Identification of Tumor Mutation Burden-Related Hub Genes and the Underlying Mechanism in Melanoma
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

ZMYND10 Functions in a Chaperone Relay During Axonemal Dynein Assembly
bioRxiv preprint doi: https://doi.org/10.1101/233718; this version posted December 13, 2017. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. 1 ZMYND10 functions in a chaperone relay during axonemal dynein assembly. 2 3 Girish R Mali1,9 , Patricia Yeyati1, Seiya Mizuno2, Margaret A Keighren1, Petra zur Lage3, Amaya 4 Garcia-Munoz4, Atsuko Shimada5, Hiroyuki Takeda5, Frank Edlich6, Satoru Takahashi2,7, Alex von 5 Kreigsheim4,8, Andrew Jarman3 and Pleasantine Mill1,*. 6 7 1. MRC Human Genetics Unit, Institute of Genetics and Molecular Medicine, University of 8 Edinburgh, Edinburgh, UK, EH4 2XU 9 2. Laboratory Animal Resource Centre, University of Tsukuba, Tsukuba, Japan, 305-8575 10 3. Centre for Integrative Physiology, University of Edinburgh, Edinburgh, UK, EH8 9XD 11 4. Systems Biology Ireland, University College Dublin, Dublin, Ireland 12 5. Department of Biological Sciences, University of Tokyo, Tokyo, Japan, 113-0033 13 6. Institute for Biochemistry and Molecular Biology, University of Freiburg, Freiburg, Germany, 14 79104 15 7. Department of Anatomy and Embryology, Faculty of Medicine, University of Tsukuba, Tsukuba, 16 Japan, 305-8575 17 8. Edinburgh Cancer Research UK Centre, Institute of Genetics and Molecular Medicine, University 18 of Edinburgh, Edinburgh, UK, EH4 2XU 19 9. Current address: MRC Laboratory of Molecular Biology, Cambridge, UK, CB2 0QH 20 * Corresponding author: [email protected] 21 22 23 24 25 26 27 28 29 30 31 1 bioRxiv preprint doi: https://doi.org/10.1101/233718; this version posted December 13, 2017. -

Supplementary Materials Functional Characterization of Rare RAB12
1 Supplementary materials Functional characterization of rare RAB12 variants and their role in musician’s and other dystonias Eva Hebert et al. Figure S1. Photograph of Individual L-10289 (mildly affected mother of the index patient from Family D) showing a 15-degree tilt of the trunk to the right as well as dystonic posturing of the right hand (involuntary flexion of the third to fifth finger and thumb and extension of the index finger). 2 Figure S2. TFRC colocalized with wildtype and mutant FLAG-RAB12. Immunofluorescent staining of fibroblasts expressing FLAG-RAB12 WT, p.Gly13Asp, or p.Ile196Val revealed predominant perinuclear localization of TFRC (red) which overlaps with the localization of FLAG-RAB12 (green) in all three cell lines (WT, p.Gly13Asp, p.Ile196Val). The nucleus was stained with DAPI (blue). Scale bar: 20µm. 3 Figure S3. Lysosomal degradation of the physiological dimeric TFRC was not affected by the RAB12 mutations. Western Blot analysis revealed the degradation of TFRC in patient fibroblasts with endogenous expression of RAB12 (a, b) in fibroblasts ectopically expressing FLAG-RAB12 (c, d), and in SH-SY5Y cells ectopically expressing FLAG-RAB12 (e, f). Cells were treated with Bafilomycin A1 for 24h. ß-actin served as loading control and for normalization. Bars in B, D, and F indicate means of three independent experiments ± SEM. ctrl control, WT wildtype 4 Figure S4. Relative LC3-II protein levels are marginally increased in SH-SY5Y cells overexpressing RAB12 Gly13Asp protein and p62 levels remained constant. a) Western Blot of proteins extracted from stably transfected SH-SY5Y cells. Expression of FLAG-tagged RAB12 WT equals the expression of mutated RAB12 proteins (Gly13Asp, I196Val) (lane 3, 5, 7). -

RESEARCH ARTICLE Body Weight-Dependent Troponin T Alternative Splicing Is Evolutionarily Conserved from Insects to Mammals and I
1523 The Journal of Experimental Biology 214, 1523-1532 © 2011. Published by The Company of Biologists Ltd doi:10.1242/jeb.051763 RESEARCH ARTICLE Body weight-dependent troponin T alternative splicing is evolutionarily conserved from insects to mammals and is partially impaired in skeletal muscle of obese rats Rudolf J. Schilder1,*, Scot R. Kimball1, James H. Marden2 and Leonard S. Jefferson1 1Department of Cellular and Molecular Physiology, The Pennsylvania State University College of Medicine, 500 University Drive, Hershey, PA 17033, USA and 2Department of Biology, The Pennsylvania State University, 208 Mueller Lab, University Park, PA 16802, USA *Author for correspondence ([email protected]) Accepted 19 January 2011 SUMMARY Do animals know at a physiological level how much they weigh, and, if so, do they make homeostatic adjustments in response to changes in body weight? Skeletal muscle is a likely tissue for such plasticity, as weight-bearing muscles receive mechanical feedback regarding body weight and consume ATP in order to generate forces sufficient to counteract gravity. Using rats, we examined how variation in body weight affected alternative splicing of fast skeletal muscle troponin T (Tnnt3), a component of the thin filament that regulates the actin–myosin interaction during contraction and modulates force output. In response to normal growth and experimental body weight increases, alternative splicing of Tnnt3 in rat gastrocnemius muscle was adjusted in a quantitative fashion. The response depended on weight per se, as externally attached loads had the same effect as an equal change in actual body weight. Examining the association between Tnnt3 alternative splicing and ATP consumption rate, we found that the Tnnt3 splice form profile had a significant association with nocturnal energy expenditure, independently of effects of weight. -

TNNT3 Rabbit Pab
Leader in Biomolecular Solutions for Life Science TNNT3 Rabbit pAb Catalog No.: A15323 Basic Information Background Catalog No. The binding of Ca(2+) to the trimeric troponin complex initiates the process of muscle A15323 contraction. Increased Ca(2+) concentrations produce a conformational change in the troponin complex that is transmitted to tropomyosin dimers situated along actin Observed MW filaments. The altered conformation permits increased interaction between a myosin 37kDa head and an actin filament which, ultimately, produces a muscle contraction. The troponin complex has protein subunits C, I, and T. Subunit C binds Ca(2+) and subunit I Calculated MW binds to actin and inhibits actin-myosin interaction. Subunit T binds the troponin 29kDa/30kDa/31kDa complex to the tropomyosin complex and is also required for Ca(2+)-mediated activation of actomyosin ATPase activity. There are 3 different troponin T genes that Category encode tissue-specific isoforms of subunit T for fast skeletal-, slow skeletal-, and cardiac-muscle. This gene encodes fast skeletal troponin T protein; also known as Primary antibody troponin T type 3. Alternative splicing results in multiple transcript variants encoding additional distinct troponin T type 3 isoforms. A developmentally regulated switch Applications between fetal/neonatal and adult troponin T type 3 isoforms occurs. Additional splice WB variants have been described but their biological validity has not been established. Mutations in this gene may cause distal arthrogryposis multiplex congenita type 2B Cross-Reactivity (DA2B). Mouse, Rat Recommended Dilutions Immunogen Information WB 1:200 - 1:2000 Gene ID Swiss Prot 7140 P45378 Immunogen Recombinant fusion protein containing a sequence corresponding to amino acids 147-256 of human TNNT3 (NP_001036246.1). -

Establishment of the Early Cilia Preassembly Protein Complex
Establishment of the early cilia preassembly protein PNAS PLUS complex during motile ciliogenesis Amjad Horania,1, Alessandro Ustioneb, Tao Huangc, Amy L. Firthd, Jiehong Panc, Sean P. Gunstenc, Jeffrey A. Haspelc, David W. Pistonb, and Steven L. Brodyc aDepartment of Pediatrics, Washington University School of Medicine, St. Louis, MO 63110; bDepartment of Cell Biology and Physiology, Washington University School of Medicine, St. Louis, MO 63110; cDepartment of Medicine, Washington University School of Medicine, St. Louis, MO 63110; and dDepartment of Medicine, University of Southern California, Keck School of Medicine, Los Angeles, CA 90033 Edited by Kathryn V. Anderson, Sloan Kettering Institute, New York, NY, and approved December 27, 2017 (received for review September 9, 2017) Motile cilia are characterized by dynein motor units, which preas- function of these proteins is unknown; however, missing dynein semble in the cytoplasm before trafficking into the cilia. Proteins motor complexes in the cilia of mutants and cytoplasmic locali- required for dynein preassembly were discovered by finding human zation (or absence in the cilia proteome) suggest a role in the mutations that result in absent ciliary motors, but little is known preassembly of dynein motor complexes. Studies in C. reinhardtii about their expression, function, or interactions. By monitoring show motor components in the cell body before transport to ciliogenesis in primary airway epithelial cells and MCIDAS-regulated flagella (22–25). However, the expression, interactions, and induced pluripotent stem cells, we uncovered two phases of expres- functions of preassembly proteins, as well as the steps required sion of preassembly proteins. An early phase, composed of HEATR2, for preassembly, are undefined. -

De Novo, Systemic, Deleterious Amino Acid Substitutions Are Common in Large Cytoskeleton‑Related Protein Coding Regions
BIOMEDICAL REPORTS 6: 211-216, 2017 De novo, systemic, deleterious amino acid substitutions are common in large cytoskeleton‑related protein coding regions REBECCA J. STOLL1, GRACE R. THOMPSON1, MOHAMMAD D. SAMY1 and GEORGE BLANCK1,2 1Department of Molecular Medicine, Morsani College of Medicine, University of South Florida; 2Immunology Program, H. Lee Moffitt Cancer Center and Research Institute, Tampa, FL 33612, USA Received June 13, 2016; Accepted October 31, 2016 DOI: 10.3892/br.2016.826 Abstract. Human mutagenesis is largely random, thus large Introduction coding regions, simply on the basis of probability, represent relatively large mutagenesis targets. Thus, we considered Genetic damage is largely random and therefore tends to the possibility that large cytoskeletal-protein related coding affect the larger, functional regions of the human genome regions (CPCRs), including extra-cellular matrix (ECM) more frequently than the smaller regions (1). For example, coding regions, would have systemic nucleotide variants that a systematic study has revealed that cancer fusion genes, on are not present in common SNP databases. Presumably, such average, are statistically, significantly larger than other human variants arose recently in development or in recent, preceding genes (2,3). The large introns of potential cancer fusion genes generations. Using matched breast cancer and blood-derived presumably allow for many different productive recombina- normal datasets from the cancer genome atlas, CPCR single tion opportunities, i.e., many recombinations that would allow nucleotide variants (SNVs) not present in the All SNPs(142) for exon juxtaposition and the generation of hybrid proteins. or 1000 Genomes databases were identified. Using the Protein Smaller cancer fusion genes tend to be associated with the rare Variation Effect Analyzer internet-based tool, it was discov- types of cancer, for example EWS RNA binding protein 1 in ered that apparent, systemic mutations (not shared among Ewing's sarcoma. -
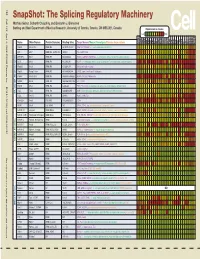
Snapshot: the Splicing Regulatory Machinery Mathieu Gabut, Sidharth Chaudhry, and Benjamin J
192 Cell SnapShot: The Splicing Regulatory Machinery Mathieu Gabut, Sidharth Chaudhry, and Benjamin J. Blencowe 133 Banting and Best Department of Medical Research, University of Toronto, Toronto, ON M5S 3E1, Canada Expression in mouse , April4, 2008©2008Elsevier Inc. Low High Name Other Names Protein Domains Binding Sites Target Genes/Mouse Phenotypes/Disease Associations Amy Ceb Hip Hyp OB Eye SC BM Bo Ht SM Epd Kd Liv Lu Pan Pla Pro Sto Spl Thy Thd Te Ut Ov E6.5 E8.5 E10.5 SRp20 Sfrs3, X16 RRM, RS GCUCCUCUUC SRp20, CT/CGRP; −/− early embryonic lethal E3.5 9G8 Sfrs7 RRM, RS, C2HC Znf (GAC)n Tau, GnRH, 9G8 ASF/SF2 Sfrs1 RRM, RS RGAAGAAC HipK3, CaMKIIδ, HIV RNAs; −/− embryonic lethal, cond. KO cardiomyopathy SC35 Sfrs2 RRM, RS UGCUGUU AChE; −/− embryonic lethal, cond. KO deficient T-cell maturation, cardiomyopathy; LS SRp30c Sfrs9 RRM, RS CUGGAUU Glucocorticoid receptor SRp38 Fusip1, Nssr RRM, RS ACAAAGACAA CREB, type II and type XI collagens SRp40 Sfrs5, HRS RRM, RS AGGAGAAGGGA HipK3, PKCβ-II, Fibronectin SRp55 Sfrs6 RRM, RS GGCAGCACCUG cTnT, CD44 DOI 10.1016/j.cell.2008.03.010 SRp75 Sfrs4 RRM, RS GAAGGA FN1, E1A, CD45; overexpression enhances chondrogenic differentiation Tra2α Tra2a RRM, RS GAAARGARR GnRH; overexpression promotes RA-induced neural differentiation SR and SR-Related Proteins Tra2β Sfrs10 RRM, RS (GAA)n HipK3, SMN, Tau SRm160 Srrm1 RS, PWI AUGAAGAGGA CD44 SWAP Sfrs8 RS, SWAP ND SWAP, CD45, Tau; possible asthma susceptibility gene hnRNP A1 Hnrnpa1 RRM, RGG UAGGGA/U HipK3, SMN2, c-H-ras; rheumatoid arthritis, systemic lupus -

NICU Gene List Generator.Xlsx
Neonatal Crisis Sequencing Panel Gene List Genes: A2ML1 - B3GLCT A2ML1 ADAMTS9 ALG1 ARHGEF15 AAAS ADAMTSL2 ALG11 ARHGEF9 AARS1 ADAR ALG12 ARID1A AARS2 ADARB1 ALG13 ARID1B ABAT ADCY6 ALG14 ARID2 ABCA12 ADD3 ALG2 ARL13B ABCA3 ADGRG1 ALG3 ARL6 ABCA4 ADGRV1 ALG6 ARMC9 ABCB11 ADK ALG8 ARPC1B ABCB4 ADNP ALG9 ARSA ABCC6 ADPRS ALK ARSL ABCC8 ADSL ALMS1 ARX ABCC9 AEBP1 ALOX12B ASAH1 ABCD1 AFF3 ALOXE3 ASCC1 ABCD3 AFF4 ALPK3 ASH1L ABCD4 AFG3L2 ALPL ASL ABHD5 AGA ALS2 ASNS ACAD8 AGK ALX3 ASPA ACAD9 AGL ALX4 ASPM ACADM AGPS AMELX ASS1 ACADS AGRN AMER1 ASXL1 ACADSB AGT AMH ASXL3 ACADVL AGTPBP1 AMHR2 ATAD1 ACAN AGTR1 AMN ATL1 ACAT1 AGXT AMPD2 ATM ACE AHCY AMT ATP1A1 ACO2 AHDC1 ANK1 ATP1A2 ACOX1 AHI1 ANK2 ATP1A3 ACP5 AIFM1 ANKH ATP2A1 ACSF3 AIMP1 ANKLE2 ATP5F1A ACTA1 AIMP2 ANKRD11 ATP5F1D ACTA2 AIRE ANKRD26 ATP5F1E ACTB AKAP9 ANTXR2 ATP6V0A2 ACTC1 AKR1D1 AP1S2 ATP6V1B1 ACTG1 AKT2 AP2S1 ATP7A ACTG2 AKT3 AP3B1 ATP8A2 ACTL6B ALAS2 AP3B2 ATP8B1 ACTN1 ALB AP4B1 ATPAF2 ACTN2 ALDH18A1 AP4M1 ATR ACTN4 ALDH1A3 AP4S1 ATRX ACVR1 ALDH3A2 APC AUH ACVRL1 ALDH4A1 APTX AVPR2 ACY1 ALDH5A1 AR B3GALNT2 ADA ALDH6A1 ARFGEF2 B3GALT6 ADAMTS13 ALDH7A1 ARG1 B3GAT3 ADAMTS2 ALDOB ARHGAP31 B3GLCT Updated: 03/15/2021; v.3.6 1 Neonatal Crisis Sequencing Panel Gene List Genes: B4GALT1 - COL11A2 B4GALT1 C1QBP CD3G CHKB B4GALT7 C3 CD40LG CHMP1A B4GAT1 CA2 CD59 CHRNA1 B9D1 CA5A CD70 CHRNB1 B9D2 CACNA1A CD96 CHRND BAAT CACNA1C CDAN1 CHRNE BBIP1 CACNA1D CDC42 CHRNG BBS1 CACNA1E CDH1 CHST14 BBS10 CACNA1F CDH2 CHST3 BBS12 CACNA1G CDK10 CHUK BBS2 CACNA2D2 CDK13 CILK1 BBS4 CACNB2 CDK5RAP2 -

Next Generation Massively Parallel Sequencing of Targeted
ARTICLE Next generation massively parallel sequencing of targeted exomes to identify genetic mutations in primary ciliary dyskinesia: Implications for application to clinical testing Jonathan S. Berg, MD, PhD1,2, James P. Evans, MD, PhD1,2, Margaret W. Leigh, MD3, Heymut Omran, MD4, Chris Bizon, PhD5, Ketan Mane, PhD5, Michael R. Knowles, MD2, Karen E. Weck, MD1,6, and Maimoona A. Zariwala, PhD6 Purpose: Advances in genetic sequencing technology have the potential rimary ciliary dyskinesia (PCD) is an autosomal recessive to enhance testing for genes associated with genetically heterogeneous Pdisorder involving abnormalities of motile cilia, resulting in clinical syndromes, such as primary ciliary dyskinesia. The objective of a range of manifestations including situs inversus, neonatal this study was to investigate the performance characteristics of exon- respiratory distress at full-term birth, recurrent otitis media, capture technology coupled with massively parallel sequencing for chronic sinusitis, chronic bronchitis that may result in bronchi- 1–3 clinical diagnostic evaluation. Methods: We performed a pilot study of ectasis, and male infertility. The disorder is genetically het- four individuals with a variety of previously identified primary ciliary erogeneous, rendering molecular diagnosis challenging given dyskinesia mutations. We designed a custom array (NimbleGen) to that mutations in nine different genes (DNAH5, DNAH11, capture 2089 exons from 79 genes associated with primary ciliary DNAI1, DNAI2, KTU, LRRC50, RSPH9, RSPH4A, and TX- dyskinesia or ciliary function and sequenced the enriched material using NDC3) account for only approximately 1/3 of PCD cases.4 the GS FLX Titanium (Roche 454) platform. Bioinformatics analysis DNAH5 and DNAI1 account for the majority of known muta- was performed in a blinded fashion in an attempt to detect the previ- tions, and the other genes each account for a small number of ously identified mutations and validate the process. -

A Missense Mutation in the RSRSP Stretch of Rbm20 Causes Dilated
www.nature.com/scientificreports OPEN A missense mutation in the RSRSP stretch of Rbm20 causes dilated cardiomyopathy and atrial fbrillation in mice Kensuke Ihara1,2*, Tetsuo Sasano2, Yuichi Hiraoka3, Marina Togo‑Ohno4, Yurie Soejima5, Motoji Sawabe5, Megumi Tsuchiya6, Hidesato Ogawa6, Tetsushi Furukawa1 & Hidehito Kuroyanagi4* Dilated cardiomyopathy (DCM) is a fatal heart disease characterized by left ventricular dilatation and cardiac dysfunction. Recent genetic studies on DCM have identifed causative mutations in over 60 genes, including RBM20, which encodes a regulator of heart‑specifc splicing. DCM patients with RBM20 mutations have been reported to present with more severe cardiac phenotypes, including impaired cardiac function, atrial fbrillation (AF), and ventricular arrhythmias leading to sudden cardiac death, compared to those with mutations in the other genes. An RSRSP stretch of RBM20, a hotspot of missense mutations found in patients with idiopathic DCM, functions as a crucial part of its nuclear localization signals. However, the relationship between mutations in the RSRSP stretch and cardiac phenotypes has never been assessed in an animal model. Here, we show that Rbm20 mutant mice harboring a missense mutation S637A in the RSRSP stretch, mimicking that in a DCM patient, demonstrated severe cardiac dysfunction and spontaneous AF and ventricular arrhythmias mimicking the clinical state in patients. In contrast, Rbm20 mutant mice with frame‑shifting deletion demonstrated less severe phenotypes, although loss of RBM20‑dependent alternative splicing was indistinguishable. RBM20S637A protein cannot be localized to the nuclear speckles, but accumulated in cytoplasmic, perinuclear granule‑like structures in cardiomyocytes, which might contribute to the more severe cardiac phenotypes. Dilated cardiomyopathy (DCM) is a fatal cardiac disease characterized by enlargement of the cardiac chambers and impaired systolic function1. -

Antibody List 2014
Antibody List 2014 Office of Technology Transfer, DKFZ Terms and Conditions Terms and conditions for licensing DKFZ’s antibodies are available upon request. Disclaimer This antibody list is neither exhaustive nor qualifies as a commercial catalogue. DKFZ has prepared the contents of this list to the best of its knowledge and confirms to the best of its knowledge that the listed antibodies are created at or in cooperation with DKFZ. DKFZ does not warrant that any listed antibody is available. Moreover DKFZ does not warrant that any listed antibody and / or the use thereof is not protected by any third party rights. Contact: Office of Technology Transfer (T010) Deutsches Krebsforschungszentrum (DKFZ) German Cancer Research Center Im Neuenheimer Feld 280 69120 Heidelberg Germany E-mail: [email protected] Index MMP9 AATF Hexa-His MPO ABCB5 HIP128 Myo II AID HIP150 NET1 AGO1 HIPK-2 Nephrin AuroraC HIPPI NF1 APC Histone H1.0 NM-1 (NMy1) BPAG Histone H1.2 NMUR-2 BPV1 L1 Histone H1.3 NO-52 BPV1 E7 Histone H1.4 NO-66 BTG2 HLA-DM NO-145 CCDC103 HLA-DO NPHP1 CCR4 HLA-DP NPHP3 CCR8 HLA-DQ NPHP4 CCR9 HLA-DR NPM CCR10 HPV11 L1 Nup88 CD3 HPV11 E2 Nus/S-tag CD19 HPV16 L1 OCAB CD22 HPV16 L2 ODA7 CD23 HPV16 E5 PACS-1 CD37 HPV16 E6 PACT CD53 HPV16 E7 p16 CD56 HPV18 E6 p27KIP CD71 HPV18 E7 p53 CD171 (L1CAM) HPV20 E6 Pick CEACAM1 HPV20 E7 PKD1 Cep170 HPV27 E6 PIF Densin HPV27 E7 Plasminogen acitvator (PA) DLL4 HSC70 Podocin DNAH3 HSP70 PSA DNAH5 Jade1 PSMA DNAH8 JAM4 PTEN DNAH11 Ki67 RFP2 DNAse1 KTU RGS7 DSC2 LaminA RIP140 EGFR LaminB1 -

Supplementary Table S4. FGA Co-Expressed Gene List in LUAD
Supplementary Table S4. FGA co-expressed gene list in LUAD tumors Symbol R Locus Description FGG 0.919 4q28 fibrinogen gamma chain FGL1 0.635 8p22 fibrinogen-like 1 SLC7A2 0.536 8p22 solute carrier family 7 (cationic amino acid transporter, y+ system), member 2 DUSP4 0.521 8p12-p11 dual specificity phosphatase 4 HAL 0.51 12q22-q24.1histidine ammonia-lyase PDE4D 0.499 5q12 phosphodiesterase 4D, cAMP-specific FURIN 0.497 15q26.1 furin (paired basic amino acid cleaving enzyme) CPS1 0.49 2q35 carbamoyl-phosphate synthase 1, mitochondrial TESC 0.478 12q24.22 tescalcin INHA 0.465 2q35 inhibin, alpha S100P 0.461 4p16 S100 calcium binding protein P VPS37A 0.447 8p22 vacuolar protein sorting 37 homolog A (S. cerevisiae) SLC16A14 0.447 2q36.3 solute carrier family 16, member 14 PPARGC1A 0.443 4p15.1 peroxisome proliferator-activated receptor gamma, coactivator 1 alpha SIK1 0.435 21q22.3 salt-inducible kinase 1 IRS2 0.434 13q34 insulin receptor substrate 2 RND1 0.433 12q12 Rho family GTPase 1 HGD 0.433 3q13.33 homogentisate 1,2-dioxygenase PTP4A1 0.432 6q12 protein tyrosine phosphatase type IVA, member 1 C8orf4 0.428 8p11.2 chromosome 8 open reading frame 4 DDC 0.427 7p12.2 dopa decarboxylase (aromatic L-amino acid decarboxylase) TACC2 0.427 10q26 transforming, acidic coiled-coil containing protein 2 MUC13 0.422 3q21.2 mucin 13, cell surface associated C5 0.412 9q33-q34 complement component 5 NR4A2 0.412 2q22-q23 nuclear receptor subfamily 4, group A, member 2 EYS 0.411 6q12 eyes shut homolog (Drosophila) GPX2 0.406 14q24.1 glutathione peroxidase