semiconductor metrology, electron microscopy, failure analysis, diagnostics, system hardware support and pattern recognition • developed statistical software for surface metrology assessment, characterization and generation of statistically similar surfaces to assist development of new optical systems • documented, published and patented original results helping employers technical communications • supported sales with prototypes and presentations • worked well with people – colleagues, customers, researchers, scientists, engineers Tools MATLAB, Octave, OpenCV, ImageJ, Scion Image, Aphelion Image, Gimp, PhotoShop, C/C++, (Visual C environment), GNU development tools, UNIX (Solaris, SGI IRIX), Linux, Windows, MS DOS. Positions and Experience Second Star Algonumerixs – 2008-present, founder and CEO http://www.secondstaralgonumerix.com/ 1) Developed a method of statistical assessment, characterisation and generation of random surface metrology for sper precision X-ray mirror manufacturing in collaboration with Lawrence Berkeley National Laboratory of University of California Berkeley. -

Medical Images Research Framework
Medical Images Research Framework Sabrina Musatian Alexander Lomakin Angelina Chizhova Saint Petersburg State University Saint Petersburg State University Saint Petersburg State University Saint Petersburg, Russia Saint Petersburg, Russia Saint Petersburg, Russia Email: [email protected] Email: [email protected] Email: [email protected] Abstract—with a growing interest in medical research problems for the development of medical instruments and to show and the introduction of machine learning methods for solving successful applications of this library on some real medical those, a need in an environment for integrating modern solu- cases. tions and algorithms into medical applications developed. The main goal of our research is to create medical images research 2. Existing systems for medical image process- framework (MIRF) as a solution for the above problem. MIRF ing is a free open–source platform for the development of medical tools with image processing. We created it to fill in the gap be- There are many open–source packages and software tween innovative research with medical images and integrating systems for working with medical images. Some of them are it into real–world patients treatments workflow. Within a short specifically dedicated for these purposes, others are adapted time, a developer can create a rich medical tool, using MIRF's to be used for medical procedures. modular architecture and a set of included features. MIRF Many of them comprise a set of instruments, dedicated takes the responsibility of handling common functionality for to solving typical tasks, such as images pre–processing medical images processing. The only thing required from the and analysis of the results – ITK [1], visualization – developer is integrating his functionality into a module and VTK [2], real–time pre–processing of images and video – choosing which of the other MIRF's features are needed in the OpenCV [3]. -

An Open Source Freeware Software for Ultrasound Imaging and Elastography
Proceedings of the eNTERFACE’07 Workshop on Multimodal Interfaces, Istanbul,˙ Turkey, July 16 - August 10, 2007 USIMAGTOOL: AN OPEN SOURCE FREEWARE SOFTWARE FOR ULTRASOUND IMAGING AND ELASTOGRAPHY Ruben´ Cardenes-Almeida´ 1, Antonio Tristan-Vega´ 1, Gonzalo Vegas-Sanchez-Ferrero´ 1, Santiago Aja-Fernandez´ 1, Veronica´ Garc´ıa-Perez´ 1, Emma Munoz-Moreno˜ 1, Rodrigo de Luis-Garc´ıa 1, Javier Gonzalez-Fern´ andez´ 2, Dar´ıo Sosa-Cabrera 2, Karl Krissian 2, Suzanne Kieffer 3 1 LPI, University of Valladolid, Spain 2 CTM, University of Las Palmas de Gran Canaria 3 TELE Laboratory, Universite´ catholique de Louvain, Louvain-la-Neuve, Belgium ABSTRACT • Open source code: to be able for everyone to modify and reuse the source code. UsimagTool will prepare specific software for the physician to change parameters for filtering and visualization in Ultrasound • Efficiency, robust and fast: using a standard object ori- Medical Imaging in general and in Elastography in particular, ented language such as C++. being the first software tool for researchers and physicians to • Modularity and flexibility for developers: in order to chan- compute elastography with integrated algorithms and modular ge or add functionalities as fast as possible. coding capabilities. It will be ready to implement in different • Multi-platform: able to run in many Operating systems ecographic systems. UsimagTool is based on C++, and VTK/ITK to be useful for more people. functions through a hidden layer, which means that participants may import their own functions and/or use the VTK/ITK func- • Usability: provided with an easy to use GUI to interact tions. as easy as possible with the end user. -
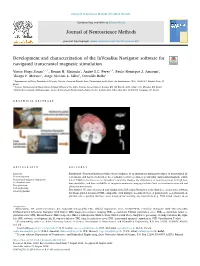
Development and Characterization of the Invesalius Navigator Software for Navigated Transcranial Magnetic Stimulation T ⁎ Victor Hugo Souzaa, ,1, Renan H
Journal of Neuroscience Methods 309 (2018) 109–120 Contents lists available at ScienceDirect Journal of Neuroscience Methods journal homepage: www.elsevier.com/locate/jneumeth Development and characterization of the InVesalius Navigator software for navigated transcranial magnetic stimulation T ⁎ Victor Hugo Souzaa, ,1, Renan H. Matsudaa, André S.C. Peresa,b, Paulo Henrique J. Amorimc, Thiago F. Moraesc, Jorge Vicente L. Silvac, Oswaldo Baffaa a Departamento de Física, Faculdade de Filosofia, Ciências e Letras de Ribeirão Preto, Universidade de São Paulo, Av. Bandeirantes, 3900, 14040-901, Ribeirão Preto, SP, Brazil b Instituto Internacional de Neurociência de Natal Edmond e Lily Safra, Instituto Santos Dumont, Rodovia RN 160 Km 03, 3003, 59280-000, Macaíba, RN, Brazil c Núcleo de Tecnologias Tridimensionais, Centro de Tecnologia da Informação Renato Archer, Rodovia Dom Pedro I Km 143, 13069-901, Campinas, SP, Brazil GRAPHICAL ABSTRACT ARTICLE INFO ABSTRACT Keywords: Background: Neuronavigation provides visual guidance of an instrument during procedures of neurological in- Neuronavigation terventions, and has been shown to be a valuable tool for accurately positioning transcranial magnetic stimu- Transcranial magnetic stimulation lation (TMS) coils relative to an individual’s anatomy. Despite the importance of neuronavigation, its high cost, Localization error low portability, and low availability of magnetic resonance imaging facilities limit its insertion in research and Co-registration clinical environments. Coil positioning New method: We have developed and validated the InVesalius Navigator as the first free, open-source software Surgical planning for image-guided navigated TMS, compatible with multiple tracking devices. A point-based, co-registration al- gorithm and a guiding interface were designed for tracking any instrument (e.g.