Chromosome 6
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

The Genetical Society of Great Britain
Heredity 59 (1987) 151—160 The Genetical Society of Great Britain THEGENETICAL SOCIETY (Abstracts of papers presented at the TVVO HUNDRED AND FIFTH MEETING of the Society held on Friday, 14th and Saturday, 15th November 1986 at UNIVERSITY COLLEGE, LONDON) 1. Selection of somatic cell D. J. Porteous, P. A. Boyd, N. D. Hastie and hybrids with specific chromosome V. van Heyningen content for mapping the WAGR MAC Clinical and Population Cytogenetics Unit, Western General Hospital, Crewe Road, syndrome Edinburgh EH4 2XU. J. M. Fletcher, H. Morrison, J. A. Fantes, Clonedprobes for a number of available chromo- A. Seawright, S. Christie, D. J. Porteous, some ii assigned genes were used to define the N. D. Hastie and V. van Heyningen extent of deletions associated with the Wilms' MAC Clinical and Population Cytogenetics Unit, tumour, aniridia, genitourinary abnormalities and Western General Hospital, Crewe Road, mental retardation (WAGR) syndrome. Establish- Edinburgh EH4 2XU. ing reliable dosage studies for a number of different probes has proved difficult. We have therefore WAGR(Wilms tumour, aniridia, genitourinary abnormalities and mental retardation) syndrome concentrated on segregating the deleted chromo- is frequently associated with deletions on the short some 11 from a number of patients in somatic cell arm of chromosome 11. The deletions vary in size hybrids and analysing DNA from these to produce but always include part of band lipl3. To home a consistent map of chromosome lip. At the same in on the Wilms tumour and aniridia loci the end time we have determined the deletion breakpoints points of the different deletion breakpoints need at a molecular level and shown that the results are to be defined at the DNA level. -

Long-Read Cdna Sequencing Identifies Functional Pseudogenes in the Human Transcriptome Robin-Lee Troskie1, Yohaann Jafrani1, Tim R
Troskie et al. Genome Biology (2021) 22:146 https://doi.org/10.1186/s13059-021-02369-0 SHORT REPORT Open Access Long-read cDNA sequencing identifies functional pseudogenes in the human transcriptome Robin-Lee Troskie1, Yohaann Jafrani1, Tim R. Mercer2, Adam D. Ewing1*, Geoffrey J. Faulkner1,3* and Seth W. Cheetham1* * Correspondence: adam.ewing@ mater.uq.edu.au; faulknergj@gmail. Abstract com; [email protected]. au Pseudogenes are gene copies presumed to mainly be functionless relics of evolution 1Mater Research Institute-University due to acquired deleterious mutations or transcriptional silencing. Using deep full- of Queensland, TRI Building, QLD length PacBio cDNA sequencing of normal human tissues and cancer cell lines, we 4102 Woolloongabba, Australia Full list of author information is identify here hundreds of novel transcribed pseudogenes expressed in tissue-specific available at the end of the article patterns. Some pseudogene transcripts have intact open reading frames and are translated in cultured cells, representing unannotated protein-coding genes. To assess the biological impact of noncoding pseudogenes, we CRISPR-Cas9 delete the nucleus-enriched pseudogene PDCL3P4 and observe hundreds of perturbed genes. This study highlights pseudogenes as a complex and dynamic component of the human transcriptional landscape. Keywords: Pseudogene, PacBio, Long-read, lncRNA, CRISPR Background Pseudogenes are gene copies which are thought to be defective due to frame- disrupting mutations or transcriptional silencing [1, 2]. Most human pseudogenes (72%) are derived from retrotransposition of processed mRNAs, mediated by proteins encoded by the LINE-1 retrotransposon [3, 4]. Due to the loss of parental cis-regula- tory elements, processed pseudogenes were initially presumed to be transcriptionally silent [1] and were excluded from genome-wide functional screens and most transcrip- tome analyses [2]. -

MECHANISMS in ENDOCRINOLOGY: Novel Genetic Causes of Short Stature
J M Wit and others Genetics of short stature 174:4 R145–R173 Review MECHANISMS IN ENDOCRINOLOGY Novel genetic causes of short stature 1 1 2 2 Jan M Wit , Wilma Oostdijk , Monique Losekoot , Hermine A van Duyvenvoorde , Correspondence Claudia A L Ruivenkamp2 and Sarina G Kant2 should be addressed to J M Wit Departments of 1Paediatrics and 2Clinical Genetics, Leiden University Medical Center, PO Box 9600, 2300 RC Leiden, Email The Netherlands [email protected] Abstract The fast technological development, particularly single nucleotide polymorphism array, array-comparative genomic hybridization, and whole exome sequencing, has led to the discovery of many novel genetic causes of growth failure. In this review we discuss a selection of these, according to a diagnostic classification centred on the epiphyseal growth plate. We successively discuss disorders in hormone signalling, paracrine factors, matrix molecules, intracellular pathways, and fundamental cellular processes, followed by chromosomal aberrations including copy number variants (CNVs) and imprinting disorders associated with short stature. Many novel causes of GH deficiency (GHD) as part of combined pituitary hormone deficiency have been uncovered. The most frequent genetic causes of isolated GHD are GH1 and GHRHR defects, but several novel causes have recently been found, such as GHSR, RNPC3, and IFT172 mutations. Besides well-defined causes of GH insensitivity (GHR, STAT5B, IGFALS, IGF1 defects), disorders of NFkB signalling, STAT3 and IGF2 have recently been discovered. Heterozygous IGF1R defects are a relatively frequent cause of prenatal and postnatal growth retardation. TRHA mutations cause a syndromic form of short stature with elevated T3/T4 ratio. Disorders of signalling of various paracrine factors (FGFs, BMPs, WNTs, PTHrP/IHH, and CNP/NPR2) or genetic defects affecting cartilage extracellular matrix usually cause disproportionate short stature. -

Chromosome 6 Abnormalities in Ovarian Surface Epithelial Tumors of Borderline Malignancy Suggest a Genetic Continuum in the Progression Model of Ovarian Neoplasms1
3404 Vol. 7, 3404–3409, November 2001 Clinical Cancer Research Chromosome 6 Abnormalities in Ovarian Surface Epithelial Tumors of Borderline Malignancy Suggest a Genetic Continuum in the Progression Model of Ovarian Neoplasms1 Maria Grazia Tibiletti, Barbara Bernasconi, Conclusion: Alterations in the above-mentioned inter- Daniela Furlan, Paola Bressan, Roberta Cerutti, val are a common finding in advanced ovarian carcinomas but also in benign ovarian cysts, implying that some tumors Carla Facco, Massimo Franchi, Cristina Riva, of borderline malignancy may arise from benign tumors and Raffaella Cinquetti, Carlo Capella, and that malignant ones may evolve from tumors of borderline Roberto Taramelli2 malignancy. Genes located in 6q27 seem to be crucial for this Laboratorio di Anatomia Patologica Ospedale di Circolo, Fondazione mechanism of early events in ovarian tumorigenesis. Macchi, 21100 Varese [M. G. T., C. F., C. R.]; and Dipartimento di Scienze Cliniche e Biologiche [B. B., D. F., P. B., R. Ce., C. C.], INTRODUCTION Clinica Ostetrica e Ginecologica [M. F.], and Dipartimento di Biologia Strutturale e Funzionale [R. Ci., R. T.], Universita` Ovarian cancer represents the most lethal of the gyneco- dell’Insubria, 21100 Varese, Italy logical tumors, and its pathogenesis is poorly understood. More than 80% of malignant tumors of the ovary are of epithelial origin and arise from the surface epithelium (1). Ovarian surface ABSTRACT epithelial tumors are divided into three histological subtypes: Purpose: We used conventional cytogenetics, molecular benign cystoadenoma, TBM,3 and AC, which are related to a cytogenetics, and molecular genetic analyses to study the different biological behavior (1). The molecular events under- pattern of allelic loss on chromosome 6q in a cohort of lying the development of ovarian tumorigenesis are still largely borderline epithelial ovarian tumors. -
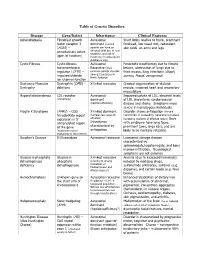
Table of Genetic Disorders Disease Gene/Defect Inheritance Clinical
Table of Genetic Disorders Disease Gene/Defect Inheritance Clinical Features Achondroplasia Fibroblast growth Autosomal Short limbs relative to trunk, prominent factor receptor 3 dominant (normal forehead, low nasal root, redundant (FGR3) – parents can have an skin folds on arms and legs constitutively active affected child due to new mutation, and risk of (gain of function) recurrence in subsequent children is low) Cystic Fibrosis Cystic fibrosis Autosomal Pancreatic insufficiency due to fibrotic transmembrane Recessive (most lesions, obstruction of lungs due to regulator (CFTR) – common genetic disorder thick mucus, lung infections (Staph, impaired chloride among Caucasians in aureus, Pseud. aeruginosa) North America) ion channel function Duchenne Muscular Dystrophin (DMD) - X-linked recessive Gradual degeneration of skeletal Dystrophy deletions muscle, impaired heart and respiratory musculature Hypercholesterolemia LDL receptor Autosomal Impaired uptake of LDL, elevated levels (commonly) dominant of LDL cholesterol, cardiovascular (haploinsufficiency) disease and stroke. Symptoms more severe in homozygous individuals Fragile X Syndrome (FMR1) – CGG X-linked dominant Disorder shows anticipation (female trinucleotide repeat (females less severely transmitters in succeeding generations produce expansion in 5’ affected) increasing numbers of affected males) Boys untranslated region Inheritance with syndrome have long faces, of the gene characterized by prominent jaws, large ears, and are (expansion occurs anticipation likely to be mentally retarded. exclusively in the mother) Gaucher’s Disease Β-Glucosidase Autosomal recessive Lysosomal storage disease characterized by splenomegaly,hepatomegaly, and bone marrow infiltration. Neurological symptoms are not common Glucose 6-phosphate Glucose 6- X-linked recessive Anemia (due to increased hemolysis) dehydrogenase phosphate (prominent among induced by oxidizing drugs, deficiency dehydrogenase individuals of sulfonamide antibiotics, sulfones (e.g. -

Molecular Cytogenetic Characterisation of a Novel De Novo Ring
Pace et al. Molecular Cytogenetics (2017) 10:9 DOI 10.1186/s13039-017-0311-y CASEREPORT Open Access Molecular cytogenetic characterisation of a novel de novo ring chromosome 6 involving a terminal 6p deletion and terminal 6q duplication in the different arms of the same chromosome Nikolai Paul Pace1, Frideriki Maggouta2, Melissa Twigden2 and Isabella Borg1,3,4* Abstract Background: Ring chromosome 6 is a rare sporadic chromosomal abnormality, associated with extreme variability in clinical phenotypes. Most ring chromosomes are known to have deletions on one or both chromosomal arms. Here, we report an atypical and unique ring chromosome 6 involving both a distal deletion and a distal duplication on the different arms of the same chromosome. Case presentation: In a patient with intellectual disability, short stature, microcephaly, facial dysmorphology, congenital heart defects and renovascular disease, a ring chromosome 6 was characterised using array-CGH and dual-colour FISH. The de-novo ring chromosome 6 involved a 1.8 Mb terminal deletion in the distal short arm and a 2.5 Mb duplication in the distal long arm of the same chromosome 6. This results in monosomy for the region 6pter to 6p25.3 and trisomy for the region 6q27 to 6qter. Analysis of genes in these chromosomal regions suggests that haploinsufficiency for FOXC1 and GMDS genes accounts for the cardiac and neurodevelopmental phenotypes in the proband. The ring chromosome 6 reported here is atypical as it involves a unique duplication of the distal long arm. Furthermore, the presence of renovascular disease is also a unique feature identified in this patient. -

WNT16 Is a New Marker of Senescence
Table S1. A. Complete list of 177 genes overexpressed in replicative senescence Value Gene Description UniGene RefSeq 2.440 WNT16 wingless-type MMTV integration site family, member 16 (WNT16), transcript variant 2, mRNA. Hs.272375 NM_016087 2.355 MMP10 matrix metallopeptidase 10 (stromelysin 2) (MMP10), mRNA. Hs.2258 NM_002425 2.344 MMP3 matrix metallopeptidase 3 (stromelysin 1, progelatinase) (MMP3), mRNA. Hs.375129 NM_002422 2.300 HIST1H2AC Histone cluster 1, H2ac Hs.484950 2.134 CLDN1 claudin 1 (CLDN1), mRNA. Hs.439060 NM_021101 2.119 TSPAN13 tetraspanin 13 (TSPAN13), mRNA. Hs.364544 NM_014399 2.112 HIST2H2BE histone cluster 2, H2be (HIST2H2BE), mRNA. Hs.2178 NM_003528 2.070 HIST2H2BE histone cluster 2, H2be (HIST2H2BE), mRNA. Hs.2178 NM_003528 2.026 DCBLD2 discoidin, CUB and LCCL domain containing 2 (DCBLD2), mRNA. Hs.203691 NM_080927 2.007 SERPINB2 serpin peptidase inhibitor, clade B (ovalbumin), member 2 (SERPINB2), mRNA. Hs.594481 NM_002575 2.004 HIST2H2BE histone cluster 2, H2be (HIST2H2BE), mRNA. Hs.2178 NM_003528 1.989 OBFC2A Oligonucleotide/oligosaccharide-binding fold containing 2A Hs.591610 1.962 HIST2H2BE histone cluster 2, H2be (HIST2H2BE), mRNA. Hs.2178 NM_003528 1.947 PLCB4 phospholipase C, beta 4 (PLCB4), transcript variant 2, mRNA. Hs.472101 NM_182797 1.934 PLCB4 phospholipase C, beta 4 (PLCB4), transcript variant 1, mRNA. Hs.472101 NM_000933 1.933 KRTAP1-5 keratin associated protein 1-5 (KRTAP1-5), mRNA. Hs.534499 NM_031957 1.894 HIST2H2BE histone cluster 2, H2be (HIST2H2BE), mRNA. Hs.2178 NM_003528 1.884 CYTL1 cytokine-like 1 (CYTL1), mRNA. Hs.13872 NM_018659 tumor necrosis factor receptor superfamily, member 10d, decoy with truncated death domain (TNFRSF10D), 1.848 TNFRSF10D Hs.213467 NM_003840 mRNA. -

Linkage and Association Analysis of Obesity Traits Reveals Novel Loci and Interactions with Dietary N-3 Fatty Acids in an Alaska Native (Yup’Ik) Population
METABOLISM CLINICAL AND EXPERIMENTAL XX (2015) XXX– XXX Available online at www.sciencedirect.com Metabolism www.metabolismjournal.com Linkage and association analysis of obesity traits reveals novel loci and interactions with dietary n-3 fatty acids in an Alaska Native (Yup’ik) population Laura Kelly Vaughan a, Howard W. Wiener b, Stella Aslibekyan b, David B. Allison c, Peter J. Havel d, Kimber L. Stanhope d, Diane M. O’Brien e, Scarlett E. Hopkins e, Dominick J. Lemas f, Bert B. Boyer e,⁎, Hemant K. Tiwari c a Department of Biology, King University, 1350 King College Rd, Bristol, TN 37620, USA b Department of Epidemiology, University of Alabama at Birmingham, 1665 University Blvd, Birmingham, AL 35294, USA c Section on Statistical Genetics, Department of Biostatistics, University of Alabama at Birmingham, 1665 University Blvd, Birmingham, AL 35294, USA d Departments of Nutrition and Molecular Biosciences, University of California at Davis, 1 Shields Ave, Davis, CA 95616, USA e USACenter for Alaska Native Health Research, Institute of Arctic Biology, 311 Irving I Building, University of Alaska Fairbanks, Fairbanks, AK 99775, USA f Department of Pediatrics, Section of Neonatology, University of Colorado Anschutz Medical Campus, 13123 East 16th Ave, Aurora, CO 80045, USA ARTICLE INFO ABSTRACT Article history: Objective. To identify novel genetic markers of obesity-related traits and to identify gene- Received 16 June 2014 diet interactions with n-3 polyunsaturated fatty acid (n-3 PUFA) intake in Yup’ik people. Accepted 28 February 2015 Material and methods. We measured body composition, plasma adipokines and ghrelin in 982 participants enrolled in the Center for Alaska Native Health Research (CANHR) Study. -

Isolation of Chromosome-Specific Ests by Microdissection-Mediated Cdna Capture Edgardo Gracia, 1-3 Michael E
Downloaded from genome.cshlp.org on September 26, 2021 - Published by Cold Spring Harbor Laboratory Press RESEARCH Isolation of Chromosome-Specific ESTs by Microdissection-Mediated cDNA Capture Edgardo Gracia, 1-3 Michael E. Ray, 1-3 Mihael H. Polymeropoulos, 4 Anindya Dehejia, 4 Paul S. Meltzer, 3 and Jeffrey M. Trent 3's 1Department of Human Genetics, The University of Michigan Medical School, Medical Science II M4708, Ann Arbor, Michigan 48109; 3Laboratory of Cancer Genetics, 4 Laboratory of Genetic Disease Research, National Center for Human Genome Research, National Institutes of Health, Bethesda, Maryland 20892 Despite dramatic advances in the identification of human expressed sequence tags (ESTs), techniques that facilitate isolation of chromosome or chromosome band-specific ESTs would be of considerable value. This report demonstrates the feasibility of identifying chromosome-specific ESTs following microdissection of a single-copy chromosome region. For this study, a reduced complexity cDNA library was linkered and hybridized to normal human metaphase chromosomes. After stringency washes, the entire long arm of chromosome 6 (6q) was microdissected. Following PCR amplification using linker-specific primers, captured cDNAs were subcloned and 187 individual clones picked at random. These 187 clones were then sorted by filter cross-hybridization into 34 unique groups. Of these 34 groups, 19 (56%) mapped to chromosome 6 by Southern blot. We identified three previously known genes, human cytovillin (ezrin) mapped previously to 6q25-26, human cardiac gap junction protein (connexin 43) mapped previously to 6q21-23.2 and prolyloligopeptidase, which had not been mapped previously. BLASTN identified three clone groups with homology to known ESTs and 12 representing novel cDNA sequences. -

Gene and Alternative Splicing Annotation with AIR
Downloaded from genome.cshlp.org on May 8, 2012 - Published by Cold Spring Harbor Laboratory Press Gene and alternative splicing annotation with AIR Liliana Florea, Valentina Di Francesco, Jason Miller, et al. Genome Res. 2005 15: 54-66 Access the most recent version at doi:10.1101/gr.2889405 Supplemental http://genome.cshlp.org/content/suppl/2004/12/08/15.1.54.DC1.html Material References This article cites 49 articles, 34 of which can be accessed free at: http://genome.cshlp.org/content/15/1/54.full.html#ref-list-1 Article cited in: http://genome.cshlp.org/content/15/1/54.full.html#related-urls Creative This article is distributed exclusively by Cold Spring Harbor Laboratory Press Commons for the first six months after the full-issue publication date (see License http://genome.cshlp.org/site/misc/terms.xhtml). After six months, it is available under a Creative Commons License (Attribution-NonCommercial 3.0 Unported License), as described at http://creativecommons.org/licenses/by-nc/3.0/. Email alerting Receive free email alerts when new articles cite this article - sign up in the box at the service top right corner of the article or click here To subscribe to Genome Research go to: http://genome.cshlp.org/subscriptions © 2005, Published by Cold Spring Harbor Laboratory Press Downloaded from genome.cshlp.org on May 8, 2012 - Published by Cold Spring Harbor Laboratory Press Methods Gene and alternative splicing annotation with AIR Liliana Florea,1,4,5 Valentina Di Francesco,2 Jason Miller,1 Russell Turner,1 Alison Yao,2 Michael Harris,2 Brian Walenz,1 Clark Mobarry,1 Gennady V. -

Spring 2003 Final3
NCBI News National Center for Biotechnology Information National Library of Medicine National Institutes of Health Department of Health and Human Services Spring 2003 [ A Field Guide to GenBank® and NCBI Resources: First Version of Human NCBI’s Scientific Outreach and Training Program Genome Reference Sequence Debuts on Biological sequence and structure use of NCBI databases and tools. DNA’s 50th information are now used in nearly The course, called “A Field Guide every field of biological research. A to GenBank and NCBI Resources”, working knowledge of these resources is designed especially for biologists April 14, 2003 marked the and standard computational biology who work at the bench or in the field 50th anniversary of the tools are an essential part of every but use sequence and structure data description of the structure biologist’s toolkit. However, keeping in their research. All researchers, of DNA and also saw the up with these databases and tools can educators and students who work release of the first version of be challenging in this period of rapidly with biological sequence and structure the 3 billion base pair refer- changing bioinformatics resources. data should find this to be a useful ence sequence of the human introduction and survey of the genome. Annotations to the In order to help researchers keep available NCBI tools and databases. raw sequence made public on April 14 abreast of enhancements and the Because of the rapid expansion of were released on April 29 when the increasing diversity of NCBI molecu- the resources, even experienced NCBI reference genome, NCBI build 33, lar biology resources, the NCBI users will likely learn something new appeared in the NCBI Map Viewer. -

A Literature-Based Database for Cell Senescence Genes and Its Application to Identify Critical Cell Aging Pathways and Associated Diseases
Citation: Cell Death and Disease (2016) 7, e2053; doi:10.1038/cddis.2015.414 OPEN & 2016 Macmillan Publishers Limited All rights reserved 2041-4889/16 www.nature.com/cddis CSGene: a literature-based database for cell senescence genes and its application to identify critical cell aging pathways and associated diseases M Zhao1, L Chen2 and H Qu*,2 Cell senescence is a cellular process in which normal diploid cells cease to replicate and is a major driving force for human cancers and aging-associated diseases. Recent studies on cell senescence have identified many new genetic components and pathways that control cell aging. However, there is no comprehensive resource for cell senescence that integrates various genetic studies and relationships with cell senescence, and the risk associated with complex diseases such as cancer is still unexplored. We have developed the first literature-based gene resource for exploring cell senescence genes, CSGene. We complied 504 experimentally verified genes from public data resources and published literature. Pathway analyses highlighted the prominent roles of cell senescence genes in the control of rRNA gene transcription and unusual rDNA repeat that constitute a center for the stability of the whole genome. We also found a strong association of cell senescence with HIV-1 infection and viral carcinogenesis that are mainly related to promoter/enhancer binding and chromatin modification processes. Moreover, pan-cancer mutation and network analysis also identified common cell aging mechanisms in cancers and uncovered a highly modular network structure. These results highlight the utility of CSGene for elucidating the complex cellular events of cell senescence.