Fe-S Protein Synthesis in Green Algae Mitochondria
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

A Spatial Interactome Reveals the Protein Organization of the Algal
HHS Public Access Author manuscript Author ManuscriptAuthor Manuscript Author Cell. Author Manuscript Author manuscript; Manuscript Author available in PMC 2018 September 21. Published in final edited form as: Cell. 2017 September 21; 171(1): 133–147.e14. doi:10.1016/j.cell.2017.08.044. A Spatial Interactome Reveals the Protein Organization of the Algal CO2 Concentrating Mechanism Luke C.M. Mackinder1,2, Chris Chen1,3, Ryan D. Leib4, Weronika Patena1, Sean R. Blum5, Matthew Rodman3, Silvia Ramundo6, Christopher M. Adams4, and Martin C. Jonikas1,3,7,8,* 1Department of Plant Biology, Carnegie Institution for Science, Stanford, CA 94305, USA 3Department of Biology, Stanford University, Stanford, CA 94305, USA 4Stanford University Mass Spectrometry, Stanford University, Stanford, CA, USA 5Department of Biomolecular Engineering, UC Santa Cruz, Santa Cruz, CA 95064, USA 6Department of Biochemistry and Biophysics, University of California, San Francisco, CA 94158, USA 7Department of Molecular Biology, Princeton University, Princeton, NJ 08544, USA SUMMARY Approximately one-third of global CO2 fixation is performed by eukaryotic algae. Nearly all algae enhance their carbon assimilation by operating a CO2-concentrating mechanism (CCM) built around an organelle called the pyrenoid, whose protein composition is largely unknown. Here, we developed tools in the model alga Chlamydomonas reinhardtii to determine the localizations of 135 candidate CCM proteins, and physical interactors of 38 of these proteins. Our data reveal the identity of 89 pyrenoid proteins, including Rubisco-interacting proteins, photosystem I assembly factor candidates and inorganic carbon flux components. We identify three previously un- described protein layers of the pyrenoid: a plate-like layer, a mesh layer and a punctate layer. -
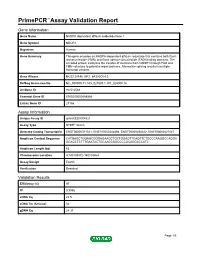
Primepcr™Assay Validation Report
PrimePCR™Assay Validation Report Gene Information Gene Name NADPH dependent diflavin oxidoreductase 1 Gene Symbol NDOR1 Organism Human Gene Summary This gene encodes an NADPH-dependent diflavin reductase that contains both flavin mononucleotide (FMN) and flavin adenine dinucleotide (FAD) binding domains. The encoded protein catalyzes the transfer of electrons from NADPH through FAD and FMN cofactors to potential redox partners. Alternative splicing results in multiple transcript variants. Gene Aliases MGC138148, NR1, bA350O14.9 RefSeq Accession No. NC_000009.11, NG_027801.1, NT_024000.16 UniGene ID Hs.512564 Ensembl Gene ID ENSG00000188566 Entrez Gene ID 27158 Assay Information Unique Assay ID qHsaCED0004821 Assay Type SYBR® Green Detected Coding Transcript(s) ENST00000371521, ENST00000344894, ENST00000458322, ENST00000427047 Amplicon Context Sequence CATGAGCTGGAGCGGGAGAAGCTGCTGGAGTTCAGTTCTGCCCAAGGCCAGGA GGAGCTCTTTGAATACTGCAACCGGCCCCGCAGGACCATC Amplicon Length (bp) 63 Chromosome Location 9:140109372-140109464 Assay Design Exonic Purification Desalted Validation Results Efficiency (%) 97 R2 0.9996 cDNA Cq 22.5 cDNA Tm (Celsius) 82 gDNA Cq 24.37 Page 1/5 PrimePCR™Assay Validation Report Specificity (%) 100 Information to assist with data interpretation is provided at the end of this report. Page 2/5 PrimePCR™Assay Validation Report NDOR1, Human Amplification Plot Amplification of cDNA generated from 25 ng of universal reference RNA Melt Peak Melt curve analysis of above amplification Standard Curve Standard curve generated using 20 million copies of template diluted 10-fold to 20 copies Page 3/5 PrimePCR™Assay Validation Report Products used to generate validation data Real-Time PCR Instrument CFX384 Real-Time PCR Detection System Reverse Transcription Reagent iScript™ Advanced cDNA Synthesis Kit for RT-qPCR Real-Time PCR Supermix SsoAdvanced™ SYBR® Green Supermix Experimental Sample qPCR Human Reference Total RNA Data Interpretation Unique Assay ID This is a unique identifier that can be used to identify the assay in the literature and online. -

Bacterial-Type Plant Ferroxidases Tune Local Phosphate Sensing in Root Development
bioRxiv preprint doi: https://doi.org/10.1101/2021.03.19.436157; this version posted March 21, 2021. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Bacterial-type plant ferroxidases tune local phosphate sensing in root development Christin Naumann1, Marcus Heisters1,2, Wolfgang Brandt3, Philipp Janitza4, Carolin Alfs1, Nancy Tang1, Alicia Toto Nienguesso1,5, Joerg Ziegler1, Richard Imre6,7, Karl Mechtler6,7, Yasin Dagdas6, Wolfgang Hoehenwarter8, Gary Sawers9, Marcel Quint4, and Steffen Abel1,10,11 1Department of Molecular Signal Processing, Leibniz Institute of Plant Biochemistry, Halle (Saale), Germany 2VEROVACCiNES GmbH, Halle (Saale), Germany 3Department of Bioorganic Chemistry, Leibniz Institute of Plant Biochemistry, Halle (Saale), Germany 4Institute of Agricultural and Nutritional Sciences, Martin Luther University Halle- Wittenberg, Halle (Saale), Germany 5Department of Anatomy and Cell Biology, University Hospital Halle, Halle (Saale), Germany 6Gregor Mendel Institute of Molecular Plant Biology, Vienna, Austria 7Research Institute of Molecular Pathology (IMP), Vienna BioCenter (VBC), Vienna, Austria 9Institute of Biology/Microbiology, Martin Luther University Halle-Wittenberg, Halle (Saale), Germany 8Proteome Analytics, Leibniz Institute of Plant Biochemistry, Halle (Saale), Germany 10Institute of Biochemistry and Biotechnology, Martin Luther University Halle-Wittenberg, Halle (Saale), Germany 11Department of Plant Sciences, University of California, Davis, CA, USA 1 bioRxiv preprint doi: https://doi.org/10.1101/2021.03.19.436157; this version posted March 21, 2021. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Abstract Fluctuating bioavailability of inorganic phosphate (Pi), often caused by complex Pi-metal interactions, guide root tip growth and root system architecture for maximizing the foraged soil volume. -

Structure of the Human Frataxin-Bound Iron-Sulfur Cluster Assembly Complex Provides Insight Into Its Activation Mechanism
bioRxiv preprint doi: https://doi.org/10.1101/561795; this version posted February 28, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY 4.0 International license. Structure of the human frataxin-bound iron-sulfur cluster assembly complex provides insight into its activation mechanism Nicholas G. Fox1,4, Xiaodi Yu2,4, Xidong Feng2, Henry J. Bailey1, Alain Martelli3, Joseph F. Nabhan3, Claire Strain-Damerell1, Christine Bulawa3, Wyatt W. Yue1,*, Seungil Han2,* 1Structural Genomics Consortium, Nuffield Department of Clinical Medicine, University of Oxford, UK OX3 7DQ 2Discovery Sciences, Worldwide Research and Development, Pfizer Inc., Eastern Point Road, Groton, CT, 06340, United States 3Rare Disease Research Unit, Worldwide Research and Development, Pfizer Inc., 610 Main Street, Cambridge, MA, 02139, United States 4These authors contributed equally. *Correspondence should be addressed to Wyatt W. Yue, [email protected] Seungil Han, [email protected] 1 bioRxiv preprint doi: https://doi.org/10.1101/561795; this version posted February 28, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY 4.0 International license. Abstract Iron-sulfur clusters (ISC) are essential in all life forms and carry out many crucial cellular functions. The core machinery for de novo ISC biosynthesis, located in the mitochondria matrix, is a five- protein complex containing the cysteine desulfurase NFS1 that is activated by frataxin (FXN), scaffold protein ISCU, accessory protein ISD11, and acyl-carrier protein ACP. -

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

Geometric and Electronic Structure Differences Between the Type 3 Copper Sites of the Multicopper Oxidases and Hemocyanin/Tyrosinase
Geometric and electronic structure differences between the type 3 copper sites of the multicopper oxidases and hemocyanin/tyrosinase Jungjoo Yoon, Satoshi Fujii1, and Edward I. Solomon2 Department of Chemistry, Stanford University, Stanford, CA 94305-5080 Contributed by Edward I. Solomon, February 25, 2009 (sent for review January 31, 2009) The coupled binuclear ‘‘type 3’’ Cu sites are found in hemocyanin (Hc), tyrosinase (Tyr), and the multicopper oxidases (MCOs), such as laccase (Lc), and play vital roles in O2 respiration. Although all type 3 Cu sites share the same ground state features, those of Hc/Tyr have very different ligand-binding properties relative to those of the MCOs. In particular, the type 3 Cu site in the MCOs (LcT3)isa part of the trinuclear Cu cluster, and if the third (i.e., type 2) Cu is T3 removed, the Lc site does not react with O2. Density functional ؊1 theory calculations indicate that O2 binding in Hc is Ϸ9 kcal mol more favorable than for LcT3. The difference is mostly found in the Ϸ ؊1 total energy difference of the deoxy states ( 7 kcal mol ), where Fig. 1. O2 binding in the type 3 Cu sites of Hc/Tyr and MCOs. the stabilization of deoxy LcT3 derives from its long equilibrium Cu–Cu distance of Ϸ5.5–6.5 Å, relative to Ϸ4.2 Å in deoxy Hc/Tyr. The O2 binding in Hc is driven by the electrostatic destabilization Cu–Cu distance of Ϸ3.6 Å (Fig. 1A) (7, 8, 10). The side-on 2Ϫ of the deoxy Hc site, in which the two Cu(I) centers are kept close geometry promotes a large overlap between the O2 * and the together by the protein for facile 2-electron reduction of O2. -

Anti-NFS1 Antibody (ARG56331)
Product datasheet [email protected] ARG56331 Package: 100 μl anti-NFS1 antibody Store at: -20°C Summary Product Description Rabbit Polyclonal antibody recognizes NFS1 Tested Reactivity Hu, Ms, Rat Tested Application ICC/IF, IHC-P, WB Host Rabbit Clonality Polyclonal Isotype IgG Target Name NFS1 Antigen Species Human Immunogen Recombinant protein of Human NFS1 Conjugation Un-conjugated Alternate Names HUSSY-08; NIFS; EC 2.8.1.7; IscS; Cysteine desulfurase, mitochondrial Application Instructions Application table Application Dilution ICC/IF 1:50 - 1:200 IHC-P 1:50 - 1:200 WB 1:500 - 1:2000 Application Note * The dilutions indicate recommended starting dilutions and the optimal dilutions or concentrations should be determined by the scientist. Positive Control Rat liver Calculated Mw 50 kDa Properties Form Liquid Purification Affinity purification with immunogen. Buffer PBS (pH 7.3), 0.02% Sodium azide and 50% Glycerol. Preservative 0.02% Sodium azide Stabilizer 50% Glycerol Storage instruction For continuous use, store undiluted antibody at 2-8°C for up to a week. For long-term storage, aliquot and store at -20°C. Storage in frost free freezers is not recommended. Avoid repeated freeze/thaw cycles. Suggest spin the vial prior to opening. The antibody solution should be gently mixed before use. www.arigobio.com 1/3 Note For laboratory research only, not for drug, diagnostic or other use. Bioinformation Database links GeneID: 18041 Mouse GeneID: 9054 Human Swiss-port # Q9Y697 Human Swiss-port # Q9Z1J3 Mouse Gene Symbol NFS1 Gene Full Name NFS1 cysteine desulfurase Background Iron-sulfur clusters are required for the function of many cellular enzymes. -

The Plankton Lifeform Extraction Tool: a Digital Tool to Increase The
Discussions https://doi.org/10.5194/essd-2021-171 Earth System Preprint. Discussion started: 21 July 2021 Science c Author(s) 2021. CC BY 4.0 License. Open Access Open Data The Plankton Lifeform Extraction Tool: A digital tool to increase the discoverability and usability of plankton time-series data Clare Ostle1*, Kevin Paxman1, Carolyn A. Graves2, Mathew Arnold1, Felipe Artigas3, Angus Atkinson4, Anaïs Aubert5, Malcolm Baptie6, Beth Bear7, Jacob Bedford8, Michael Best9, Eileen 5 Bresnan10, Rachel Brittain1, Derek Broughton1, Alexandre Budria5,11, Kathryn Cook12, Michelle Devlin7, George Graham1, Nick Halliday1, Pierre Hélaouët1, Marie Johansen13, David G. Johns1, Dan Lear1, Margarita Machairopoulou10, April McKinney14, Adam Mellor14, Alex Milligan7, Sophie Pitois7, Isabelle Rombouts5, Cordula Scherer15, Paul Tett16, Claire Widdicombe4, and Abigail McQuatters-Gollop8 1 10 The Marine Biological Association (MBA), The Laboratory, Citadel Hill, Plymouth, PL1 2PB, UK. 2 Centre for Environment Fisheries and Aquacu∑lture Science (Cefas), Weymouth, UK. 3 Université du Littoral Côte d’Opale, Université de Lille, CNRS UMR 8187 LOG, Laboratoire d’Océanologie et de Géosciences, Wimereux, France. 4 Plymouth Marine Laboratory, Prospect Place, Plymouth, PL1 3DH, UK. 5 15 Muséum National d’Histoire Naturelle (MNHN), CRESCO, 38 UMS Patrinat, Dinard, France. 6 Scottish Environment Protection Agency, Angus Smith Building, Maxim 6, Parklands Avenue, Eurocentral, Holytown, North Lanarkshire ML1 4WQ, UK. 7 Centre for Environment Fisheries and Aquaculture Science (Cefas), Lowestoft, UK. 8 Marine Conservation Research Group, University of Plymouth, Drake Circus, Plymouth, PL4 8AA, UK. 9 20 The Environment Agency, Kingfisher House, Goldhay Way, Peterborough, PE4 6HL, UK. 10 Marine Scotland Science, Marine Laboratory, 375 Victoria Road, Aberdeen, AB11 9DB, UK. -

Yeast Genome Gazetteer P35-65
gazetteer Metabolism 35 tRNA modification mitochondrial transport amino-acid metabolism other tRNA-transcription activities vesicular transport (Golgi network, etc.) nitrogen and sulphur metabolism mRNA synthesis peroxisomal transport nucleotide metabolism mRNA processing (splicing) vacuolar transport phosphate metabolism mRNA processing (5’-end, 3’-end processing extracellular transport carbohydrate metabolism and mRNA degradation) cellular import lipid, fatty-acid and sterol metabolism other mRNA-transcription activities other intracellular-transport activities biosynthesis of vitamins, cofactors and RNA transport prosthetic groups other transcription activities Cellular organization and biogenesis 54 ionic homeostasis organization and biogenesis of cell wall and Protein synthesis 48 plasma membrane Energy 40 ribosomal proteins organization and biogenesis of glycolysis translation (initiation,elongation and cytoskeleton gluconeogenesis termination) organization and biogenesis of endoplasmic pentose-phosphate pathway translational control reticulum and Golgi tricarboxylic-acid pathway tRNA synthetases organization and biogenesis of chromosome respiration other protein-synthesis activities structure fermentation mitochondrial organization and biogenesis metabolism of energy reserves (glycogen Protein destination 49 peroxisomal organization and biogenesis and trehalose) protein folding and stabilization endosomal organization and biogenesis other energy-generation activities protein targeting, sorting and translocation vacuolar and lysosomal -
![NFS1 Mouse Monoclonal Antibody [Clone ID: OTI5D1] Product Data](https://docslib.b-cdn.net/cover/9613/nfs1-mouse-monoclonal-antibody-clone-id-oti5d1-product-data-549613.webp)
NFS1 Mouse Monoclonal Antibody [Clone ID: OTI5D1] Product Data
OriGene Technologies, Inc. 9620 Medical Center Drive, Ste 200 Rockville, MD 20850, US Phone: +1-888-267-4436 [email protected] EU: [email protected] CN: [email protected] Product datasheet for TA805820 NFS1 Mouse Monoclonal Antibody [Clone ID: OTI5D1] Product data: Product Type: Primary Antibodies Clone Name: OTI5D1 Applications: IHC, WB Recommended Dilution: WB 1:500, IHC 1:150 Reactivity: Human, Mouse, Rat Host: Mouse Isotype: IgG2a Clonality: Monoclonal Immunogen: Human recombinant protein fragment corresponding to amino acids 1-299 of human NFS1(NP_066923) produced in E.coli. Formulation: PBS (PH 7.3) containing 1% BSA, 50% glycerol and 0.02% sodium azide. Concentration: 1 mg/ml Purification: Purified from mouse ascites fluids or tissue culture supernatant by affinity chromatography (protein A/G) Conjugation: Unconjugated Storage: Store at -20°C as received. Stability: Stable for 12 months from date of receipt. Predicted Protein Size: 50 kDa Gene Name: NFS1 cysteine desulfurase Database Link: NP_066923 Entrez Gene 18041 MouseEntrez Gene 84594 RatEntrez Gene 9054 Human Q9Y697 This product is to be used for laboratory only. Not for diagnostic or therapeutic use. View online » ©2021 OriGene Technologies, Inc., 9620 Medical Center Drive, Ste 200, Rockville, MD 20850, US 1 / 3 NFS1 Mouse Monoclonal Antibody [Clone ID: OTI5D1] – TA805820 Background: Iron-sulfur clusters are required for the function of many cellular enzymes. The proteins encoded by this gene supply inorganic sulfur to these clusters by removing the sulfur from cysteine, creating alanine in the process. This gene uses alternate in-frame translation initiation sites to generate mitochondrial forms and cytoplasmic/nuclear forms. -

Downloaded Using the Database on Presence of This Metal for 24 H
fmicb-11-01834 August 10, 2020 Time: 12:25 # 1 ORIGINAL RESEARCH published: 10 August 2020 doi: 10.3389/fmicb.2020.01834 Metabolic Adaptation of Paracoccidioides brasiliensis in Response to in vitro Copper Deprivation Guilherme Petito1,2†, Juliana Santana de Curcio1†, Maristela Pereira1, Alexandre Melo Bailão1, Juliano Domiraci Paccez1, Gabriel Brum Tristão1, Camila Oliveira Barbosa de Morais1, Marcelo Valle de Souza3, Agenor de Castro Moreira Santos Junior3, Wagner Fontes3, Carlos André Ornelas Ricart3 and Célia Maria de Almeida Soares1* Edited by: 1 Laboratório de Biologia Molecular, Instituto de Ciências Biológicas, Universidade Federal de Goiás, Goiânia, Brazil, Carlos Pelleschi Taborda, 2 Programa de Pós-graduação em Genética e Biologia Molecular, Universidade Federal de Goiás, Goiânia, Brazil, University of São Paulo, Brazil 3 Departamento de Biologia Celular, Instituto de Biologia, Universidade de Brasília, Brasília, Brazil Reviewed by: Rosana Puccia, Federal University of São Paulo, Brazil Copper is an essential micronutrient for the performance of important biochemical Sandro Rogerio Almeida, processes such as respiration detoxification, and uptake of metals like iron. Studies University of São Paulo, Brazil Elizabeth R. Ballou, have shown that copper deprivation is a strategy used by the host against pathogenic University of Birmingham, fungi such as Cryptoccocus neoformans and Candida albicans during growth and United Kingdom development of infections in the lungs and kidneys. Although there are some *Correspondence: studies, little is known about the impact of copper deprivation in members of the Célia Maria de Almeida Soares [email protected] Paracoccidioides genus. Therefore, using isobaric tag labeling (iTRAQ)-Based proteomic †These authors have contributed approach and LC-MS/MS, we analyzed the impact of in vitro copper deprivation in equally to this work the metabolism of Paracoccidioides brasiliensis. -

Supplementary Table S4. FGA Co-Expressed Gene List in LUAD
Supplementary Table S4. FGA co-expressed gene list in LUAD tumors Symbol R Locus Description FGG 0.919 4q28 fibrinogen gamma chain FGL1 0.635 8p22 fibrinogen-like 1 SLC7A2 0.536 8p22 solute carrier family 7 (cationic amino acid transporter, y+ system), member 2 DUSP4 0.521 8p12-p11 dual specificity phosphatase 4 HAL 0.51 12q22-q24.1histidine ammonia-lyase PDE4D 0.499 5q12 phosphodiesterase 4D, cAMP-specific FURIN 0.497 15q26.1 furin (paired basic amino acid cleaving enzyme) CPS1 0.49 2q35 carbamoyl-phosphate synthase 1, mitochondrial TESC 0.478 12q24.22 tescalcin INHA 0.465 2q35 inhibin, alpha S100P 0.461 4p16 S100 calcium binding protein P VPS37A 0.447 8p22 vacuolar protein sorting 37 homolog A (S. cerevisiae) SLC16A14 0.447 2q36.3 solute carrier family 16, member 14 PPARGC1A 0.443 4p15.1 peroxisome proliferator-activated receptor gamma, coactivator 1 alpha SIK1 0.435 21q22.3 salt-inducible kinase 1 IRS2 0.434 13q34 insulin receptor substrate 2 RND1 0.433 12q12 Rho family GTPase 1 HGD 0.433 3q13.33 homogentisate 1,2-dioxygenase PTP4A1 0.432 6q12 protein tyrosine phosphatase type IVA, member 1 C8orf4 0.428 8p11.2 chromosome 8 open reading frame 4 DDC 0.427 7p12.2 dopa decarboxylase (aromatic L-amino acid decarboxylase) TACC2 0.427 10q26 transforming, acidic coiled-coil containing protein 2 MUC13 0.422 3q21.2 mucin 13, cell surface associated C5 0.412 9q33-q34 complement component 5 NR4A2 0.412 2q22-q23 nuclear receptor subfamily 4, group A, member 2 EYS 0.411 6q12 eyes shut homolog (Drosophila) GPX2 0.406 14q24.1 glutathione peroxidase