Genomic and Transcriptomic Investigations Into the Feed Efficiency Phenotype of Beef Cattle
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-
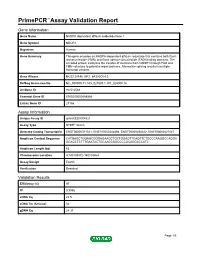
Primepcr™Assay Validation Report
PrimePCR™Assay Validation Report Gene Information Gene Name NADPH dependent diflavin oxidoreductase 1 Gene Symbol NDOR1 Organism Human Gene Summary This gene encodes an NADPH-dependent diflavin reductase that contains both flavin mononucleotide (FMN) and flavin adenine dinucleotide (FAD) binding domains. The encoded protein catalyzes the transfer of electrons from NADPH through FAD and FMN cofactors to potential redox partners. Alternative splicing results in multiple transcript variants. Gene Aliases MGC138148, NR1, bA350O14.9 RefSeq Accession No. NC_000009.11, NG_027801.1, NT_024000.16 UniGene ID Hs.512564 Ensembl Gene ID ENSG00000188566 Entrez Gene ID 27158 Assay Information Unique Assay ID qHsaCED0004821 Assay Type SYBR® Green Detected Coding Transcript(s) ENST00000371521, ENST00000344894, ENST00000458322, ENST00000427047 Amplicon Context Sequence CATGAGCTGGAGCGGGAGAAGCTGCTGGAGTTCAGTTCTGCCCAAGGCCAGGA GGAGCTCTTTGAATACTGCAACCGGCCCCGCAGGACCATC Amplicon Length (bp) 63 Chromosome Location 9:140109372-140109464 Assay Design Exonic Purification Desalted Validation Results Efficiency (%) 97 R2 0.9996 cDNA Cq 22.5 cDNA Tm (Celsius) 82 gDNA Cq 24.37 Page 1/5 PrimePCR™Assay Validation Report Specificity (%) 100 Information to assist with data interpretation is provided at the end of this report. Page 2/5 PrimePCR™Assay Validation Report NDOR1, Human Amplification Plot Amplification of cDNA generated from 25 ng of universal reference RNA Melt Peak Melt curve analysis of above amplification Standard Curve Standard curve generated using 20 million copies of template diluted 10-fold to 20 copies Page 3/5 PrimePCR™Assay Validation Report Products used to generate validation data Real-Time PCR Instrument CFX384 Real-Time PCR Detection System Reverse Transcription Reagent iScript™ Advanced cDNA Synthesis Kit for RT-qPCR Real-Time PCR Supermix SsoAdvanced™ SYBR® Green Supermix Experimental Sample qPCR Human Reference Total RNA Data Interpretation Unique Assay ID This is a unique identifier that can be used to identify the assay in the literature and online. -

Mayr, Annu Rev Genet 2017
GE51CH09-Mayr ARI 12 October 2017 10:21 Annual Review of Genetics Regulation by 3-Untranslated Regions Christine Mayr Department of Cancer Biology and Genetics, Memorial Sloan Kettering Cancer Center, New York, NY 10065, USA; email: [email protected] Annu. Rev. Genet. 2017. 51:171–94 Keywords First published as a Review in Advance on August noncoding RNA, regulatory RNA, alternative 3-UTRs, alternative 30, 2017 polyadenylation, cotranslational protein complex formation, cellular The Annual Review of Genetics is online at organization, mRNA localization, RNA-binding proteins, cooperativity, genet.annualreviews.org accessibility of regulatory elements https://doi.org/10.1146/annurev-genet-120116- 024704 Abstract Copyright c 2017 by Annual Reviews. 3 -untranslated regions (3 -UTRs) are the noncoding parts of mRNAs. Com- All rights reserved pared to yeast, in humans, median 3-UTR length has expanded approx- imately tenfold alongside an increased generation of alternative 3-UTR isoforms. In contrast, the number of coding genes, as well as coding region length, has remained similar. This suggests an important role for 3-UTRs in the biology of higher organisms. 3-UTRs are best known to regulate ANNUAL REVIEWS Further diverse fates of mRNAs, including degradation, translation, and localiza- Click here to view this article's tion, but they can also function like long noncoding or small RNAs, as has Annu. Rev. Genet. 2017.51:171-194. Downloaded from www.annualreviews.org online features: • Download figures as PPT slides been shown for whole 3 -UTRs as well as for cleaved fragments. Further- • Navigate linked references • Download citations more, 3 -UTRs determine the fate of proteins through the regulation of Access provided by Memorial Sloan-Kettering Cancer Center on 11/30/17. -

Rule Mining on Microrna Expression Profiles For
Faculty of Engineering and Information Technology University of Technology, Sydney Rule Mining on MicroRNA Expression Profiles for Human Disease Understanding A thesis submitted in partial fulfillment of the requirements for the degree of Doctor of Philosophy by Renhua Song July 2016 CERTIFICATE OF AUTHORSHIP/ORIGINALITY I certify that the work in this thesis has not previously been submitted for a degree nor has it been submitted as part of requirements for a degree except as fully acknowledged within the text. I also certify that the thesis has been written by me. Any help that I have received in my research work and the preparation of the thesis itself has been acknowledged. In addition, I certify that all information sources and literature used are indicated in the thesis. Signature of Candidate i Acknowledgments First, and foremost, I would like to express my gratitude to my chief supervisor, Assoc. Prof. Jinyan Li, and to my co-supervisors Assoc. Prof. Paul Kennedy and Assoc. Prof. Daniel Catchpoole. I am extremely grateful for all the advice and guidance so unselfishly given to me over the last three and half years by these three distinguished academics. This research would not have been possible without their high order supervision, support, assistance and leadership. The wonderful support and assistance provided by many people during this research is very much appreciated by me and my family. I am very grateful for the help that I received from all of these people but there are some special individuals that I must thank by name. A very sincere thank you is definitely owed to my loving husband Jing Liu, not only for his tremendous support during the last three and half years, but also his unfailing and often sorely tested patience. -

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

Cryptococcus Neoformans Can Form Titan-Like Cells in Vitro in Response to Multiple Signals
RESEARCH ARTICLE Cryptococcus neoformans can form titan-like cells in vitro in response to multiple signals Nuria Trevijano-Contador1¤, Haroldo Cesar de Oliveira1,2, RocõÂo GarcõÂa-Rodas1, SueÂlen Andreia Rossi1, Irene Llorente1, A ngel Zaballos3, Guilhem Janbon4, JoaquõÂn Ariño5, O scar Zaragoza1* 1 Mycology Reference Laboratory, National Centre for Microbiology, Instituto de Salud Carlos III, Majadahonda, Madrid, Spain, 2 Universidade Estadual Paulista (UNESP), Faculdade de Ciências Farmacêuticas, CaÃmpus Araraquara, Departamento de AnaÂlises ClõÂnicas, LaboratoÂrio de Micologia ClõÂnica, Araraquara, São Paulo, Brazil, 3 Genomics Unit, Core Scientific Services, Instituto de Salud Carlos III, a1111111111 Majadahonda, Madrid, Spain, 4 Institut Pasteur, Unite Biologie des ARN des Pathogènes Fongiques, a1111111111 DeÂpartement de Mycologie, Paris, France, 5 Institut de Biotecnologia i Biomedicina and Departament de a1111111111 BioquõÂmica i Biologia Molecular, Universitat Autònoma de Barcelona, Cerdanyola del Vallès, Spain a1111111111 a1111111111 ¤ Current address: Albert Einstein College of Medicine, Bronx, New York, United States of America. * [email protected] Abstract OPEN ACCESS Cryptococcus neoformans is an encapsulated pathogenic yeast that can change the size of Citation: Trevijano-Contador N, de Oliveira HC, GarcõÂa-Rodas R, Rossi SA, Llorente I, Zaballos AÂ, the cells during infection. In particular, this process can occur by enlarging the size of the et al. (2018) Cryptococcus neoformans can form capsule without modifying the size of the cell body, or by increasing the diameter of the cell titan-like cells in vitro in response to multiple body, which is normally accompanied by an increase of the capsule too. This last process signals. PLoS Pathog 14(5): e1007007. https://doi. leads to the formation of cells of an abnormal enlarged size denominated titan cells. -

Exploring Prostate Cancer Genome Reveals Simultaneous Losses of PTEN, FAS and PAPSS2 in Patients with PSA Recurrence After Radical Prostatectomy
Int. J. Mol. Sci. 2015, 16, 3856-3869; doi:10.3390/ijms16023856 OPEN ACCESS International Journal of Molecular Sciences ISSN 1422-0067 www.mdpi.com/journal/ijms Article Exploring Prostate Cancer Genome Reveals Simultaneous Losses of PTEN, FAS and PAPSS2 in Patients with PSA Recurrence after Radical Prostatectomy Chinyere Ibeawuchi 1, Hartmut Schmidt 2, Reinhard Voss 3, Ulf Titze 4, Mahmoud Abbas 5, Joerg Neumann 6, Elke Eltze 7, Agnes Marije Hoogland 8, Guido Jenster 9, Burkhard Brandt 10 and Axel Semjonow 1,* 1 Prostate Center, Department of Urology, University Hospital Muenster, Albert-Schweitzer-Campus 1, Gebaeude 1A, Muenster D-48149, Germany; E-Mail: [email protected] 2 Center for Laboratory Medicine, University Hospital Muenster, Albert-Schweitzer-Campus 1, Gebaeude 1A, Muenster D-48149, Germany; E-Mail: [email protected] 3 Interdisciplinary Center for Clinical Research, University of Muenster, Albert-Schweitzer-Campus 1, Gebaeude D3, Domagkstrasse 3, Muenster D-48149, Germany; E-Mail: [email protected] 4 Pathology, Lippe Hospital Detmold, Röntgenstrasse 18, Detmold D-32756, Germany; E-Mail: [email protected] 5 Institute of Pathology, Mathias-Spital-Rheine, Frankenburg Street 31, Rheine D-48431, Germany; E-Mail: [email protected] 6 Institute of Pathology, Klinikum Osnabrueck, Am Finkenhuegel 1, Osnabrueck D-49076, Germany; E-Mail: [email protected] 7 Institute of Pathology, Saarbrücken-Rastpfuhl, Rheinstrasse 2, Saarbrücken D-66113, Germany; E-Mail: [email protected] 8 Department -

Genome-Wide Association Study for Circulating Fibroblast Growth Factor
www.nature.com/scientificreports OPEN Genome‑wide association study for circulating fbroblast growth factor 21 and 23 Gwo‑Tsann Chuang1,2,14, Pi‑Hua Liu3,4,14, Tsui‑Wei Chyan5, Chen‑Hao Huang5, Yu‑Yao Huang4,6, Chia‑Hung Lin4,7, Jou‑Wei Lin8, Chih‑Neng Hsu8, Ru‑Yi Tsai8, Meng‑Lun Hsieh9, Hsiao‑Lin Lee9, Wei‑shun Yang2,9, Cassianne Robinson‑Cohen10, Chia‑Ni Hsiung11, Chen‑Yang Shen12,13 & Yi‑Cheng Chang2,9,12* Fibroblast growth factors (FGFs) 21 and 23 are recently identifed hormones regulating metabolism of glucose, lipid, phosphate and vitamin D. Here we conducted a genome‑wide association study (GWAS) for circulating FGF21 and FGF23 concentrations to identify their genetic determinants. We enrolled 5,000 participants from Taiwan Biobank for this GWAS. After excluding participants with diabetes mellitus and quality control, association of single nucleotide polymorphisms (SNPs) with log‑transformed FGF21 and FGF23 serum concentrations adjusted for age, sex and principal components of ancestry were analyzed. A second model additionally adjusted for body mass index (BMI) and a third model additionally adjusted for BMI and estimated glomerular fltration rate (eGFR) were used. A total of 4,201 participants underwent GWAS analysis. rs67327215, located within RGS6 (a gene involved in fatty acid synthesis), and two other SNPs (rs12565114 and rs9520257, located between PHC2-ZSCAN20 and ARGLU1-FAM155A respectively) showed suggestive associations with serum FGF21 level (P = 6.66 × 10–7, 6.00 × 10–7 and 6.11 × 10–7 respectively). The SNPs rs17111495 and rs17843626 were signifcantly associated with FGF23 level, with the former near PCSK9 gene and the latter near HLA-DQA1 gene (P = 1.04 × 10–10 and 1.80 × 10–8 respectively). -

UCLA UCLA Electronic Theses and Dissertations
UCLA UCLA Electronic Theses and Dissertations Title Global and Local Regulation of Gene Expression in the Human Brain Permalink https://escholarship.org/uc/item/83g7t3g1 Author Hartl, Christopher Publication Date 2019 Peer reviewed|Thesis/dissertation eScholarship.org Powered by the California Digital Library University of California UNIVERSITY OF CALIFORNIA Los Angeles Global and Local Regulation of Gene Expression in the Human Brain A dissertation submitted in partial satisfaction of the requirements for the degree of Doctor of Philosophy in Bioinformatics by Christopher Hartl 2019 © Copyright by Christopher Hartl 2019 ABSTRACT OF THE DISSERTATION Global and Local Regulation of Gene Expression in the Human Brain by Christopher Hartl Doctor of Philosophy in Bioinformatics University of California, Los Angeles, 2019 Professor Daniel H. Geschwind, Chair Neuropsychiatric disorders are behavioral conditions marked by intellectual, social, or emotional deficits that can be linked to diseases of the nervous system. Autism spectrum disorder (ASD), schizophrenia (SCZ), bipolar disorder (BP), major depressive disorder (MDD), and attention deficit and hyperactivity disorder (ADHD) are common, heritable diseases each with a prevalence exceeding 1% of the population, none of which can be characterized by discernable anatomical or neurological pathologies. Genetic association studies have identified mutations in hundreds of genes that contribute to risk for at least one of these disorders, and have shown that a substantial fraction of the genetic liability is shared between many of these neuropsychiatric diseases. It has long been hoped that with enough genetic evidence we will identify the biological pathways, developmental time points, and brain regions that, when disrupted, give rise to neuropsychiatric disorders. -

Modulating Hallmarks of Cholangiocarcinoma
University of Nebraska Medical Center DigitalCommons@UNMC Theses & Dissertations Graduate Studies Fall 12-14-2018 Modulating Hallmarks of Cholangiocarcinoma Cody Wehrkamp University of Nebraska Medical Center Follow this and additional works at: https://digitalcommons.unmc.edu/etd Part of the Molecular Biology Commons Recommended Citation Wehrkamp, Cody, "Modulating Hallmarks of Cholangiocarcinoma" (2018). Theses & Dissertations. 337. https://digitalcommons.unmc.edu/etd/337 This Dissertation is brought to you for free and open access by the Graduate Studies at DigitalCommons@UNMC. It has been accepted for inclusion in Theses & Dissertations by an authorized administrator of DigitalCommons@UNMC. For more information, please contact [email protected]. MODULATING HALLMARKS OF CHOLANGIOCARCINOMA by Cody J. Wehrkamp A DISSERTATION Presented to the Faculty of the University of Nebraska Graduate College in Partial Fulfillment of the Requirements for the Degree of Doctor of Philosophy Biochemistry and Molecular Biology Graduate Program Under the Supervision of Professor Justin L. Mott University of Nebraska Medical Center Omaha, Nebraska November 2018 Supervisory Committee: Kaustubh Datta, Ph.D. Melissa Teoh‐Fitzgerald, Ph.D. Richard G. MacDonald, Ph.D. Acknowledgements This endeavor has led to scientific as well as personal growth for me. I am indebted to many for their knowledge, influence, and support along the way. To my mentor, Dr. Justin L. Mott, you have been an incomparable teacher and invaluable guide. You upheld for me the concept that science is intrepid, even when the experience is trying. Through my training, and now here at the end, I can say that it has been an honor to be your protégé. When you have shaped your future graduates to be and do great, I will be privileged to say that I was your first one. -

Controls Homeostatic Splicing of ARGLU1 Mrna Stephan P
Published online 28 November 2016 Nucleic Acids Research, 2017, Vol. 45, No. 6 3473–3486 doi: 10.1093/nar/gkw1140 An Ultraconserved Element (UCE) controls homeostatic splicing of ARGLU1 mRNA Stephan P. Pirnie, Ahmad Osman, Yinzhou Zhu and Gordon G. Carmichael* Department of Genetics and Genome Sciences, UCONN Health Center, 400 Farmington Avenue, Farmington, CT 06030, USA Received January 13, 2016; Revised October 25, 2016; Editorial Decision October 28, 2016; Accepted October 31, 2016 ABSTRACT or inhibit the usage of a particular splice site (1,2). The ex- pression of trans-acting factors in a developmental and tis- Arginine and Glutamate-Rich protein 1 (ARGLU1) is sue specific manner results in regulated splicing that isof- a protein whose function is poorly understood, but ten cell specific. Most notably, trans-acting proteins such as may act in both transcription and pre-mRNA splicing. the NOVA (3–6), the RBFOX (7) and SR protein (8–11) We demonstrate that the ARGLU1 gene expresses families, and a number of hnRNP (12,13) proteins compete at least three distinct RNA splice isoforms – a fully to bind nascent RNAs at specific motifs and drive regula- spliced isoform coding for the protein, an isoform tion of alternative splicing in a tissue and developmentally containing a retained intron that is detained in the specific manner. Alternative splicing is an important pro- nucleus, and an isoform containing an alternative cess that is seen in at least 95% of multi-exon genes in the exon that targets the transcript for nonsense medi- human transcriptome (14). Furthermore, alternative splic- ated decay. -

Reconstructing Cell Cycle Pseudo Time-Series Via Single-Cell Transcriptome Data—Supplement
School of Natural Sciences and Mathematics Reconstructing Cell Cycle Pseudo Time-Series Via Single-Cell Transcriptome Data—Supplement UT Dallas Author(s): Michael Q. Zhang Rights: CC BY 4.0 (Attribution) ©2017 The Authors Citation: Liu, Zehua, Huazhe Lou, Kaikun Xie, Hao Wang, et al. 2017. "Reconstructing cell cycle pseudo time-series via single-cell transcriptome data." Nature Communications 8, doi:10.1038/s41467-017-00039-z This document is being made freely available by the Eugene McDermott Library of the University of Texas at Dallas with permission of the copyright owner. All rights are reserved under United States copyright law unless specified otherwise. File name: Supplementary Information Description: Supplementary figures, supplementary tables, supplementary notes, supplementary methods and supplementary references. CCNE1 CCNE1 CCNE1 CCNE1 36 40 32 34 32 35 30 32 28 30 30 28 28 26 24 25 Normalized Expression Normalized Expression Normalized Expression Normalized Expression 26 G1 S G2/M G1 S G2/M G1 S G2/M G1 S G2/M Cell Cycle Stage Cell Cycle Stage Cell Cycle Stage Cell Cycle Stage CCNE1 CCNE1 CCNE1 CCNE1 40 32 40 40 35 30 38 30 30 28 36 25 26 20 20 34 Normalized Expression Normalized Expression Normalized Expression 24 Normalized Expression G1 S G2/M G1 S G2/M G1 S G2/M G1 S G2/M Cell Cycle Stage Cell Cycle Stage Cell Cycle Stage Cell Cycle Stage Supplementary Figure 1 | High stochasticity of single-cell gene expression means, as demonstrated by relative expression levels of gene Ccne1 using the mESC-SMARTer data. For every panel, 20 sample cells were randomly selected for each of the three stages, followed by plotting the mean expression levels at each stage. -

Molecular Genetic Characterization of Ptr Toxc-Tsc1 Interaction
MOLECULAR GENETIC CHARACTERIZATION OF PTR TOXC-TSC1 INTERACTION AND COMPARATIVE GENOMICS OF PYRENOPHORA TRITICI-REPENTIS A Dissertation Submitted to the Graduate Faculty of the North Dakota State University of Agriculture and Applied Science By Gayan Kanishka Kariyawasam In Partial Fulfillment of the Requirements for the Degree of DOCTOR OF PHILOSOPHY Major Department: Plant Pathology November 2018 Fargo, North Dakota North Dakota State University Graduate School Title MOLECULAR GENETIC CHARACTERIZATION OF PTR TOXC-TSC1 INTERACTION AND COMPARATIVE GENOMICS OF PYRENOPHORA TRITICI-REPENTIS By Gayan Kanishka Kariyawasam The Supervisory Committee certifies that this disquisition complies with North Dakota State University’s regulations and meets the accepted standards for the degree of DOCTOR OF PHILOSOPHY SUPERVISORY COMMITTEE: Dr. Zhaohui Liu Chair Dr. Shaobin Zhong Dr. Justin D. Faris Dr. Phillip E. McClean Dr. Timothy L. Friesen Approved: November 7, 2018 Jack Rasmussen Date Department Chair ABSTRACT Tan spot of wheat, caused by Pyrenophora tritici-repentis, is an economically important disease worldwide. The disease system is known to involve three pairs of interactions between fungal-produced necrotrophic effectors (NEs) and the wheat sensitivity genes, namely Ptr ToxA- Tsn1, Ptr ToxB-Tsc2 and Ptr ToxC-Tsc1, all of which result in susceptibility. Many lines of evidence also suggested the involvement of additional fungal virulence and host resistance factors. Due to the non-proteinaceous nature, Ptr ToxC, has not been purified and the fungal gene (s) controlling Ptr ToxC production is unknown. The objective for the first part of research is to map the fungal gene (s) controlling Ptr ToxC production. Therefore, A bi-parental fungal population segregating for Ptr ToxC production was first developed from genetically modified heterothallic strains of AR CrossB10 (Ptr ToxC producer) and 86-124 (Ptr ToxC non-producer), and then was genotyped and phenotyped.