Detection of Infectious Brome Mosaic Virus in Irrigation Ditches and Draining Strands in Poland
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Systematic, Genome-Wide Identification of Host Genes Affecting Replication of a Positive-Strand RNA Virus
Systematic, genome-wide identification of host genes affecting replication of a positive-strand RNA virus David B. Kushner*†, Brett D. Lindenbach*‡§, Valery Z. Grdzelishvili*, Amine O. Noueiry*, Scott M. Paul*¶, and Paul Ahlquist*‡ʈ *Institute for Molecular Virology and ‡Howard Hughes Medical Institute, University of Wisconsin, Madison, WI 53706 Contributed by Paul Ahlquist, October 23, 2003 Positive-strand RNA viruses are the largest virus class and include serves as a template for synthesis of a subgenomic (sg) mRNA, many pathogens such as hepatitis C virus and the severe acute RNA4, which encodes the viral coat protein (Fig. 1A). respiratory syndrome coronavirus (SARS). Brome mosaic virus The yeast Saccharomyces cerevisiae has proven a valuable (BMV) is a representative positive-strand RNA virus whose RNA model for normal and disease processes in human and other replication, gene expression, and encapsidation have been repro- cells. The unusual ability of BMV to direct its genomic RNA duced in the yeast Saccharomyces cerevisiae. By using traditional replication, gene expression, encapsidation, and other processes yeast genetics, host genes have been identified that function in in this yeast (7, 8) has allowed traditional yeast mutagenic controlling BMV translation, selecting BMV RNAs as replication analyses that have identified host genes involved in multiple steps templates, activating the replication complex, maintaining a lipid of BMV RNA replication and gene expression. Such host genes composition required for membrane-associated RNA replication, encode a wide variety of functions and contribute to diverse and other steps. To more globally and systematically identify such replication steps, including supporting and regulating viral trans- host factors, we used engineered BMV derivatives to assay viral lation, selecting and recruiting viral RNAs as replication RNA replication in each strain of an ordered, genome-wide set of templates, activating the RNA replication complex through yeast single-gene deletion mutants. -

UC Riverside UC Riverside Previously Published Works
UC Riverside UC Riverside Previously Published Works Title Viral RNAs are unusually compact. Permalink https://escholarship.org/uc/item/6b40r0rp Journal PloS one, 9(9) ISSN 1932-6203 Authors Gopal, Ajaykumar Egecioglu, Defne E Yoffe, Aron M et al. Publication Date 2014 DOI 10.1371/journal.pone.0105875 Peer reviewed eScholarship.org Powered by the California Digital Library University of California Viral RNAs Are Unusually Compact Ajaykumar Gopal1, Defne E. Egecioglu1, Aron M. Yoffe1, Avinoam Ben-Shaul2, Ayala L. N. Rao3, Charles M. Knobler1, William M. Gelbart1* 1 Department of Chemistry & Biochemistry, University of California Los Angeles, Los Angeles, California, United States of America, 2 Institute of Chemistry & The Fritz Haber Research Center, The Hebrew University of Jerusalem, Givat Ram, Jerusalem, Israel, 3 Department of Plant Pathology, University of California Riverside, Riverside, California, United States of America Abstract A majority of viruses are composed of long single-stranded genomic RNA molecules encapsulated by protein shells with diameters of just a few tens of nanometers. We examine the extent to which these viral RNAs have evolved to be physically compact molecules to facilitate encapsulation. Measurements of equal-length viral, non-viral, coding and non-coding RNAs show viral RNAs to have among the smallest sizes in solution, i.e., the highest gel-electrophoretic mobilities and the smallest hydrodynamic radii. Using graph-theoretical analyses we demonstrate that their sizes correlate with the compactness of branching patterns in predicted secondary structure ensembles. The density of branching is determined by the number and relative positions of 3-helix junctions, and is highly sensitive to the presence of rare higher-order junctions with 4 or more helices. -

Virus Particle Structures
Virus Particle Structures Virus Particle Structures Palmenberg, A.C. and Sgro, J.-Y. COLOR PLATE LEGENDS These color plates depict the relative sizes and comparative virion structures of multiple types of viruses. The renderings are based on data from published atomic coordinates as determined by X-ray crystallography. The international online repository for 3D coordinates is the Protein Databank (www.rcsb.org/pdb/), maintained by the Research Collaboratory for Structural Bioinformatics (RCSB). The VIPER web site (mmtsb.scripps.edu/viper), maintains a parallel collection of PDB coordinates for icosahedral viruses and additionally offers a version of each data file permuted into the same relative 3D orientation (Reddy, V., Natarajan, P., Okerberg, B., Li, K., Damodaran, K., Morton, R., Brooks, C. and Johnson, J. (2001). J. Virol., 75, 11943-11947). VIPER also contains an excellent repository of instructional materials pertaining to icosahedral symmetry and viral structures. All images presented here, except for the filamentous viruses, used the standard VIPER orientation along the icosahedral 2-fold axis. With the exception of Plate 3 as described below, these images were generated from their atomic coordinates using a novel radial depth-cue colorization technique and the program Rasmol (Sayle, R.A., Milner-White, E.J. (1995). RASMOL: biomolecular graphics for all. Trends Biochem Sci., 20, 374-376). First, the Temperature Factor column for every atom in a PDB coordinate file was edited to record a measure of the radial distance from the virion center. The files were rendered using the Rasmol spacefill menu, with specular and shadow options according to the Van de Waals radius of each atom. -

The Crystallographic Structure of Brome Mosaic Virus Robertw.Lucas,Stevenb.Larsonandalexandermcpherson*
doi:10.1006/jmbi.2001.5389availableonlineathttp://www.idealibrary.comon J. Mol. Biol. (2002) 317, 95±108 The Crystallographic Structure of Brome Mosaic Virus RobertW.Lucas,StevenB.LarsonandAlexanderMcPherson* University of California, Irvine The structure of brome mosaic virus (BMV), the type member of the 560 Steinhaus Hall, Irvine bromoviridae family, has been determined from a single rhombohedral CA 92697-3900, USA crystal by X-ray diffraction, and re®ned to an R value of 0.237 for data in the range 3.4-40.0 AÊ . The structure, which represents the native, compact form at pH 5.2 in the presence of 0.1 M Mg2, was solved by molecular replacement using the model of cowpea chlorotic mottle virus (CCMV), which BMV closely resembles. The BMV model contains amino acid resi- dues 41-189 for the pentameric capsid A subunits, and residues 25-189 and 1-189 for the B and C subunits, respectively, which compose the hex- americ capsomeres. In the model there are two Mg ions and one mol- ecule of polyethylene glycol (PEG). The ®rst 25 amino acid residues of the C subunit are modeled as polyalanine. The coat protein has the cano- nical ``jellyroll'' b-barrel topology with extended amino-terminal polypep- tides as seen in other icosahedral plant viruses. Mass spectrometry shows that in native BMV virions, a signi®cant fraction of the amino-terminal peptides are apparently cleaved. No recognizable nucleic acid residue is visible in the electron density maps except at low resolution where it appears to exhibit a layered arrangement in the virion interior. -

Parallels Among Positive-Strand RNA Viruses, Reverse-Transcribing Viruses and Double-Stranded RNA Viruses
REVIEWS Parallels among positive-strand RNA viruses, reverse-transcribing viruses and double-stranded RNA viruses Paul Ahlquist Abstract | Viruses are divided into seven classes on the basis of differing strategies for storing and replicating their genomes through RNA and/or DNA intermediates. Despite major differences among these classes, recent results reveal that the non-virion, intracellular RNA- replication complexes of some positive-strand RNA viruses share parallels with the structure, assembly and function of the replicative cores of extracellular virions of reverse-transcribing viruses and double-stranded RNA viruses. Therefore, at least four of seven principal virus classes share several underlying features in genome replication and might have emerged from common ancestors. This has implications for virus function, evolution and control. Positive-strand RNA virus Despite continuing advances, established and emerging ssRNA or dsRNA. Other viruses replicate by intercon- A virus, the infectious virions of viruses remain major causes of human disease, with verting their genomes between RNA and DNA. The viri- which contain the genome in a dramatic costs in mortality, morbidity and economic ons of such reverse-transcribing viruses always initially single-stranded, messenger- terms. In addition to acute diseases, viruses cause at package the RNA forms of their genomes, and either sense RNA form. least 15–20% of human cancers1,2 and are implicated in might (for example, hepadnaviruses and foamy retro- neurological and other chronic disorders. One of many viruses) or might not (for example, orthoretroviruses) challenges in controlling viruses and virus-mediated dis- reverse-transcribe the RNA into DNA before the virion eases is that viruses show an amazing diversity in basic exits the initially infected producer cell. -

Cowpea Chlorotic Mottle Bromovirus Replication Proteins Support Template- Selective RNA Replication in Saccharomyces Cerevisiae
RESEARCH ARTICLE Cowpea chlorotic mottle bromovirus replication proteins support template- selective RNA replication in Saccharomyces cerevisiae Bryan S. Sibert1,2, Amanda K. Navine1,3, Janice Pennington1,2, Xiaofeng Wang1¤, Paul Ahlquist1,2,3* a1111111111 1 Institute for Molecular Virology, University of Wisconsin-Madison, Madison, Wisconsin, United States of America, 2 Howard Hughes Medical Institute, University of Wisconsin-Madison, Madison, Wisconsin, United a1111111111 States of America, 3 John W. and Jeanne M. Rowe Center for Research in Virology, Morgridge Institute for a1111111111 Research, University of Wisconsin-Madison, Madison, Wisconsin, United States of America a1111111111 a1111111111 ¤ Current address: Department of Plant Pathology, Physiology, and Weed Science, Virginia Tech University, Blacksburg, Virginia, United States of America * [email protected] OPEN ACCESS Abstract Citation: Sibert BS, Navine AK, Pennington J, Wang X, Ahlquist P (2018) Cowpea chlorotic Positive-strand RNA viruses generally assemble RNA replication complexes on rearranged mottle bromovirus replication proteins support host membranes. Alphaviruses, other members of the alpha-like virus superfamily, and template-selective RNA replication in many other positive-strand RNA viruses invaginate host membrane into vesicular RNA repli- Saccharomyces cerevisiae. PLoS ONE 13(12): cation compartments, known as spherules, whose interior is connected to the cytoplasm. e0208743. https://doi.org/10.1371/journal. pone.0208743 Brome mosaic virus (BMV) and its close relative, cowpea chlorotic mottle virus (CCMV), form spherules along the endoplasmic reticulum. BMV spherule formation and RNA replication Editor: Sebastien Pfeffer, Institut de Biologie Moleculaire et Cellulaire, FRANCE can be fully reconstituted in S. cerevisiae, enabling many studies identifying host factors and viral interactions essential for these processes. -

Comparison of Plant‐Adapted Rhabdovirus Protein Localization and Interactions
University of Kentucky UKnowledge University of Kentucky Doctoral Dissertations Graduate School 2011 COMPARISON OF PLANT‐ADAPTED RHABDOVIRUS PROTEIN LOCALIZATION AND INTERACTIONS Kathleen Marie Martin University of Kentucky, [email protected] Right click to open a feedback form in a new tab to let us know how this document benefits ou.y Recommended Citation Martin, Kathleen Marie, "COMPARISON OF PLANT‐ADAPTED RHABDOVIRUS PROTEIN LOCALIZATION AND INTERACTIONS" (2011). University of Kentucky Doctoral Dissertations. 172. https://uknowledge.uky.edu/gradschool_diss/172 This Dissertation is brought to you for free and open access by the Graduate School at UKnowledge. It has been accepted for inclusion in University of Kentucky Doctoral Dissertations by an authorized administrator of UKnowledge. For more information, please contact [email protected]. ABSTRACT OF DISSERTATION Kathleen Marie Martin The Graduate School University of Kentucky 2011 COMPARISON OF PLANT‐ADAPTED RHABDOVIRUS PROTEIN LOCALIZATION AND INTERACTIONS ABSTRACT OF DISSERTATION A dissertation submitted in partial fulfillment of the requirements for the Degree of Doctor of Philosophy in the College of Agriculture at the University of Kentucky By Kathleen Marie Martin Lexington, Kentucky Director: Dr. Michael M Goodin, Associate Professor of Plant Pathology Lexington, Kentucky 2011 Copyright © Kathleen Marie Martin 2011 ABSTRACT OF DISSERTATION COMPARISON OF PLANT‐ADAPTED RHABDOVIRUS PROTEIN LOCALIZATION AND INTERACTIONS Sonchus yellow net virus (SYNV), Potato yellow dwarf virus (PYDV) and Lettuce Necrotic yellows virus (LNYV) are members of the Rhabdoviridae family that infect plants. SYNV and PYDV are Nucleorhabdoviruses that replicate in the nuclei of infected cells and LNYV is a Cytorhabdovirus that replicates in the cytoplasm. LNYV and SYNV share a similar genome organization with a gene order of Nucleoprotein (N), Phosphoprotein (P), putative movement protein (Mv), Matrix protein (M), Glycoprotein (G) and Polymerase protein (L). -

Tically Expands Our Understanding on Virosphere in Temperate Forest Ecosystems
Preprints (www.preprints.org) | NOT PEER-REVIEWED | Posted: 21 June 2021 doi:10.20944/preprints202106.0526.v1 Review Towards the forest virome: next-generation-sequencing dras- tically expands our understanding on virosphere in temperate forest ecosystems Artemis Rumbou 1,*, Eeva J. Vainio 2 and Carmen Büttner 1 1 Faculty of Life Sciences, Albrecht Daniel Thaer-Institute of Agricultural and Horticultural Sciences, Humboldt-Universität zu Berlin, Ber- lin, Germany; [email protected], [email protected] 2 Natural Resources Institute Finland, Latokartanonkaari 9, 00790, Helsinki, Finland; [email protected] * Correspondence: [email protected] Abstract: Forest health is dependent on the variability of microorganisms interacting with the host tree/holobiont. Symbiotic mi- crobiota and pathogens engage in a permanent interplay, which influences the host. Thanks to the development of NGS technol- ogies, a vast amount of genetic information on the virosphere of temperate forests has been gained the last seven years. To estimate the qualitative/quantitative impact of NGS in forest virology, we have summarized viruses affecting major tree/shrub species and their fungal associates, including fungal plant pathogens, mutualists and saprotrophs. The contribution of NGS methods is ex- tremely significant for forest virology. Reviewed data about viral presence in holobionts, allowed us to address the role of the virome in the holobionts. Genetic variation is a crucial aspect in hologenome, significantly reinforced by horizontal gene transfer among all interacting actors. Through virus-virus interplays synergistic or antagonistic relations may evolve, which may drasti- cally affect the health of the holobiont. Novel insights of these interplays may allow practical applications for forest plant protec- tion based on endophytes and mycovirus biocontrol agents. -

Synthesis and Characterization of a Full-Length Infectious Cdna Clone of Tomato Mottle Mosaic Virus
viruses Article Synthesis and Characterization of a Full-Length Infectious cDNA Clone of Tomato Mottle Mosaic Virus Liqin Tu 1,2 , Shuhua Wu 2, Danna Gao 1, Yong Liu 3, Yuelin Zhu 1,* and Yinghua Ji 2,* 1 College of Horticulture, Nanjing Agricultural University, Nanjing 210095, China; [email protected] (L.T.); [email protected] (D.G.) 2 Institute of Plant Protection, Jiangsu Academy of Agricultural Sciences/Key Lab of Food Quality and Safety of Jiangsu Province-State Key Laboratory Breeding Base, Nanjing 210014, China; [email protected] 3 Institute of Plant Protection, Hunan Academy of Agricultural Sciences, Changsha 410125, China; [email protected] * Correspondence: [email protected] (Y.Z.); [email protected] (Y.J.); Tel.: +86-25-84396472 (Y.Z.); +86-25-84390394 (Y.J.) Abstract: Tomato mottle mosaic virus (ToMMV) is a noteworthy virus which belongs to the Virgaviridae family and causes serious economic losses in tomato. Here, we isolated and cloned the full-length genome of a ToMMV Chinese isolate (ToMMV-LN) from a naturally infected tomato (Solanum lycopersicum L.). Sequence analysis showed that ToMMV-LN contains 6399 nucleotides (nts) and is most closely related to a ToMMV Mexican isolate with a sequence identity of 99.48%. Next, an infectious cDNA clone of ToMMV was constructed by a homologous recombination approach. Both the model host N. benthamiana and the natural hosts tomato and pepper developed severe symptoms upon agroinfiltration with pToMMV, which had a strong infectivity. Electron micrographs indicated that a large number of rigid rod-shaped ToMMV virions were observed from the agroinfiltrated N. -

Crystallization of Brome Mosaic Virus and T = 1 Brome Mosaic
Virology 286, 290–303 (2001) doi:10.1006/viro.2000.0897, available online at http://www.idealibrary.com on Crystallization of Brome Mosaic Virus and T ϭ 1 Brome Mosaic Virus Particles Following a Structural Transition Robert W. Lucas, Yurii G. Kuznetsov, Steven B. Larson, and Alexander McPherson1 University of California, Irvine, Department of Molecular Biology and Biochemistry, Irvine, California 92697-3900 Received November 9, 2000; returned to author for revision January 17, 2001; accepted March 6, 2001 Brome mosaic virus (BMV), a T ϭ 3 icosahedral plant virus, can be dissociated into coat protein subunits and subunit oligomers at pH 7.5 in the presence of concentrated salts. We have found that during the course of this treatment the coat protein subunits are cleaved, presumably by plant cell proteases still present in the preparation, between amino acids 35 and 36. The truncated protein subunits will then reorganize into T ϭ 1 icosahedral particles and can be crystallized from sodium malonate. Quasi elastic light scattering and atomic force microscopy results suggest that the transition from T ϭ 3toT ϭ 1 particles can occur by separate pathways, dissociation into coat protein subunits and oligomers and reassembly into T ϭ 1 particles, or direct condensation of the T ϭ 3 virions to T ϭ 1 particles with the shedding of hexameric capsomeres. The latter process has been directly visualized using atomic force microscopy. Native T ϭ 3 virions have been crystallized in several different crystal forms, but neither a rhombohedral form nor either of two orthorhombic forms diffract beyond about 3.4 Å. -
Eukaryotic Cell Eukaryotic Cells Are Defined As Cells Containing Organized Nucleus and Organelles Which Are Enveloped by Membrane-Bound Organelles
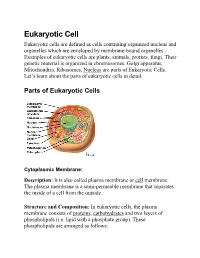
Eukaryotic Cell Eukaryotic cells are defined as cells containing organized nucleus and organelles which are enveloped by membrane-bound organelles. Examples of eukaryotic cells are plants, animals, protists, fungi. Their genetic material is organized in chromosomes. Golgi apparatus, Mitochondria, Ribosomes, Nucleus are parts of Eukaryotic Cells. Let’s learn about the parts of eukaryotic cells in detail. Parts ot Eukaryotic Cells Cytoplasmic Membrane: Description: It is also called plasma membrane or cell membrane. The plasma membrane is a semi-permeable membrane that separates the inside of a cell from the outside. Structure and Composition: In eukaryotic cells, the plasma membrane consists of proteins , carbohydrates and two layers of phospholipids (i.e. lipid with a phosphate group). These phospholipids are arranged as follows: • The polar, hydrophilic (water-loving) heads face the outside and inside of the cell. These heads interact with the aqueous environment outside and within a cell. • The non-polar, hydrophobic (water-repelling) tails are sandwiched between the heads and are protected from the aqueous environments. Scientists Singer and Nicolson(1972) described the structure of the phospholipid bilayer as the ‘Fluid Mosaic Model’. The reason is that the bi-layer looks like a mosaic and has a semi-fluid nature that allows lateral movement of proteins within the bilayer. Image: Fluid mosaic model. Orange circles – Hydrophilic heads; Lines below – Hydrophobic tails. Functions • The plasma membrane is selectively permeable i.e. it allows only selected substances to pass through. • It protects the cells from shock and injuries. • The fluid nature of the membrane allows the interaction of molecules within the membrane. -

Positive-Strand RNA Viruses Stimulate Host Phosphatidylcholine Synthesis at Viral Replication Sites
Positive-strand RNA viruses stimulate host phosphatidylcholine synthesis at viral replication sites Jiantao Zhanga,1, Zhenlu Zhanga,b, Vineela Chukkapallic, Jules A. Nchoutmboubed, Jianhui Lia, Glenn Randallc, George A. Belovd, and Xiaofeng Wanga,2 aDepartment of Plant Pathology, Physiology, and Weed Science, Virginia Tech, Blacksburg, VA 24061; bDepartment of Plant Protection, Fujian Agriculture and Forestry University, Fuzhou 350002, People’s Republic of China; cDepartment of Microbiology, The University of Chicago, Chicago, IL 60637; and dVirginia-Maryland Regional College of Veterinary Medicine, University of Maryland, College Park, MD 20742 Edited by Peter Palese, Icahn School of Medicine at Mount Sinai, New York, NY, and approved January 12, 2016 (received for review October 6, 2015) All positive-strand RNA viruses reorganize host intracellular mem- expression of 1a alone in yeast induces spherule formation (6), branes to assemble their viral replication complexes (VRCs); how- which requires 1a’s amphipathic α-helix (1a amino acids 392–407) ever, how these viruses modulate host lipid metabolism to (8), helix A, and 1a–1a interactions (9, 10). In addition, several accommodate such membrane proliferation and rearrangements is host proteins, including membrane-shaping reticulons (RTNs) not well defined. We show that a significantly increased phospha- (11) and an ESCRT (endosomal sorting complex required for tidylcholine (PC) content is associated with brome mosaic virus transport) component, Snf7p (sucrose nonfermenting7) (12), are (BMV) replication in both natural host barley and alternate host recruited by 1a to form spherules. yeast based on a lipidomic analysis. Enhanced PC levels are primarily Similar to other (+)RNA viruses, BMV promotes host lipid associated with the perinuclear ER membrane, where BMV replica- synthesis and requires balanced lipids for the formation and ac- tion takes place.